1. INTRODUCTION
Gene therapy has swiftly become a fundamental aspect of precision medicine by directly modifying, replacing, or silencing faulty genes [1]. The domain gained clinical significance following the inaugural successful human gene transfer in 1989, followed by the pivotal correction of adenosine deaminase–severe combined immunodeficiency in 1990, which exhibited lasting immune reconstitution and validated the concept of genomic intervention [2]. Since that time, regulatory and commercial investments have intensified, particularly due to the European Commission’s 2023 reclassification of Gene Therapy Medicinal Products (GTMPs), indicating the integration of genome-modifying medicines into standard clinical pipelines [3].
Notwithstanding significant advancements, the clinical efficacy of gene therapy remains limited by the lack of a universal delivery system that can accommodate diverse cargo types, achieve high editing efficiency, and ensure safe biodistribution [4]. Viral vectors continue to be the cornerstone of the field, offering clinical advantages in hemophilia, retinal dystrophies, and spinal muscular atrophy (SMA); nonetheless, their application is hindered by insertional mutagenesis, immunogenicity, manufacturing challenges, and rigorous regulatory scrutiny [5,6]. Non-viral systems, such as synthetic nanoparticles and lipoplexes, offer enhanced design flexibility and diminished immunogenicity; however, they frequently encounter challenges related to limited endosomal escape, brief circulation durations, and inadequate delivery of substantial cargos such as CRISPR/Cas9 plasmids or messenger RNA (mRNA) expression systems [7]. These obstacles have gained significance as gene therapy advances into intricate disease categories.
In oncology, gene delivery technologies now focus on KRAS^G12D (Kirsten rat sarcoma viral oncogene homolog, G12D variant), BRAF (B-Raf proto-oncogene, serine/threonine kinase), EGFR (epidermal growth factor receptor), and TP53 (tumor protein p53) pathways utilizing CRISPR, small interfering RNA (siRNA), and mRNA replacement methodologies in preclinical and early clinical development [8-10]. In uncommon diseases, therapies for Duchenne muscular dystrophy (DMD; Duchenne muscular dystrophy and dystrophin), SMA (SMN1/2; survival motor neuron 1/2), transthyretin amyloidosis (TTR; transthyretin), and cystic fibrosis (CFTR; cystic fibrosis transmembrane conductance regulator) necessitate extensive coding sequences that surpass the packaging capacities of numerous viral vectors [11]. Neurological illnesses present a further obstacle – the blood-brain barrier (BBB) – which limits absorption by neurons and glia, necessitating low-toxicity carriers suitable for repeated administration [12].
The unaddressed requirements have rekindled interest in exosomes – endogenous extracellular vesicles (EVs) derived from endosomes that demonstrate little immunogenicity, inherent biocompatibility, and natural affinity for particular tissues [13]. Exosomes contain and safeguard nucleic acids, proteins, and lipids from degradation, circulate persistently in serum, and possess the distinctive capability to traverse endothelial and brain barriers [14]. Research has shown that engineered exosomes can effectively transport CRISPR/Cas9 ribonucleoproteins (RNPs), tumor-suppressive microRNA (miRNAs), and mRNA-encoded therapeutic proteins, exhibiting enhanced cellular uptake, diminished cytokine induction, and advantageous biodistribution relative to numerous synthetic carriers [15-17]. Initial preclinical models demonstrate the inhibition of oncogenic signaling, restoration of protein-deficiency symptoms, and control of neuroinflammation, underscoring their adaptability as a gene delivery platform [18].
This paper posits that modified exosomes can alleviate significant translational challenges of gene therapy, such as cargo degradation, inadequate tissue specificity, and dose-limiting tolerability. We assess progress in cargo loading, membrane functionalization, and the creation of distribution strategies across several clinical situations. We examine how artificial intelligence (AI) can facilitate variant interpretation and treatment selection by correlating genomic information with modality selection and delivery parameters. The research collectively endorses a cohesive developmental trajectory for exosome-mediated gene delivery that is progressively customized for specific diseases and focused toward translation.
Recent evaluations on exosome-based CRISPR delivery and AI in nanomedicine frequently lack a clear integration that links genetics to vector design and translational controls. This article integrates the convergence of AI and exosomes by linking AI-assisted whole-exome sequencing (WES) prioritization to specific engineering decisions (cargo modality, loading approach, targeting design, and potency assays) and structuring these components into a workflow designed to enhance reproducibility and clinical preparedness [15,19,20].
2. EXOSOMES: BIOGENESIS, COMPOSITION, AND FUNCTIONAL ROLES
This section summarizes the synthesis of EVs/exosomes and the sorting of their payload, providing adequate information for engineering and delivery design. Comprehensive mechanistic pathways are not thoroughly examined to preserve concentration on the AI-driven treatment process. Exosomes are a specific subtype of EVs, measuring 50–150 nm, originating from the endosomal pathway and discharged by most cell types into bodily fluids. Exosome biogenesis, selective cargo loading, and regulated secretion are essential for their function as intercellular messengers and innovative therapeutic vehicles [21]. This section consolidates the essential mechanistic elements of exosome formation, delineates the specific molecular constituents that provide stability and tropism, and elucidates how these components influence their functions in physiology and pathology. According to International Society for Extracellular Vesicles (ISEV)’s Minimal information for studies of extracellular vesicles (MISEV) criteria, we employ the word EVs as a general designation and reserve the term “exosomes” for instances where endosomal origin is empirically validated [22,23].
A systematic literature search was performed on PubMed/MEDLINE, Scopus, and Web of Science for publications spanning 2014–2025. Boolean keyword combinations comprised exosome OR EV AND (gene therapy OR CRISPR OR mRNA OR siRNA OR plasmid DNA [pDNA]) AND (loading OR targeting OR surface modification OR delivery). Supplementary records were discovered by reference snowballing from pivotal review articles and original research investigations.
Eligible research needed to document the preclinical or clinical application of exosomes for genetic material delivery, outline a loading or targeting method, or present in vivo efficacy or biodistribution data. Research solely concentrating on diagnostic biomarkers, non-vesicular nanoparticles, or non-genetic therapies was omitted. Data from the included publications were retrieved regarding cargo type, loading method, cell source, surface-engineering strategy, illness model, and treatment outcome. This methodical methodology was employed to deliver a representative and equitable overview of the present status of synthetic exosome research.
2.1. Biogenesis, Molecular Structure, and Cargo Composition
Exosomes are nanoscale EVs (50–150 nm) derived from the endosomal system and secreted by several mammalian cell types into bodily fluids [24]. Their creation initiates with the inward budding of the plasma membrane, resulting in the formation of early endosomes that subsequently mature into multivesicular bodies (MVBs). In MVBs, the repetitive invagination of the limiting membrane generates intraluminal vesicles (ILVs), which serve as progenitors of secreted exosomes [25]. Two well-defined cargo-sorting processes govern this operation. The ESCRT-dependent process identifies ubiquitinated proteins and selectively integrates them into ILVs, whereas ESCRT-independent methods utilize ceramides, tetraspanins, and lipid microdomains to facilitate vesicle formation [26,27]. Collectively, these mechanisms establish a controlled mechanism for the encapsulation of nucleic acids and proteins, differentiating exosomes from other subclasses of EVs.
The molecular structure of exosomes indicates their endosomal origin. Their membranes are abundant in cholesterol, sphingomyelin, and phosphatidylserine, which contribute to the stabilization of circulating vesicles [28]. Canonical exosomal markers, including CD9, CD63, CD81, Alix, and TSG101, are commonly employed for identification and have a role in trafficking and uptake [29]. Exosomes contain mRNAs, miRNAs, siRNAs, long non-coding RNAs, DNA fragments, and a diverse proteome that includes signaling molecules, metabolic enzymes, heat shock proteins, and disease-related factors [21]. This cargo is meticulously organized. Tumor-derived exosomes selectively accumulate oncogenic transcripts and proteins that facilitate angiogenesis and immune evasion, while immune-cell exosomes transport immunoregulatory miRNAs and cytokine-modulating proteins [30]. Neural exosomes are distinguished by their ability to traverse the BBB, a characteristic not possessed by the majority of synthesized nanoparticles [13].
These biological characteristics provide direct therapeutic significance. The lipid bilayer safeguards nucleic acids against degradation by circulating endonucleases, facilitating the systemic transport of mRNA, miRNA, or CRISPR/Cas9 components [31]. Surface tetraspanins and integrins affect biodistribution, facilitating preferential tropism for liver, brain, lung, or tumor tissues [32]. In preclinical models, engineered exosomes have been utilized to deliver tumor-suppressive miRNAs (e.g., miR-34a and miR-143), CRISPR components targeting KRAS^G12D or EGFR mutations, and mRNA encoding functional proteins, exhibiting improved cellular uptake and diminished immunogenicity relative to numerous synthetic carriers [33].
Exosomes transport a varied array of physiologically active substances, such as nucleic acids, proteins, lipids, and metabolites, allowing them to serve as specialized mediators of intercellular communication [24]. In contrast to synthesized nanoparticles, exosomes selectively encapsulate genetic and protein content through directed sorting mechanisms rather than through passive encapsulation. The selective loading is regulated by RNA-binding proteins, lipid raft domains, tetraspanins, and ESCRT machinery, all of which enhance the concentration of functional molecules [26,27]. Consequently, exosomal cargos possess biological significance and frequently indicate the metabolic and pathological condition of the originating cell.
Nucleic acid cargo comprises mRNA, miRNA, siRNA, long non-coding RNA, and double-stranded DNA fragments [21]. These molecules are enclosed within a lipid bilayer that safeguards them against nuclease breakdown in circulation [28]. Numerous treatment trials have proven this benefit. Exosomal administration of miR-34a and miR-143 inhibits oncogenic pathways in breast and colorectal cancer models, whereas exosome-mediated siRNA targeting KRAS^G12D, BRAF, or EGFR reduces tumor growth and spreading potential [33]. In metabolic and neurological disorders, exosomes with mRNA that encodes defective or malfunctioning proteins have reinstated physiological expression in vivo, demonstrating proof-of-concept for protein replacement by exosomal mRNA [13].
Protein cargo additionally enhances therapeutic potential. The exosomal proteome includes tetraspanins (CD9, CD63, and CD81), heat-shock proteins, signal transduction mediators, metabolic enzymes, and disease-specific factors [29]. Tumor-derived exosomes convey oncogenic proteins and immunosuppressive ligands that alter the milieu to promote tumor viability [30]. In contrast, modified exosomes can be infused with therapeutic proteins – such as pro-apoptotic factors, cytokine modulators, and CRISPR-associated nucleases – to alter target-cell behaviors. The stable encapsulation of Cas9 protein and single-guide RNA within exosomes facilitates non-viral CRISPR delivery, minimizing the danger of genome integration and decreasing immunogenicity relative to viral vectors [31].
The composition of lipids is a significant factor influencing cargo stability and cellular absorption. Elevated concentrations of cholesterol, sphingomyelin, and phosphatidylserine enhance membrane stiffness, extend circulation duration, and facilitate interactions with scavenger receptors and integrins on recipient cells [32]. This inherent biodistribution has been utilized to accomplish tailored delivery. Macrophage-derived exosomes preferentially aggregate in inflamed tissues, but neural exosomes possess the capability to traverse the BBB, a feat seldom accomplished by synthetic nanocarriers [13].
Collectively, these characteristics position exosomes as a physiologically specialized medium for the secure transport of nucleic acids and proteins, facilitating precise regulation of gene expression in target tissues. The selective packing, stability in circulation, and inherent tissue tropism present a persuasive justification for the creation of exosomes as advanced gene therapy vectors [34].
2.2. Functional Roles of Exosomes in Gene Transfer
Exosomes have emerged as a very promising biological platform for gene transport due to their inherent ability to encapsulate nucleic acids, safeguard them from destruction, and convey functional genetic material to destination cells [13]. Their lipid bilayer confers resistance to endonucleases and promotes fusion or endocytosis, hence facilitating the effective cytoplasmic release of mRNA, miRNA, siRNA, and CRISPR-associated components [31]. In contrast to typical synthetic nanoparticles, exosomes possess inherent surface ligands – tetraspanins, integrins, and adhesion molecules – that direct biodistribution and improve cellular uptake across physiological barriers, such as the BBB [29,32].
These systems have demonstrated therapeutic significance across various illness categories. In cancer models, exosome-mediated delivery of siRNA and miRNA targeting KRAS^G12D, EGFR, and BRAF attenuates oncogenic signaling and reduces tumor proliferation [33]. Engineered exosomes transporting tumor-suppressive mRNAs reinstate functional protein expression in colorectal, lung, and breast cancers, while reducing inflammatory cytokine responses commonly associated with synthetic carriers [34]. In addition to cancer, exosomes have successfully administered therapeutic mRNA for SMN1, CFTR, and TTR in neurological and metabolic illnesses, exhibiting enhanced stability and reduced immunogenicity in vivo relative to recurrent viral dosing [21,28]. Their capacity to protect substantial cargos facilitates non-viral delivery of CRISPR/Cas9 proteins or plasmids, mitigating the dangers of genomic integration and permitting temporary gene editing [31].
Exosomes are distinguished from existing non-viral systems by their natural biocompatibility, low immunogenicity, and selective tropism. Engineered exosomes can utilize cell-type-specific surface indicators instead of necessitating artificial coatings or targeting peptides [35]. Macrophage-derived vesicles specifically target inflamed tissues, whereas neural exosomes traverse the BBB and aggregate in microglia and neurons [13]. These attributes provide exosomes versatile and scalable vehicles for genetic therapies, especially in conditions where repeated administration, immunological reactivity, or targeted tissue delivery pose significant challenges for viral vectors.
3. MECHANISMS OF GENE DELIVERY THROUGH EXOSOMES
Exosomes have emerged as intriguing vehicles for gene delivery due to their inherent biological versatility. They inherently encapsulate a diverse array of molecular cargos, such as mRNAs, short regulatory RNAs, and proteins, which can be selectively loaded and delivered to recipient cells [35]. Exosomes can traverse restrictive physiological barriers, such as the BBB, and can infiltrate poorly vascularized tissues, including cartilage. On arriving at their destination, they frequently remain in the target microenvironment for a protracted duration, facilitating prolonged therapeutic efficacy [36].
Exosomes exhibit inherent targeting capabilities, and their biological origin significantly affects their biodistribution. They demonstrate context-dependent tropism influenced by environmental signals, cellular origin, and surface markers, facilitating their application in targeted therapy [37]. For instance, exosomes produced from mesenchymal stem cells exhibit a preference for damaged tissues [38], while exosomes originating from tumors engage intensively with the tumor microenvironment [39]. Exosomes possess the capability to traverse the BBB and specifically target neurons and glial cells [40].
The precision of exosome targeting is determined by the composition of ligands and receptors found on both exosomes and recipient cells [41]. Exosomes can transport MHC-peptide complexes and tumor necrosis factor ligands, which engage co-receptors on target cells and trigger specific signaling cascades [42]. These vesicles adhere to cell surfaces mainly through receptor-ligand interactions, influencing cellular metabolism and subsequent biological reactions. A comprehensive comprehension of ligand–receptor dynamics is crucial for enhancing the therapeutic application of exosomes, especially in targeted clinical settings [43].
Cargo loading denotes the integration of therapeutic biomolecules, including proteins and nucleic acids, into exosomes for clinical or research purposes. Loading strategies are typically categorized into pre-isolation and post-isolation methods [44]. Endogenous (pre-loading) methods entail modifying donor cells to express or absorb therapeutic substances before the collection of exosomes. Transfection techniques facilitate the introduction of nucleic acids or proteins into parental cells, enabling the cargo to be spontaneously contained in exosomes during biogenesis. This strategy, while biologically pertinent, typically demonstrates reduced loading efficiency relative to exogenous strategies [21].
Exogenous (post-loading) techniques involve the introduction of medicinal compounds into pure exosomes following their isolation [21]. Various approaches have been devised to augment membrane permeability and facilitate cargo internalization. Passive incubation subjects exosomes to elevated concentrations of therapeutic compounds, facilitating diffusion through the vesicle membrane; still, the loading efficiency for nucleic acids and small molecules is typically suboptimal [45]. Electroporation temporarily breaches the exosomal membrane using an electric field, forming pores that facilitate cargo entry. This technique has been employed to encapsulate insulin, thereafter incubated at 37°C for 1 h, and generally attains superior loading efficiency compared to passive incubation [44]. Sonication utilizes ultrasonic shear pressures to transiently disrupt membrane structure, hence augmenting permeability and improve loading efficiency compared to incubation-based methods [46]. Freeze–thaw cycling, which consistently modifies membrane fluidity through temperature fluctuations, can facilitate the structural integration of therapeutic cargo, although potentially undermining vesicle stability [47]. Extrusion propels exosomes through membranes with certain pore dimensions, enabling the mechanical integration of cargo; although efficient for diverse therapeutic agents, it may compromise vesicle morphology or membrane integrity [46].
Every technique has unique benefits and drawbacks. Consequently, the choice of a suitable loading method must be informed by the therapeutic drug, cargo properties, and the intended biological effect [21].
A unique strategy combines endogenous and exogenous loading techniques to improve cargo integration into exosomes. This method involves engineering producer cells to express a DNA aptamer that binds target mRNAs, facilitating intracellular sequestration of the transcripts. The resultant mRNA–aptamer complexes are subsequently encapsulated into exosomes through a CD9–zinc finger protein fusion system [48,49]. On entering the exosomes, the mRNA is liberated from the aptamer complex, and electroporation facilitates effective delivery and translation of the mRNA in recipient cells, both in vitro and in vivo. This dual-loading technique significantly enhances the efficiency of therapeutic cargo delivery by concurrently affecting endogenous capture and exogenous insertion [21,50]. These combined loading systems utilize both endogenous and exogenous mechanisms, addressing individual constraints and enhancing the practical utility of exosome-based medicines.
Exosomes facilitate the transfer of pDNA containing therapeutic genes to recipient cells, leading to improved gene expression and decreased immunogenicity relative to numerous traditional delivery techniques [51]. They are also effective transporters of therapeutic RNA molecules, including as mRNA, miRNA, and siRNA. The intrinsic lipid and protein composition of exosomes safeguards RNA cargo from enzymatic destruction and facilitates effective uptake by target cells [52].
In cancer treatment, exosome-mediated siRNA transport has demonstrated the ability to suppress oncogene activity, while the exosomal transportation of therapeutic mRNA facilitates the synthesis of functional proteins, constituting a significant method for disorders resulting from protein shortage [45]. Exosomes can transport both DNA and RNA concurrently, presenting opportunities for combination gene therapy that targets several biological pathways, therefore enhancing therapeutic effects, especially in oncology [53].
Transporting RNAs through exosomes presents difficulties, especially in the effective loading of substantial RNA molecules. Electroporation effectively facilitates the loading of siRNA and miRNA, but frequently proves inadequate for the efficient loading of mRNA [51]. Exosomes have effectively transported siRNA to inhibit specific genes in several conditions, including malignancies and neurological disorders. Exosome-delivered siRNA can effectively target the gene with little off-target effects [54].
Exosomes contain and safeguard miRNAs, facilitating their delivery to target cells and thus influencing gene expression and cellular activity. Nonetheless, other obstacles continue to hinder exosome-mediated RNA distribution, such as the substantial size and intricate structure of RNA molecules. Future research must concentrate on enhancing techniques for exosome isolation, characterization, and purification to advance therapeutic uses. Engineering exosomes for targeted delivery and enhanced cargo loading can further improve exosome-mediated RNA transport [49]. The inherent characteristics of exosomes provide distinct benefits for RNA transport in comparison to synthetic delivery systems [17].
Exosomes are widely viewed as attractive natural vectors for pDNA and CRISPR/Cas9 components because to their biocompatibility, low immunogenicity, and inherent capacity to cross physiological barriers. The emergence of CRISPR/Cas9 has significantly progressed genome editing, and its incorporation with exosome-mediated transportation presents an effective method for conveying gene-editing tools into target cells [55]. Exosomes can be modified to incorporate CRISPR/Cas9 plasmids or RNP complexes, enhancing intracellular delivery and minimizing off-target editing occurrences. Exosomes containing CRISPR/Cas9 pDNA aimed at the oncogenic KRAS^G12D mutation in pancreatic cancer cells have demonstrated the ability to impede cellular proliferation and decrease tumor growth in vivo [53,56].
Notwithstanding their therapeutic promise, exosome-mediated transport of pDNA and CRISPR/Cas9 systems continues to encounter numerous obstacles. The efficacy of loading substantial genetic components into exosomes is still constrained, and the preservation of stability and functional integrity of these cargos during transit has yet to be thoroughly optimized. Moreover, attaining accurate and tissue-specific targeting necessitates ongoing technological progress and research [53,57].
Advancements in exosome engineering are crucial for converting exosome-mediated transport of pDNA and CRISPR/Cas9 components into viable therapeutic treatments. Figure 1 illustrates the mechanistic foundation of exosome-mediated gene delivery, detailing the various stages required for the transfer of genetic material to mammalian cells.
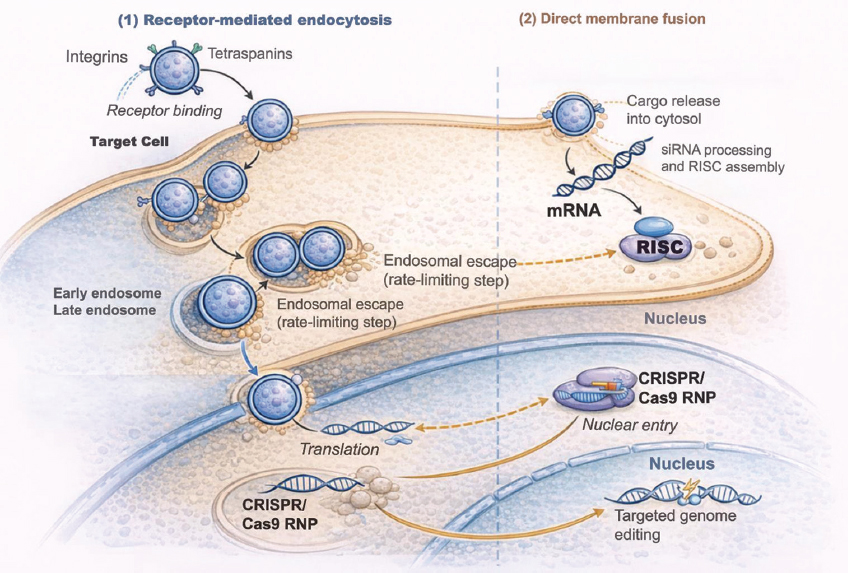 | Figure 1: Mechanistic pathway of exosome-mediated gene delivery in mammalian cells. Exosomes are nanoscale vesicles composed of lipid bilayers, loaded with surface integrins and tetraspanins (CD9, CD63, and CD81), which enable specific binding to receptors on the plasma membrane of target cells. Two principal entrance pathways are illustrated: (1) receptor-mediated endocytosis, resulting in vesicle internalization and transport through early endosomes, and (2) direct membrane fusion, facilitating prompt cytosolic release of cargo. On entering the cytoplasm, exosomes release nucleic acids, such as messenger RNA (mRNA), small interfering RNA (siRNA), and CRISPR/Cas9 RNP complexes or plasmids. These cargos facilitate specific intracellular mechanisms: Exosomal mRNA is translated by ribosomes; siRNA interacts with the RNA-induced silencing complex to execute post-transcriptional gene silencing; and CRISPR/Cas9 complexes migrate to the nucleus to conduct targeted genome editing. The diagram emphasizes organelles (nucleus, endosomes, and ribosomes) and directing arrows to depict the sequential stages of delivery and functional results. [Click here to view] |
4. METHODS AND TECHNIQUES FOR ENGINEERING EXOSOMES FOR GENE THERAPY
Exosome modification often initiates at the donor-cell level, when cells are genetically altered to express therapeutic nucleic acids [20]. Donor cells can generate exosomes that naturally incorporate mRNAs, siRNAs, or CRISPR/Cas components during exosome biogenesis through plasmid transfection or viral transduction [20]. This approach guarantees inherent packaging integrity and reduces external interference with vesicles [20]. The payload is intrinsically sorted into exosomes, resulting in vesicles that are biocompatible, stable, and possess minimal immunogenicity [58]. These exosomes preserve their membrane proteins and targeting capabilities, rendering them appealing carriers for gene therapy in comparison to synthetic vectors or viruses [59].
A multitude of studies have proven the efficacy of this strategy. For instance, mesenchymal stem cells modified to overexpress CXCR4 produced exosomes with enhanced targeting to tumor tissues [60]. These vesicles successfully administered siRNA directed against survivin, leading to gene silencing and tumor inhibition in vivo [60].
4.1. Engineering Approaches for Cargo Loading and Surface Alteration
The genetic modification of donor cells, specifically HEK293/HEK293T lines, facilitates the reliable and stable incorporation of therapeutic RNAs – such as miR-34a or miR-21 – into exosomes [61]. This approach guarantees biocompatibility and minimal immunogenicity while facilitating scalable and regulated production, as evidenced by lentiviral transduction techniques that improve RNA packaging and functional delivery in therapeutic applications such as cancer treatment and spinal cord repair [61].
An advanced technique entails an “all-in-one exosome sorter” system, wherein donor cells are co-transfected with therapeutic RNA and a sorting scaffold protein (e.g., FLAG-tagged tetraspanin (TCS)) [62]. This architecture facilitates selective RNA packaging and magnetic bead purification, resulting in enhanced treatment efficacy, as demonstrated in a mouse model of familial hypercholesterolemia [62]. The utilization of genetically modified donor cells facilitates disease-specific tailoring of exosome cargo and surface proteins, hence improving efficacy and targeting in clinical applications [36].
To address the constraints and variability of endogenous cargo encapsulation, many external loading methodologies have been developed. The methods encompass electroporation, sonication, incubation, freeze-thaw cycling, and membrane extrusion, as detailed in Table 1.
Table 1: Overview of exosome loading techniques, their underlying mechanisms, and compatible cargo types for therapeutic applications.
| Technique | Mechanism | Cargo types | References |
|---|---|---|---|
| Electroporation | Transient pore formation in exosome membranes through electric pulses | siRNA, mRNA, and plasmid DNA | [45] |
| Sonication | Ultrasonic shear force disrupts the membrane for cargo entry | Proteins, small molecules, and miRNAs | [44] |
| Freeze–Thaw Cycling | Temperature cycling induces membrane permeability | Proteins, drugs, and miRNAs | [63] |
| Extrusion | Mechanical forces cargo into exosomes through membrane pores | Catalase and drugs | [44] |
| Lipophilic Conjugation | Chemical linkage of hydrophobic cargo to membrane lipids | Lipophilic nucleic acids and small molecules | [64] |
| Microfluidic Loading | Nanochannel shear flow permeabilizes membranes for controlled entry | mRNA, antibodies, and targeted ligands | [65] |
| Dialysis-Based Loading | Osmotic pressure drives passive cargo loading through semipermeable membranes | Small molecules and proteins | [66] |
| Click Chemistry | Covalent conjugation of ligands or drugs through bioorthogonal reactions | Targeting ligands, dyes, and peptides | [20] |
| In situ Synthesis | Exogenous molecules are synthesized directly within or near vesicles | Nanoparticles and polymers | [67] |
| Hybrid exosome mimics | Fusion of natural exosomes with liposomes or synthetic nanoparticles | Hydrophobic and hydrophilic drugs | [68] |
| Aptamer-mediated conjugation | Surface modification with nucleic acid aptamers to enhance targeting | Small RNAs and tumor-specific ligands | [69] |
| ESCRT-independent sorting | Ceramide and lipid raft pathways mediate selective cargo inclusion | Endogenous proteins and miRNAs | [70] |
| Nanofluidic nanoporation | Mechanical shear in nanochannels enables high-throughput cargo entry | siRNA, mRNA, and small drugs | [71] |
siRNA: Small interfering RNA, mRNA: Messenger RNA, miRNA: MicroRNA, CD63: Cluster of Differentiation 63, GFP: Green Fluorescent Protein, ssDNA: Single-stranded DNA, gRNA: Guide RNA, RNP: Ribonucleoprotein, HBV: Hepatitis B Virus, HPV: Human papillomavirus, KRAS: Kirsten rat sarcoma viral oncogene homolog, ESCRT: Endosomal sorting complex required for transport.
The variety of exosome cargo-loading methods demonstrates advancing efforts to enhance therapeutic effectiveness, stability, and targeting accuracy [44]. Table 1 illustrates that each method presents unique advantages and disadvantages, frequently weighing loading efficiency against the maintenance of vesicle structure. Physical methods such as electroporation and sonication are frequently utilized for nucleic acid delivery due to their simplicity and moderate efficacy; however, they may jeopardize membrane integrity [44,45]. Conversely, chemical methods, such lipophilic conjugation, facilitate enhanced encapsulation of hydrophobic compounds with minimum vesicle disruption [64], but they frequently necessitate meticulous tuning to preserve cargo bioactivity.
Dialysis and extrusion offer gentler methods for passive loading, especially for small molecules and proteins, whereas novel techniques like in situ synthesis facilitate the intracellular creation of functional nanocomplexes that are inherently encapsulated in exosomes during biogenesis [36,44]. The selection of approach should be determined by the physicochemical properties of the cargo, the specific therapeutic application, and the necessary scalability for clinical implementation.
The surface engineering of exosomes has facilitated targeted delivery to certain cell types or tissues, greatly enhancing their therapeutic efficacy. An exemplary instance is the application of the CP05 peptide, which adheres to the exosomal surface marker CD63 [72]. Gao et al. conjugated CP05 with a muscle-targeting peptide and loaded the modified exosomes with splice-correcting oligonucleotides, leading to restored dystrophin expression and enhanced muscle function in dystrophic mice, underscoring the precision and efficacy of peptide-based targeting [72].
Aptamer-based therapies have demonstrated therapeutic potential. Hosseini et al. employed EDC/NHS chemistry to link the nucleolin-binding aptamer AS1411 to doxorubicin-loaded exosomes, facilitating targeted uptake by colorectal cancer cells and significant tumor reduction in vivo [73]. Chemical methodologies like click chemistry enhance the functionalization repertoire; Smyth et al. illustrated that azide–alkyne cycloaddition can securely conjugate ligands to exosome surfaces while preserving vesicle integrity, facilitating customizable bioorthogonal exosome labeling [74].
Hung and Leonard tackled peptide degradation in exosome display by conjugating targeting peptides with glycosylation motifs, thereby enhancing ligand stability and delivery to neuroblastoma cells [75]. Collectively, these cases demonstrate how peptide anchoring, aptamer functionalization, antibody conjugation, and chemical ligation constitute a diverse toolset for attaining stable, efficient, and tissue-selective exosomal targeting.
4.2. Exosome-Facilitated Transmission of CRISPR/Cas Systems
The exosome-mediated delivery of CRISPR/Cas9 systems is swiftly progressing as a non-viral, biocompatible method for precise gene editing in many disorders [76]. Numerous investigations have confirmed that exosomes may transport and distribute CRISPR/Cas9 components – including plasmids, mRNA, and RNP complexes – through both endogenous packaging and post-isolation loading techniques like as electroporation and sonication, as detailed in Table 2.
Table 2: Summary of exosome-based CRISPR/Cas9 delivery strategies, detailing loading methods, cargo types, target tissues, and gene-editing outcomes across various therapeutic models.
| Loading strategy | Cargo type | Target tissue/cells | Delivery outcome | Study |
|---|---|---|---|---|
| Electroporation | Cas9 RNP | Liver | Efficient tissue-specific gene editing in liver disease models | [57] |
| Sonication, Extrusion | Cas9 protein, RNP | Fibroblasts | Confirmed loading; future steps to assess editing efficiency | [77] |
| Protein fusion (GFP nanobody/CD63) | sgRNA: Cas9 RNP | A549stop-DsRed cells | Effective target gene knockout through functionalized exosomes | [78] |
| Engineered producer cells | RNP, sgRNA, and ssDNA | Lung | Efficient and targeted gene editing in lung cells in vivo | [79] |
| Endogenous packaging | Plasmid DNA | Ovarian cancer | Tumor-specific gene editing and enhanced chemosensitivity | [80] |
| Transfection | CRISPR/Cas9 plasmid | Pancreatic cancer | Inhibited tumor growth and mutant KRAS targeting | [81] |
| Endogenous exosomal release | Cas9 protein and gRNA | HBV/HPV models | Functional gene editing observed in vitro | [82] |
RNP: Ribonucleoprotein, sgRNA: Single-guide RNA, ssDNA: Single-stranded DNA, gRNA: Guide RNA, GFP: Green fluorescent protein, CD63: Cluster of differentiation 63, HBV: Hepatitis B virus, HPV: Human papillomavirus, KRAS: Kirsten rat sarcoma viral oncogene homolog.
The aggregated results underscore both the practicality and the functional adaptability of exosome-mediated CRISPR/Cas9 delivery methods. Electroporation of Cas9 RNPs has become a dependable technique for hepatic gene editing, providing high efficiency with reduced cytotoxicity [57]. Conversely, mechanical techniques such as sonication and extrusion, although proficient in cargo loading, may jeopardize vesicle integrity, highlighting the necessity to reconcile efficiency with structural preservation [77].
Surface-engineering methodologies, including CD63-fused nanobody systems, illustrate that exosome tropism and intracellular trafficking can be systematically altered to improve editing specificity [78]. Endogenously altered donor cells facilitate precise, cell-type-specific packing of editing complexes, resulting in in vivo functional correction in pulmonary tissue and establishing a scalable, biologically pertinent method [80]. Furthermore, utilizing exosomes produced from tumor cells for plasmid administration demonstrates the potential of leveraging tumor tropism for targeted oncogene disruption [80,83,84].
Collectively, these strategies highlight the versatility of exosomes as gene-editing vectors and stress the necessity of harmonizing cargo type, loading technique, and illness context to enhance therapeutic efficacy.
The effectiveness of delivery relies not only on the technological techniques for cargo loading and exosome engineering but also increasingly on the selection of genetic cargo, which is contingent on upstream genomic analysis [20]. Novel technologies like AI-WES can now detect actionable mutations with remarkable accuracy. These insights immediately inform the therapeutic design process, corresponding with the engineering strategies depicted in Figure 2.
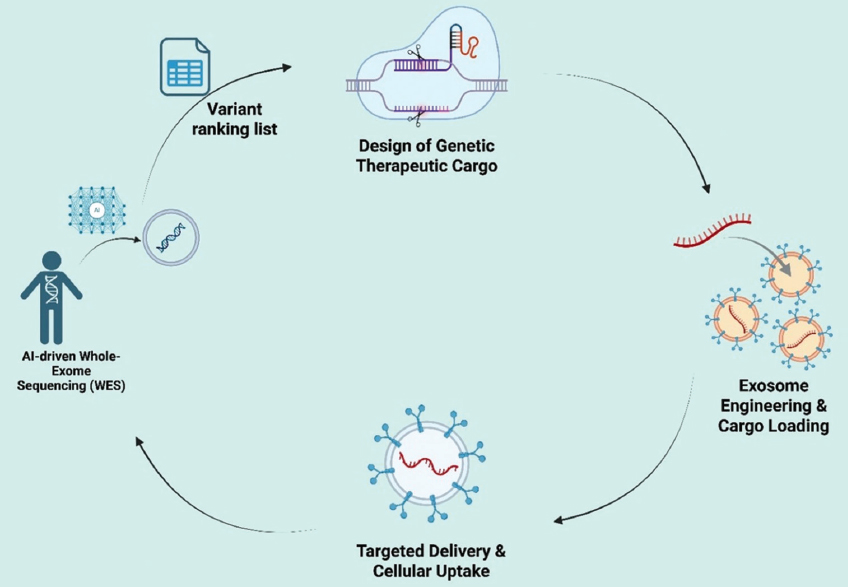 | Figure 2: Artificial intelligence (AI)-driven interpretation of whole-exome sequencing (WES) data for exosome-based gene therapy. AI-driven analysis of WES data identifies and prioritizes actionable variants, guiding the selection of therapeutic cargos (CRISPR/Cas9, small interfering RNA, and messenger RNA). These cargos are subsequently incorporated into exosomes and directed to target cells, enabling personalized gene therapy tailored to patient-specific mutational profiles. [Click here to view] |
5. THERAPEUTIC UTILIZATION OF ENGINEERED EXOSOMES IN GENE THERAPY
Engineered exosomes have garnered considerable attention as advanced vectors for gene therapy, especially in conditions where traditional viral or synthetic carriers encounter safety, immunological, or cargo-size constraints [34]. Their inherent biocompatibility, minimal immunogenicity, and capacity to transport nucleic acids with significant stability have facilitated extensive therapeutic research in oncology, genetic disorders, metabolic diseases, and neurological issues [13,28]. In contrast to synthetic nanoparticles, exosomes have the ability to traverse biological boundaries, demonstrate inherent tissue tropism, and incorporate a variety of genetic materials, including as plasmid DNA, mRNA, siRNA, miRNA, and CRISPR/Cas9 systems [31].
5.1. Gene Therapy in Cancer through Exosome Mediation
Cancer has emerged as the most sophisticated domain for exosome-mediated gene therapy, using exosomes’ capacity to selectively target tumors, transport intricate genetic payloads, and elicit multi-pathway suppression with low immunogenicity [13,34,85]. Tumors inherently secrete exosomes that target primary and metastatic locations, influenced by integrin-mediated organotropism and the metabolic congruence between tumor-derived vesicles and malignant tissue [30]. This intrinsic tropism has motivated engineering approaches that include therapeutic RNA or CRISPR elements into exosomes to inhibit oncogenes, reinstate tumor-suppressor pathways, or enhance tumor sensitivity to chemotherapeutics [35].
The exosomal transport of short RNAs is one of the best-recognized approaches in oncology. Engineered exosomes containing siRNA or miRNA can inhibit mutated or hyperactive oncogenes such as KRAS^G12D, EGFR, BRAF, HER2, and MYC, leading to significant reductions in proliferation, migration, and angiogenesis in colorectal, breast, pancreatic, and lung cancer models [21,33]. Exosomal delivery of miR-34a and miR-143 effectively inhibits anti-apoptotic signaling and reverses epithelial–mesenchymal transition, thereby diminishing metastasis more efficiently than liposomal RNA formulations, which are rapidly degraded and often provoke systemic inflammatory responses [86]. In contrast to synthetic carriers, exosomes protect short RNAs from RNases and promote cytoplasmic release by membrane fusion or endocytosis [29]. These strategies provide prolonged inhibition of carcinogenic pathways with diminished dosages and decreased toxicity.
In addition to silencing, exosomes facilitate the mRNA-mediated re-expression of tumor-suppressive genes. Vesicles transporting mRNA that encodes PTEN, P53, or pro-apoptotic proteins like BAX reinstate functional protein levels in glioblastoma and breast cancer models, resulting in cell-cycle arrest and increased susceptibility to chemotherapy [28]. Exosomal mRNA delivery significantly circumvents the dose-limiting immunogenicity linked to repetitive viral treatment, facilitating sustained expression without provoking neutralizing antibodies. This is especially pertinent for solid tumors necessitating numerous treatment cycles.
The most advanced applications utilize CRISPR/Cas9 systems. The exosomal delivery of CRISPR/Cas9 RNP complexes has effectively edited oncogenic alleles, such as KRAS^G12D in pancreatic cancer and EGFRvIII in glioblastoma, resulting in gene deletion without viral integration and markedly diminished off-target editing [31]. Exosomes can encapsulate big proteins and guide RNAs, so bypassing the cargo-size restrictions of adeno-associated virus (AAV) vectors. Moreover, temporary RNP treatment diminishes prolonged immune activation and alleviates the dangers associated with random genomic insertion.
Exosomes are progressively acknowledged as instruments for immunotherapy. Dendritic cell-derived exosomes that present tumor antigens augment T-cell priming, whereas the exosomal administration of checkpoint-silencing siRNAs (such as those targeting PD-L1) enhances anti-tumor immunity [30]. The integration of exosome-mediated gene therapy with chemotherapy or monoclonal antibodies has yielded synergistic results, exemplified by the co-delivery of doxorubicin-loaded exosomes and miR-21 inhibitors to surmount treatment resistance in triple-negative breast cancer [34]. These combo regimens demonstrate how exosomes might influence tumor genetics while improving responses to current therapies.
In contrast to synthetic nanoparticles, exosomes have enhanced circulation half-life, decreased clearance by macrophages, and improved endosomal escape attributable to their intrinsic membrane composition [26]. Viral vectors are effective gene-transfer technologies; yet, they are constrained by dosage toxicity, immunogenicity, and regulatory limitations. Exosomes offer a non-integrating, repeat-dose-compatible platform, which is essential for the persistent treatment of solid tumors rather than by a singular intervention [87]. Figure 3 illustrates the mechanistic process of tumor targeting, cellular uptake, cargo release, and subsequent reduction of oncogenic signaling through exosome-mediated transport of siRNA, mRNA, and CRISPR.
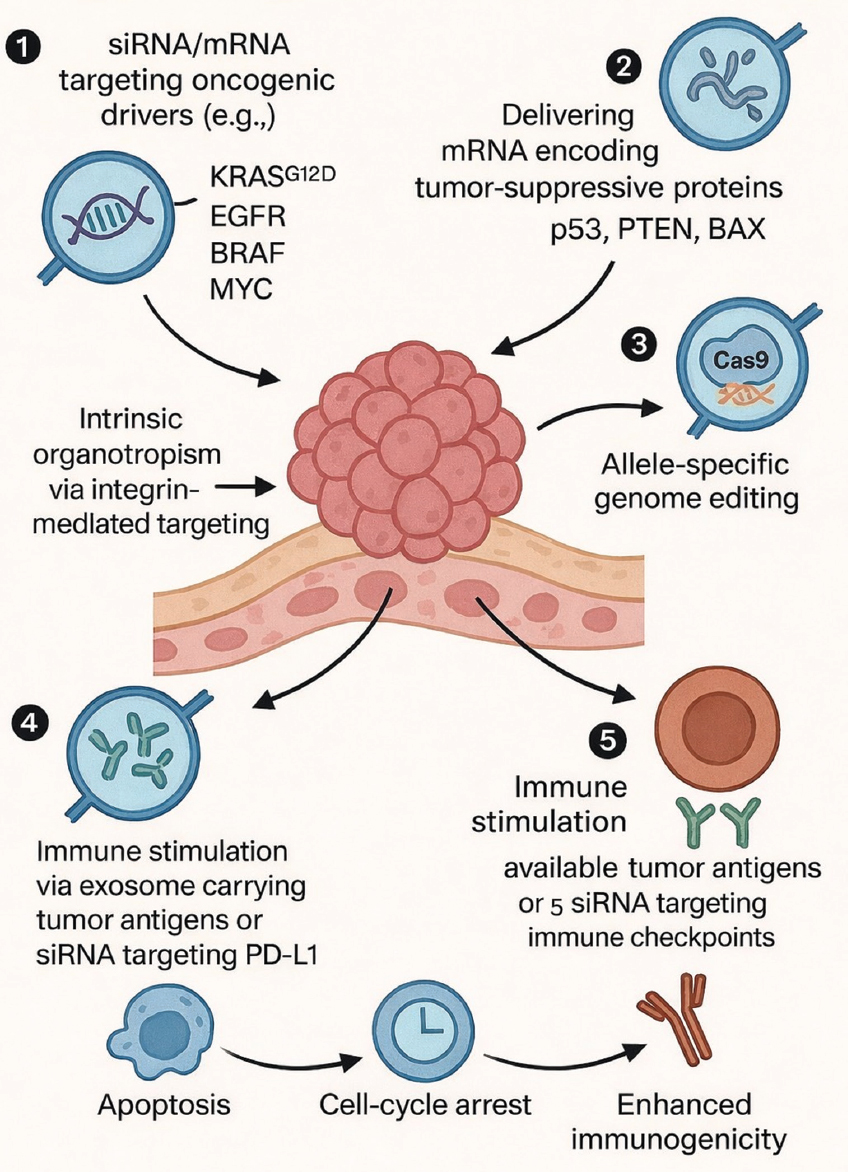 | Figure 3: Therapeutic strategies enabled by engineered exosomes in cancer and immune modulation. (1) Exosomes loaded with small interfering RNA (siRNA) or microRNA targeting oncogenic drivers, including KRAS^G12D, EGFR, BRAF, and MYC. (2) Delivery of mRNA encoding tumor-suppressive proteins such as p53, PTEN, and BAX to restore anti-tumor signaling. (3) CRISPR/Cas9 ribonucleoprotein (RNP) delivery enabling allele-specific or permanent knockout of oncogenic mutations. (4) Immune modulation mediated by exosomes transporting tumor-associated antigens or siRNA targeting immune checkpoints, including PD-L1. (5) Therapeutic outcomes following exosome uptake via tetraspanin- and integrin-mediated organotropism, endocytosis, or membrane fusion, resulting in tumor cell death, reduced proliferation, and enhanced immunological activation. [Click here to view] |
5.2. Exosome-Mediated Gene Therapy for Genetic and Hereditary Disorders
Monogenic disorders are significant targets for exosome-mediated gene therapy, as the restoration of functional protein can frequently be achieved with the delivery of a single mRNA or the induction of a precise gene edit [88]. Conventional viral vectors have demonstrated clinical effectiveness in certain illnesses; however, recurrent administration is constrained by immune responses, cargo size limitations, and insertional mutagenesis. Exosomes circumvent these obstacles due to their inherent biocompatibility, non-integrative delivery mechanism, and ability to transport substantial RNA transcripts and protein–RNA complexes [13,28].
Engineered exosomes have effectively reinstated absent or impaired proteins in various preclinical situations. The administration of mRNA encoding SMN1 in SMA elevates SMN protein levels in motor neurons and restores neuromuscular function, overcoming a constraint of AAV-based SMA therapy where immune responses impede redosing [28]. Exosomal administration of mRNA for CFTR enhances chloride-channel function in cystic fibrosis models, whereas transthyretin (TTR) mRNA delivery reinstates hepatic protein synthesis in transthyretin amyloidosis [21]. Exosomes circumvent neutralizing antibodies, allowing for repeated administration – an advantage in conditions necessitating lifetime treatment.
Certain genetic illnesses are caused not by the absence of protein, but by a deleterious gain-of-function. In Huntington’s disease, exosomes containing siRNA targeting mutant Huntington’s disease (HTT) transcripts diminish aggregation formation and enhance neuronal survival in mouse models [34]. In amyotrophic lateral sclerosis, exosomes transporting siRNA targeting SOD1 diminish motor neuron degeneration and extend survival in preclinical investigations [26]. These findings illustrate that exosome-mediated silencing can specifically suppress pathogenic alleles while preserving the healthy gene variant – an achievement challenging to realize with numerous non-viral nanoparticles.
Exosome-mediated CRISPR delivery is an emerging area in the treatment of genetic disorders. Exosomes can transport substantial Cas9 proteins and guide RNAs, so circumventing AAV’s size limitations and preventing enduring nuclease expression. In Duchenne muscular dystrophy models, the administration of Cas9 RNPs reinstates dystrophin expression in skeletal muscle without observable off-target modifications [31]. Recent investigations have employed analogous techniques for hemoglobinopathies by altering HBB (hemoglobin subunit beta) and reactivating fetal hemoglobin pathways, underscoring a potential avenue for curative therapy devoid of viral integration [89].
An important benefit of exosomes is their capacity to access areas that are unreachable by viral vectors or synthetic particles. Exosomes generated from macrophages preferentially aggregate in the liver and spleen, fibroblast exosomes target connective tissues, and neural exosomes traverse the BBB, facilitating the therapy of central nervous system illnesses [13]. Surface functionalization using ligands such as RVG (rabies virus glycoprotein) peptides or heparin-binding motifs improves tissue selectivity and minimizes systemic exposure [29]. These tropism characteristics facilitate focused distribution while reducing toxicity.
Despite still in the preclinical phase, the growing body of evidence outlines a feasible translational pathway for exosome-mediated gene repair. This pathway involves the judicious selection of therapeutic mRNA or CRISPR-based agents, strategic engineering of donor cells, targeted alteration of exosomal surface characteristics, scalable good manufacturing practice (GMP)-compliant production, and long-term assessment of protein restoration or mutation correction in circulation [28,36]. As manufacturing frameworks and regulatory standards evolve, exosomes are becoming a promising platform for curative, redosable gene therapies, especially for inherited illnesses resistant to viral delivery methods [90]. The mechanisms – mRNA replacement, allele-specific silencing, CRISPR-based correction, and tissue-targeted biodistribution – are illustrated schematically in Figure 4.
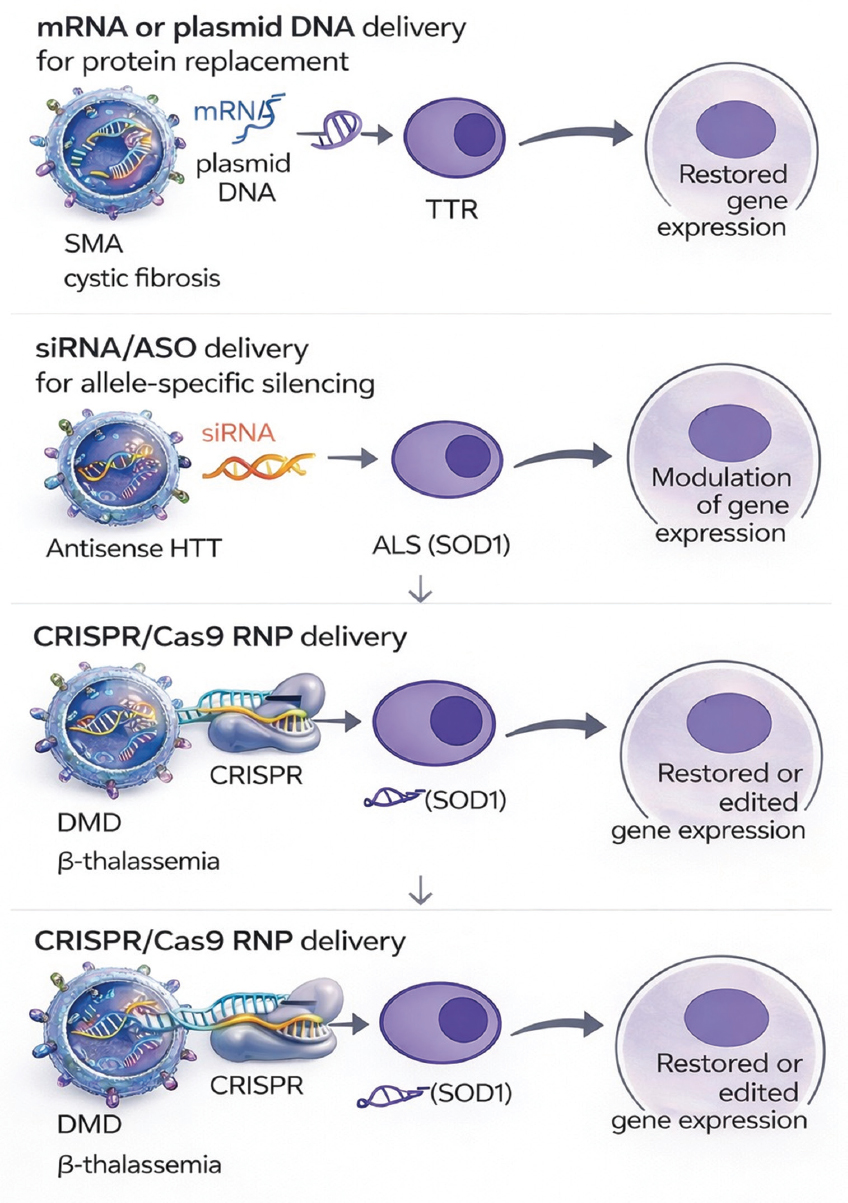 | Figure 4: Exosome-based genetic correction strategies for inherited diseases. The diagram illustrates the exosomal transport of mRNA or plasmid DNA for protein substitution in spinal muscular atrophy (SMN1), cystic fibrosis, and transthyretin amyloidosis. A second panel depicts allele-specific silencing with small interfering RNA/ASO for gain-of-function mutations in Huntington’s disease and amyotrophic lateral sclerosis (SOD1). A third panel illustrates the delivery of CRISPR/Cas9 RNP for dystrophin restoration in Duchenne muscular dystrophy or HBB editing in β-thalassemia. Exosomes safeguard nucleic acids from destruction, infiltrate target cells, discharge cargo, and restore or regulate gene expression without viral integration. [Click here to view] |
5.3. Gene Therapy Utilizing Exosomes for Neurological Disorders
Neurological disorders provide a significant obstacle for gene therapy due to the blood–BBB, which substantially restricts the administration of viral vectors and synthetic nanoparticles. Exosomes possess a distinct advantage in this situation since they can inherently traverse the BBB and transport functional genetic material to neurons, astrocytes, and microglia with low systemic toxicity [13,29]. Their intrinsic membrane composition and tetraspanin-facilitated uptake enhance effective endocytosis and cytoplasmic release of mRNA, siRNA, and CRISPR components, surmounting a major obstacle to central nervous system (CNS)-targeted therapy [32].
Numerous neurodegenerative diseases are caused by deleterious gain-of-function mutations. Exosomes with siRNA targeting mutant HTT transcripts have diminished huntingtin protein aggregation and enhanced neuronal survival in Huntington’s disease models, providing selective allele suppression without irreversible genome alteration [34]. In amyotrophic lateral sclerosis, exosomal administration of siRNA directed at SOD1 mitigates motor neuron degradation and prolongs survival in preclinical investigations [26]. Comparable methodologies have been investigated for the suppression of tau-associated transcripts in Alzheimer’s disease to mitigate neurofibrillary damage.
Some brain illnesses are defined by the depletion of vital proteins instead of the buildup of harmful species. Exosome-mediated delivery of SMN1 mRNA has been demonstrated to restore SMN protein expression in motor neurons and improve neuromuscular function in SMA [91]. These findings suggest a potentially reusable therapeutic strategy that overcomes the constraints of AAV-based methods hindered by anti-capsid immunity [28]. mRNA delivery has demonstrated potential for lysosomal storage disorders, wherein exosomes transporting enzyme-encoding transcripts rectify metabolic deficiencies in brain cells.
The exosomal delivery of CRISPR/Cas9 RNP complexes facilitates accurate genome editing in brain tissues, while mitigating the prolonged nuclease expression linked to viral vectors. Preliminary research has shown effective gene deletion in neuronal populations with minimized off-target effects and, importantly, no risk of viral genome integration [31]. This strategy is currently under investigation for various prevalent neurological disorders, including genetic epilepsies, LRRK2-associated Parkinson’s disease, and mutations linked to ALS [92].
Exosomes have been modified with targeting peptides like RVG to improve neuronal absorption by binding to nicotinic acetylcholine receptors on neuronal cells [29]. Alternative investigations employ Lamp2b-fusion constructs or ApoA-I motifs to enhance BBB penetration and cerebral accumulation. These technologies markedly enhance the delivery of therapeutic cargo while minimizing systemic exposure. Collectively, exosomes offer a biocompatible, redosable, and barrier-penetrating platform for genetic intervention in neurological diseases. Their capacity to safeguard nucleic acids, transport substantial genetic payloads, and penetrate deep brain areas establishes them as a next-generation non-viral instrument for addressing disorders previously deemed unreachable by gene therapy [93]. Figure 5 illustrates the mechanism through which modified exosomes traverse the BBB, convey genetic material, and rehabilitate neuronal function.
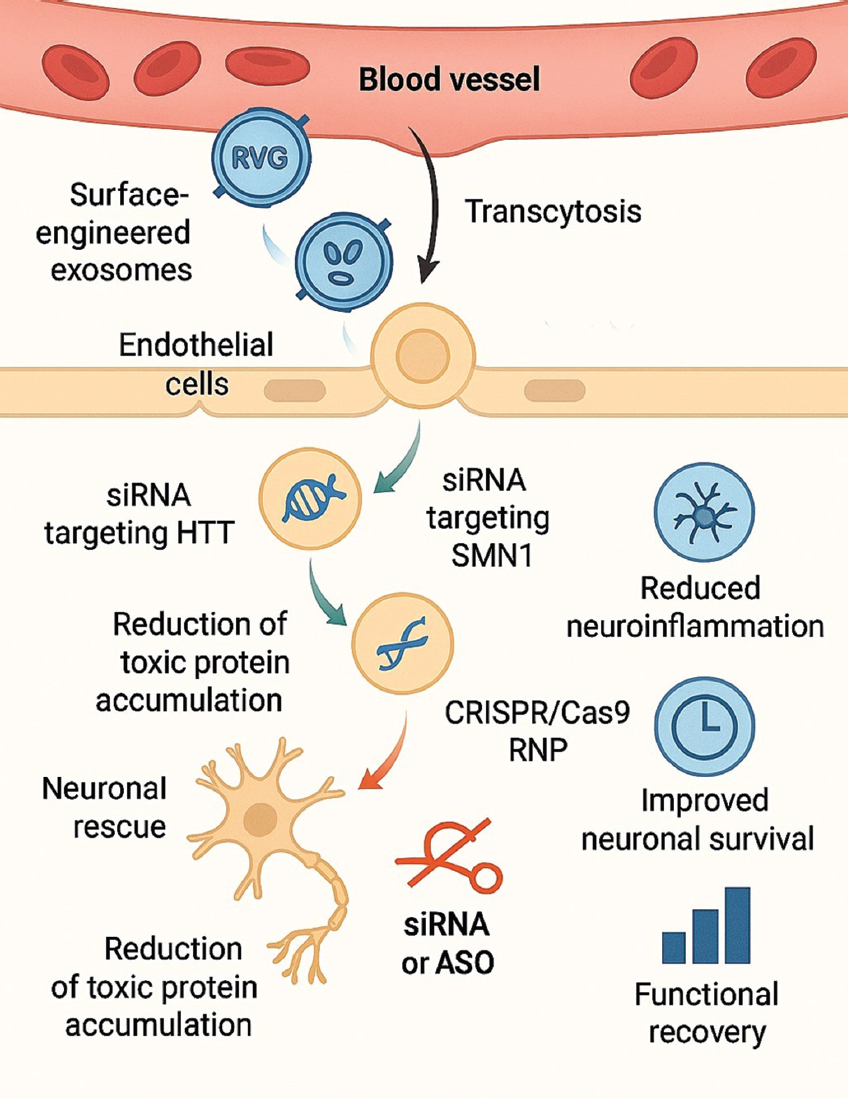 | Figure 5: Mechanism of exosome-mediated gene delivery across the blood–brain barrier (BBB). The figure demonstrates surface-engineered exosomes displaying RVG or Lamp2b targeting peptides, enabling interaction with BBB endothelial cells and transcytosis into the CNS. Once in neural tissue, exosomes release siRNA targeting Huntington’s disease or SOD1, or messenger RNA encoding SMN1, leading to neuronal rescue and reduction of toxic protein accumulation. CRISPR/Cas9 RNP delivery for mutation correction is also shown. Outcome elements include reduced neuroinflammation, improved neuronal survival, and functional recovery. [Click here to view] |
5.4. Addressing Genetic Disorders through Exosome-Mediated Gene Therapy: A Translational Framework
The therapeutic promise of exosomes is especially significant in genetic disorders, where accurate release of corrected nucleic acids can restore gene activity or inhibit deleterious alleles. In contrast to viral vectors, exosomes can transport large mRNAs, plasmid DNA, or CRISPR RNP complexes without limitations on cargo size or the hazards associated with genomic integration [28,31]. Their inherent biocompatibility and minimal immunogenicity facilitate repeated administration, which is crucial for chronic genetic conditions. A defined developmental roadmap is emerging to facilitate the clinical translation of exosome-based genetic treatments.
Step 1 – Selection of therapeutic cargo: The therapeutic cargo must be chosen based on the illness mechanism, the necessary duration of effect, and the intracellular target site. In loss-of-function scenarios, gene augmentation through mRNA or plasmid DNA can reinstate protein expression (e.g., SMN1, CFTR, and TTR), while harmful gain-of-function illnesses are typically addressed through transcript suppression utilizing siRNA or antisense oligonucleotides (e.g., HTT and SOD1). For mutations necessitating robust sequence correction, CRISPR/Cas9 RNPs or editing platforms (e.g., for HBB or DMD) may be contemplated [34]. Loading can be accomplished through electroporation, membrane permeabilization, or donor-cell engineering; however, due to the dependence of loading efficiency and vesicle integrity on the method and cargo, the selection of cargo should be aligned with the characterization of encapsulation, recovery, and functional potency in pertinent models [26].
Step 2 – Engineering exosomes for tissue tropism: While native vesicles exhibit certain targeting capabilities, modified exosomes enhance precision. Lamp2b-fused ligands, including RVG peptides, guide vesicles to neurons, RGD (arginine–glycine–aspartic acid) and integrin-binding motifs guide tumors and vasculature, while ApoA-I facilitates hepatic transport [29,32]. These alterations enhance on-target accumulation while diminishing systemic exposure – essential for gene-editing tools with significant biological potency.
Step 3 – Biodistribution and pharmacodynamics: Preclinical evaluation entails the real-time monitoring of tagged exosomes by fluorescence or bioluminescence techniques. Research indicates that modified vesicles can traverse the BBB, aggregate in motor neurons, hepatocytes, and cardiomyocytes, and release functional nucleic acids without eliciting immune activation [13]. In SMA and CFTR models, mRNA-loaded exosomes reinstate protein expression and normal cellular function, whereas CRISPR-loaded vesicles accomplish allele deletion in dystrophic muscle without viral integration [31].
Step 4 – Evaluation of efficacy and safety: The transient characteristics of exosomal mRNA and CRISPR RNPs mitigate the hazards of off-target genomic alterations and insertional mutagenesis, rendering them appealing for pediatric and long-term therapeutic applications [28]. Toxicology tests indicate negligible cytokine release and minor complement activation, in contrast to immunological responses elicited by AAV or lipid nanoparticles. The absence of powerful promoters or viral capsids in exosomes enhances long-term safety.
Step 5 – GMP manufacturing and scalability: Scalable production continues to be a significant constraint. Present methodologies encompass the bioreactor amplification of donor cells, purification through size-exclusion chromatography or tangential flow filtration, and quality evaluation using nanoparticle tracking and proteomic analysis [30]. Regulatory frameworks are increasingly acknowledging exosomes as unique biological treatments, facilitating more defined clinical paths.
Exosome-based delivery platforms provide a viable translational pathway for treating genetic disorders. Their ability to carry substantial and structurally intricate genetic materials, navigate biological barriers, and allow for repeated delivery without dose-limiting immunogenicity establishes them as a next-generation method for genome correction [94]. Continuous improvements in cargo-loading efficiency, tissue-specific targeting, and GMP-compliant production will influence the speed at which these methodologies progress into clinical evaluation and, ultimately, patient application [95].
Figure 6 illustrates the complete translational pathway, encompassing therapeutic cargo design, GMP manufacture, and preclinical validation.
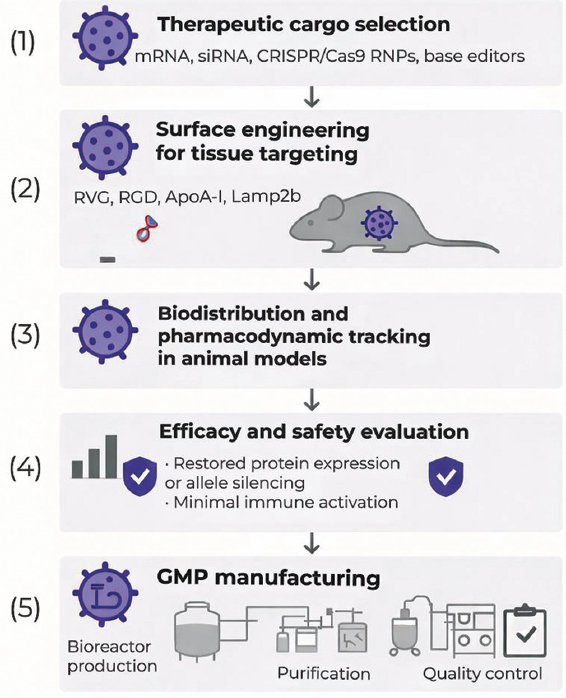 | Figure 6: Translational roadmap for exosome-mediated gene therapy in genetic diseases. The flowchart illustrates five development stages: (1) Therapeutic cargo selection (messenger RNA, small interfering RNA, CRISPR/Cas9 RNPs, and base editors); (2) surface engineering for tissue targeting (RVG, RGD, ApoA-I, and Lamp2b fusion); (3) biodistribution and pharmacodynamic tracking in animal models; (4) efficacy and safety evaluation showing restored protein expression or allele silencing with minimal immune activation; and (5) GMP manufacturing pipeline including bioreactor production, purification, and quality control. The figure emphasizes redosability, absence of genomic integration, and suitability for clinical translation. [Click here to view] |
5.5. Exosome Design Guided by AI-WES: A Pragmatic Workflow
AI-assisted analysis of WES can serve as a preliminary design engine for exosome gene treatments by transforming patient variants into precise delivery parameters. Initially, AI-assisted variant prioritization determines actionable targets and specifies whether the treatment objective is gene restoration (loss-of-function), allele silencing (gain-of-function), or sequence rectification (pathogenic variations amenable to editing) [96]. The chosen modality (mRNA/siRNA/antisense oligonucleotides (ASOs), CRISPR RNP, and base editing) imposes limitations on cargo size, necessary subcellular localization, and permissible exposure period [97]. Third, these limits inform exosome engineering decisions, encompassing donor-cell origin, loading technique (passive or active), and surface modification to mitigate predominant biodistribution sinks (e.g., liver/spleen sequestration) and enhance tropism [98]. Fourth, translation necessitates consistent characterization and potency assays linked to mechanisms (e.g., editing rate, knockdown efficiency, and protein restoration) along with a GMP-compliant manufacturing strategy that includes batch comparability requirements [99]. Ultimately, results from biodistribution, efficacy, and safety assessments should inform model enhancement, facilitating the continual advancement of both AI priority and exosome design criteria.
Positioning concerning recent reviews (2024–2025): Recent reviews have comprehensively documented exosome-mediated CRISPR/Cas delivery methodologies and their oncological applications [55,56], while concurrent reviews highlight how AI/machine learning (ML) can expedite carrier design, forecasting, and manufacturing preparedness [100-102]. Nonetheless, these strands predominantly advance concurrently – exosome–CRISPR reviews are generally focused on methodologies (loading modality, targeting ligands, and proof-of-concept editing), while AI-in-nanomedicine reviews frequently enhance physicochemical parameters without directly associating them with patient-specific pathogenic variants or clinical sequencing processes. This review links AI-assisted WES interpretation to exosome engineering decisions through an actionable decision matrix [Table 3] and presents translational failure modes (potency assays, batch variability, safety, and regulatory classification) as variables that can be modeled and mitigated within a closed-loop design–build–test–learn pipeline.
Table 3: AI–WES-guided exosome design decision matrix.
| Stage | Concrete output | AI/WES+data inputs | Exosome design levers | Mechanism-linkedassays /quality control (QC) | Common failure modes+mitigations |
|---|---|---|---|---|---|
| 0. Genomic interpretation | Ranked actionable variants+target hypothesis | WES, phenotype (Human Phenotype Ontology, HPO), family history; variant effect predictors; and literature/knowledge graphs | Defines whether therapy needs restoration, silencing, or correction | Orthogonal confirmation (Sanger/targeted NGS); expression evidence where relevant | Mis-prioritization from biased training data→transparent criteria+clinician review |
| 1. Modality selection | Chosen cargo type (mRNA/siRNA/CRISPR RNP/base editor) + dosing concept | Predicted loss-of-function/gain-of-function (LoF/GoF) expression; editability; and off-target risk models | Cargo size constraints; cytosolic versus nuclear delivery; and duration of action | In vitro target engagement (knockdown/editing/protein restoration); and off-target screens | Over-engineering (plasmid) when transient is safer→prefer RNP/mRNA; predefine go/no-go |
| 2. Donor cell/source choice | Source selected with safety rationale | Target tissue; immunogenicity risk; scalability; and producer line datasets | mesenchymal stem cell (MSC)/immune/cell-line EVs; and avoid tumor-derived sources for therapy | Minimal information for studies of extracellular vesicles-aligned markers; sterility/mycoplasma; residual DNA/RNA profiling | Oncogenic/pro-inflammatory cargo carryover→source qualification+cargo profiling |
| 3. Loading strategy | Loading method+expected loading metrics | Cargo physicochemical properties; stability; and desired copy number | Passive incubation; electroporation; transfection; microfluidics; and hybrids | Loading efficiency; nanoparticle tracking analysis/electron microscopy (NTA/EM); free-cargo removal; RNase/protease protection | Aggregation/membrane damage→optimize buffers; include free-cargo and aggregation controls |
| 4. Targeting/tropism engineering | Surface ligand plan+route of administration | Biodistribution models; receptor expression maps; and prior in vivo datasets | Display peptides/antibodies; glycoengineering; and route (IV/local) | In vivo biodistribution (barcoding/label-free); target: off-target ratios; functional potency | Liver/spleen sequestration→route+stealth; validate with label-free methods |
| 5. Manufacturing +release | GMP-compatible process+release criteria | Process analytics; batch records; and comparability data | Bioreactor expansion; size-exclusion chromatography/tangential flow filtration (SEC/TFF)-based purification; and closed-system steps | Identity/purity; potency assay; residual host DNA; endotoxin; stability | Batch variability→mechanism-linked potency assay+predefined acceptance ranges |
| 6. Feedback learning loop | Updated models+design rules | All outcomes (including negatives); pharmacovigilance | Iterative refinement of source/loading/targeting | Prospective validation; external test sets; and dataset/model documentation | Overfitting+leakage→strict governance; report negative outcomes where possible |
WES: Whole-exome sequencing, AI: Artificial intelligence, mRNA: Messenger RNA, siRNA: small interfering RNA, RNP: Ribonucleoprotein, EVs: Extracellular vesicles.
6. CHALLENGES, LIMITATIONS, AND FUTURE DIRECTIONS
Despite the rapidly growing interest in exosome-mediated gene therapy, substantial scientific, engineering, and regulatory challenges persist, hindering extensive clinical implementation. The limits do not diminish the field’s potential; rather, they create a framework of actionable priorities for advancing exosomes from experimental systems to therapeutically dependable gene-delivery platforms.
6.1. Standardization, Nomenclature, and Reporting (MISEV)
In the EV domain, uneven language and inadequate reporting of pre-analytical factors, isolation techniques, and marker panels are significant contributors to irreproducibility. The ISEV advocates for the use of “EV” as the generic designation unless endosomal origin is substantiated, and it delineates minimum reporting standards (MISEV2018 and the expanded MISEV2023 update) pertaining to sample handling, separation, characterization, and functional assertions. In translational research, adhering to these checklists for experimental design and reporting, which encompass depleted/negative indicators and, when possible, protein topology, aids in differentiating vesicle-associated activity from co-isolated contaminants and enhances batch comparability and release criteria.
6.2. Isolation, Batch Variability, and Mechanism-Associated Potency Assays
A significant challenge is standardized large-scale biomanufacturing. Exosomes are released in limited quantities, and their composition is significantly affected by the kind of donor cell, passage history, oxygen levels, food availability, and pathological conditions [28,31,34]. Minor fluctuations in cell growth conditions can modify vesicle dimensions, protein composition, RNA content, and immunological characteristics. Despite advancements in bioreactor-based expansion, tangential flow filtering, and ultracentrifugation that have enhanced yields, a widely acknowledged GMP-compliant manufacturing workflow remains absent [104]. The lack of validated potency assays, release criteria, and purity thresholds hinders regulatory approval and adds batch-to-batch variability, a worry less prevalent with synthetic nanoparticles or viral vectors [29]. Future endeavors should emphasize regulated donor-cell modification, enclosed-system manufacturing, and instantaneous vesicle analysis to guarantee consistent therapeutic efficacy.
6.3. Efficiency in Cargo Loading and Meticulous Characterization
A pertinent constraint is the inefficient and diverse loading of cargo, especially for substantial genetic constructs such as plasmids, CRISPR/Cas9 RNPs, and base editors. Passive loading utilizes natural biogenesis pathways, yet results in inconsistent encapsulation efficiency. Active loading methods – such as electroporation, sonication, extrusion, and lipid fusion – enhance cargo integration but may compromise membrane integrity or denature proteins [26,27]. Molecular techniques employing RNA-binding domains (e.g., MS2, HuR, and zipcode elements) provide enhanced selectivity but necessitate genetic alteration of donor cells and introduce regulatory complexities. The subsequent stage of exosome engineering must integrate the rational design of loading motifs, membrane-stabilization chemistries, and standardized measurement techniques for encapsulated nucleic acids [105].
6.4. Limitations in Biodistribution and Targeting
Biodistribution and targeted specificity continue to be significant translational obstacles. On systemic administration, native exosomes are preferentially sequestered by the liver, spleen, and mononuclear phagocyte system [13,32]. This rapid elimination restricts therapeutic buildup in affected tissues. Surface-engineering techniques, such as Lamp2b–RVG fusions for neuronal uptake, RGD peptides for tumor integrins, and ApoA-I motifs for endothelial transport, can improve tissue tropism in preclinical models [106]. Nonetheless, their long-term safety, immunogenicity, and metabolic stability remain ambiguous. Biodistribution varies significantly among species, rendering rat pharmacokinetics an inadequate predictor of human results. Advancements in this domain will rely on quantitative whole-body imaging, mechanistic modeling of vesicle trafficking, and the development of receptors specifically designed for human biology [107]. Key characteristics, advantages, and limitations of extracellular vesicle–based delivery strategies are summarized in Table 4.
Table 4: How this review extends recent 2024–2025 reviews relevant to exosome–CRISPR delivery and AI in nanomedicine
| Recent review (year) | Primary scope | What it establishes well | Key gap for AI–WES-guided exosome design | Extension provided here |
|---|---|---|---|---|
| Guo et al. [7] | Viral versus non-viral nanocarriers for in vivo CRISPR editing | Comparative lens across delivery vectors and editing modalities | Exosomes treated as one option; no variant-to-cargo mapping or EV-specific process controls | Deep focus on exosome engineering coupled to genomic target selection and EV QC variables |
| Serrano et al. [15] | Exosome-based drug delivery platforms | Surface engineering and targeting strategies across diseases | Limited attention to gene editing cargo logic and clinical genomics inputs | Integrates gene-editing payload selection with AI-assisted WES and a design decision matrix |
| Balaraman et al. [56] | Exosome-mediated CRISPR-Cas9 for cancer gene editing (review) | Catalogs exosome targeting, tumor microenvironment barriers, and preclinical evidence | Does not formalize AI-enabled, patient-specific variant prioritization or model-driven design loops | Proposes an AI–WES-to-exosome workflow and highlights data/assay requirements for iterative optimization |
| Agrahari et al. [102] | AI/ML to accelerate nanomedicine discovery, manufacturing and translation (perspective) | Maps where AI/ML can reduce trial-and-error and support scalable manufacturing | Not specific to EV biology; lacks EV nomenclature/heterogeneity and gene-therapy constraints | Adapts the AI tool chain to EV datasets, heterogeneity, and gene-editing safety/efficacy endpoints |
| Azimi[100] and Chou et al. [101] | AI-driven nanoparticle design and prediction (reviews) | Highlights data-driven design, interpretability issues, and translational barriers for nanoformulations | Does not address EV isolation variability, tropism, or clinical sequencing integration | Extends AI concepts to EV-specific sources of variance and to genomics-informed payload decisions |
WES: Whole-exome sequencing, AI: Artificial intelligence, EVs: Extracellular vesicles.
6.5. Safety, Failure Mechanisms, and Contradictory Evidence
Safety continues to be a paramount concern in clinical translation. While exosomes are often less immunogenic than viral vectors, vesicles originating from tumor cells or genetically altered donor lines may include oncogenic proteins, mutant nucleic acids, or immunosuppressive agents [30]. Exosomes may affect gene expression in non-target tissues, prompting worries about off-target silencing, unintentional epigenetic alterations, or horizontal RNA transfer. Comprehensive toxicological and biodistribution studies are essential to assess these concerns, especially for CRISPR-based therapies, as permanent genome alterations may result in delayed detrimental effects. The creation of entirely synthetic or semi-synthetic exosome-mimetic systems, produced through the bottom-up assembly of lipid membranes and specified cargo, may address biological heterogeneity while preserving biocompatibility [108].
An equitable evaluation must also recognize persistent failure mechanisms. Numerous studies indicate low and inconsistent loading of substantial or multicomponent cargos, such as CRISPR RNPs and base editors [109]. Aggressive loading may compromise vesicle integrity. Following systemic administration, vesicles are frequently swiftly absorbed by the mononuclear phagocyte system. Targeting motifs exhibit restricted predictability among species [110].
Safety considerations encompass more than just immediate tolerance. Extracellular vesicles may convey unexpected bioactive nucleic acids or proteins. Extracellular vesicles produced from tumors may convey pro-tumorigenic signals. Repeated administration can influence immunological responses [111]. Genome editing presents supplementary dangers, including off-target effects. Transparent reporting of adverse or contradictory data, along with uniform characterization, is essential to differentiate platform constraints from modifiable engineering variables [112].
6.6. Regulatory Categorization and Clinical Scalability
Regulatory impediments continue to pose a significant challenge. Regulatory agencies vary in their classification of modified exosomes; some categorize them as biological treatments, while others describe them as advanced therapy medicinal goods, drug–device combos, or gene therapy vectors. The absence of alignment hampers clinical trial preparation and introduces uncertainty to manufacturing approval [113]. The European Commission’s 2023 revision of its GTMPs framework is a positive advancement; yet, really harmonized international norms remain absent. Comprehensive standards for vesicle definition, release testing, potency assays, sterility, stability, and in vivo tracking remain inadequately defined. Without cohesive regulatory frameworks, industry investment and the trajectory toward clinical translation are expected to advance more slowly than the scientific developments themselves [114].
6.7. Areas of AI Assistance and Potential Misinformation
Machine learning is progressively employed to amalgamate multi-omic EV characterization, biodistribution profiles, and functional readouts into prediction models. These models may evaluate design aspects such as donor cell source, loading method, and targeting ligands [115]. They can also facilitate the formulation of mechanism-associated potency assays. Nonetheless, AI does not supplant stringent experimental control. Model efficacy must be assessed prospectively using independent datasets. Training data must be verifiable against standardized electric vehicle preparations and reporting frameworks. Algorithmic bias, such as the over-representation of particular cell lines or animal models, must be assessed and mitigated to curtail excessive translational assertions [116].
Notwithstanding these obstacles, the discipline is progressing toward definitive solutions. Innovative technologies – such as DNA barcoding for real-time monitoring, microfluidic purification, CRISPR-compatible exosome scaffolds, machine-learning-enhanced cargo selection, and WES-informed mutation prioritization – are facilitating a more systematic, disease-targeted design of exosome therapeutics [19]. Exosomes are positioned as a transformational platform for next-generation gene therapy due to their capacity to deliver substantial gene-editing complexes without genomic integration, traverse biological barriers including the BBB, and be safely re-administered. The ongoing combination of synthetic biology, biomaterials engineering, and computational genomics is expected to characterize the initial phase of clinically authorized exosome gene therapies in the forthcoming decade.
7. CONCLUSION
Engineered EVs, including exosomes when their endosomal origin is confirmed, are developing as a supplementary delivery system for gene and genome-editing therapies, with possible benefits in biocompatibility, diminished immunogenicity, and re-administrability. Simultaneously, translation will rely on addressing particular bottlenecks instead of extrapolating from preclinical potential.
Significant research deficiencies encompass: (i) A standardized nomenclature for EVs, consistent isolation reporting, and characterization in accordance with MISEV guidelines; (ii) reproducible and quantifiable metrics for loading and release of large or multicomponent cargos (e.g., plasmids, CRISPR RNPs, and base editors); (iii) predictive biodistribution and targeting methodologies that are applicable across various models; (iv) validated potency assays linked to mechanisms and release criteria that ensure batch comparability; (v) closed-system, GMP-compliant manufacturing processes featuring scalable purification and quality control; and (vi) long-term safety frameworks that consider immune responses, unintended cargo transfer, and off-target risks specific to editing.
The primary opportunity at the AI-exosome interface is to transform patient genomes into technical specifications. AI-assisted WES variant prioritization can guide target selection and modality determination, while data-driven models can enhance exosome design parameters and manufacturing quality characteristics. Prospective validation and transparent reporting are crucial for ensuring that AI serves as a dependable decision-support system rather than exacerbating dataset bias.
8. CONTRIBUTIONS OF THE AUTHORS
All authors made substantial contributions to conception and design, acquisition of data, or analysis and interpretation of data; took part in drafting the article or revising it critically for important intellectual content; agreed to submit to the current journal; gave final approval of the version to be published; and agree to be accountable for all aspects of the work. All the authors are eligible to be author as per the International Committee of Medical Journal Editors (ICMJE) requirements/guidelines.
9. FUNDING
There is no funding to report.
10. CONFLICTS OF INTEREST
The authors report no financial or any other conflicts of interest in this work.
11. ETHICAL APPROVALS
This study does not involve experiments on animals or human participants.
12. DATA AVAILABILITY
All the data is available with the authors and shall be provided upon request.
13. PUBLISHER’S NOTE
All claims expressed in this article are solely those of the authors and do not necessarily represent those of the publisher, the editors, or the reviewers. The journal remains neutral with regard to jurisdictional claims in published institutional affiliations.
14. USE OF ARTIFICIAL INTELLIGENCE (AI)-ASSISTED TECHNOLOGY
The authors declare that they have not used artificial intelligence (AI)-tools for writing and editing of the manuscript, and no images were manipulated using AI.
REFERENCES
1. Li H, Yang Y, Hong W, Huang M, Wu M, Zhao X. Applications of genome editing technology in the targeted therapy of human diseases:Mechanisms, advances and prospects. Signal Transduct Target Ther. 2020;5(1):1. [CrossRef]
2. Muul LM, Tuschong LM, Soenen SL, Jagadeesh GJ, Ramsey WJ, Long Z, et al. Persistence and expression of the adenosine deaminase gene for 12 years and immune reaction to gene transfer components:Long-term results of the first clinical gene therapy trial. Blood. 2003;101(7):2563-9. [CrossRef]
3. von Fritschen M, Janosz E, Blume C, Jägle U, Keating K, Schneider CK. What's in a word?Defining “gene therapy medicines”. Mol Ther Methods Clin Dev. 2024;32(4):101348. [CrossRef]
4. Watson-Levings RS, Palmer GD, Levings PP, Dacanay EA, Evans CH, Ghivizzani SC. Gene therapy in orthopaedics:Progress and challenges in pre-clinical development and translation. Front Bioeng Biotechnol. 2022;10:901317. [CrossRef]
5. Taha EA, Lee J, Hotta A. Delivery of CRISPR-Cas tools for in vivo genome editing therapy:Trends and challenges. J Control Release. 2022;342:345-61. [CrossRef]
6. Al-Dosari MS, Gao X. Nonviral gene delivery:Principle, limitations, and recent progress. AAPS J. 2009;11:671-81. [CrossRef]
7. Guo Z, Zhu AT, Fang RH, Zhang L. Viral and nonviral nanocarriers for in vivo CRISPR-based gene editing. Nano Res. 2024;17(10): 8904-25. [CrossRef]
8. Wang Y, Bui TA, Yang X, Hutvagner G, Deng W. Advancements in gene therapies targeting mutant KRAS in cancers. Cancer Metastasis Rev. 2025;44(1):24. [CrossRef]
9. Yang L, Li Z, Binzel DW, Guo P, Williams TM. Targeting oncogenic KRAS in non-small cell lung cancer with EGFR aptamer-conjugated multifunctional RNA nanoparticles. Mol Ther Nucleic Acids. 2023;33:559-71. [CrossRef]
10. Sayed S, Sidorova OA, Hennig A, Augsburg M, Cortes Vesga CP, Abohawya M, et al. Efficient correction of oncogenic KRAS and TP53 mutations through CRISPR base editing. Cancer Res. 2022;82(17):3002-15. [CrossRef]
11. Haque US, Yokota T. Gene editing for duchenne muscular dystrophy: From experimental models to emerging therapies. Degener Neurol Neuromuscul Dis. 2025;15:17-40. [CrossRef]
12. Jiao Y, Yang L, Wang R, Song G, Fu J, Wang J, et al. Drug delivery across the blood-brain barrier:A new strategy for the treatment of neurological diseases. Pharmaceutics. 2024;16(12):1611. [CrossRef]
13. Chen YF, Luh F, Ho YS, Yen Y. Exosomes:A review of biologic function, diagnostic and targeted therapy applications, and clinical trials. J Biomed Sci. 2024;31(1):67. [CrossRef]
14. Sharma V, Mukhopadhyay CD. Exosome as drug delivery system: Current advancements. Extracellular Vesicle. 2024;3:100032. [CrossRef]
15. Serrano DR, Juste F, Anaya BJ, Ramirez BI, Sanchez-Guirales SA, Quispillo JM, et al. Exosome-based drug delivery:A next-generation platform for cancer, infection, neurological and immunological diseases, gene therapy and regenerative medicine. Pharmaceutics. 2025;17(10):1336. [CrossRef]
16. Mehdizadeh S, Mamaghani M, Hassanikia S, Pilehvar Y, Ertas YN. Exosome-powered neuropharmaceutics: Unlocking the blood-brain barrier for next-gen therapies. J Nanobiotechnology. 2025;23(1):329. [CrossRef]
17. Lu Y, Huang W, Li M, Zheng A. Exosome-based carrier for RNA delivery: Progress and challenges. Pharmaceutics. 2023;15(2):598. [CrossRef]
18. Van Hoecke L, Lucci C, Vandenbroucke RE. Mesenchymal stromal cell extracellular vesicles as immune modulators and drug carriers in neurodegenerative disorders. Trends Neurosci. 2025;48(11):919-34. [CrossRef]
19. O'Brien TD, Campbell NE, Potter AB, Letaw JH, Kulkarni A, Richards CS. Artificial intelligence (AI)-assisted exome reanalysis greatly aids in the identification of new positive cases and reduces analysis time in a clinical diagnostic laboratory. Genet Med. 2022;24(1):192-200. [CrossRef]
20. Bahadorani M, Nasiri M, Dellinger K, Aravamudhan S, Zadegan R. Engineering exosomes for therapeutic applications: Decoding biogenesis, content modification, and cargo loading strategies. Int J Nanomed. 2024;19:7137-64. [CrossRef]
21. Lee YJ, Shin KJ, Chae YC. Regulation of cargo selection in exosome biogenesis and its biomedical applications in cancer. Exp Mol Med. 2024;56(4):877-89. [CrossRef]
22. Srivastava A, Rathore S, Munshi A, Ramesh R. Organically derived exosomes as carriers of anticancer drugs and imaging agents for cancer treatment. Semin Cancer Biol. 2022;86(Pt 1):80-100. [CrossRef]
23. Thery C, Witwer KW, Aikawa E, Alcaraz MJ, Anderson JD, Andriantsitohaina R, et al. Minimal information for studies of extracellular vesicles 2018 (MISEV2018):A position statement of the International Society for Extracellular Vesicles and update of the MISEV2014 guidelines. J Extracell Vesicles. 2018;7(1):1535750. [CrossRef]
24. Salomon C, Das S, Erdbrügger U, Kalluri R, Kiang Lim S, Olefsky JM, et al. Extracellular vesicles and their emerging roles as cellular messengers in endocrinology:An endocrine society scientific statement. Endocr Rev. 2022;43(3):441-68. [CrossRef]
25. Huang Z, Zhao X, Wen W, Shi R, Liang G. Exosome miRNA sorting controlled by RNA-binding protein-motif interactions. Extracell Vesicles Circ Nucl Acids. 2025;6(3):475-503. [CrossRef]
26. Gurung S, Perocheau D, Touramanidou L, Baruteau J. The exosome journey: From biogenesis to uptake and intracellular signalling. Cell Commun Signal. 2021;19(1):47. [CrossRef]
27. Liu M, Wen Z, Zhang T, Zhang L, Liu X, Wang M. The role of exosomal molecular cargo in exosome biogenesis and disease diagnosis. Front Immunol. 2024;15:1417758. [CrossRef]
28. Tenchov R, Sasso JM, Wang X, Liaw WS, Chen CA, Zhou QA. Exosomes-nature's lipid nanoparticles, a rising star in drug delivery and diagnostics. ACS Nano. 2022;16(11):17802-46. [CrossRef]
29. Peng X, Yang L, Ma Y, Li Y, Li H. Focus on the morphogenesis, fate and the role in tumor progression of multivesicular bodies. Cell Commun Signal. 2020;18(1):122. [CrossRef]
30. Hikita T, Uehara R, Itoh RE, Mitani F, Miyata M, Yoshida T, et al. MEK/ERK-mediated oncogenic signals promote secretion of extracellular vesicles by controlling lysosome function. Cancer Sci. 2022;113(4):1264-76. [CrossRef]
31. Boucher JM, Robich M, Scott SS, Yang X, Ryzhova L, Turner JE, et al. Rab27a regulates human perivascular adipose progenitor cell differentiation. Cardiovasc Drugs Ther. 2018;32(5):519-30. [CrossRef]
32. Kiral FR, Kohrs FE, Jin EJ, Hiesinger PR. Rab GTPases and membrane trafficking in neurodegeneration. Curr Biol. 2018;28(8):R471-86. [CrossRef]
33. El-Masry TA, El-Nagar MMF, Oriquat GA, Alotaibi BS, Saad HM, El Zahaby EI, et al. Therapeutic efficiency of Tamoxifen/Orlistat nanocrystals against solid ehrlich carcinoma via targeting TXNIP/HIF1-a/MMP-9/P27 and BAX/Bcl2/P53 signaling pathways. Biomed Pharmacother. 2024;180:117429. [CrossRef]
34. Chitti SV, Gummadi S, Kang T, Shahi S, Marzan AL, Nedeva C, et al. Vesiclepedia 2024:An extracellular vesicles and extracellular particles repository. Nucleic Acids Res. 2024;52(D1):D1694-8. [CrossRef]
35. Li J, Wang J, Chen Z. Emerging role of exosomes in cancer therapy:Progress and challenges. Mol Cancer. 2025;24(1):13. [CrossRef]
36. Duan L, Xu L, Xu X, Qin Z, Zhou X, Xiao Y, et al. Exosome-mediated delivery of gene vectors for gene therapy. Nanoscale. 2021;13(3):1387-97. [CrossRef]
37. Kalluri R, LeBleu VS. The biology, function, and biomedical applications of exosomes. Science. 2020;367(6478):eaau6977. [CrossRef]
38. Lai RC, Yeo RWY, Tan KH, Lim SK. Mesenchymal stem cell exosome ameliorates reperfusion injury through proteomic complementation. Regen Med. 2013;8(2):197-209. [CrossRef]
39. Peinado H, Ale?kovi?M, Lavotshkin S, Matei I, Costa-Silva B, Moreno-Bueno G, et al. Melanoma exosomes educate bone marrow progenitor cells toward a pro-metastatic phenotype through MET. Nat Med. 2012;18(6):883-91. [CrossRef]
40. Alvarez-Erviti L, Seow Y, Yin H, Betts C, Lakhal S, Wood MJ. Delivery of siRNA to the mouse brain by systemic injection of targeted exosomes. Nat Biotechnol. 2011;29(4):341-5. [CrossRef]
41. Mulcahy LR, Isabella VM, Lewis K. Pseudomonas aeruginosa biofilms in disease. Microb Ecol. 2014;68:1-12. [CrossRef]
42. Rui T, Wang K, Xiang A, Guo J, Tang N, Jin X, et al. Serum exosome-derived piRNAs could be promising biomarkers for HCC diagnosis. Int J Nanomed. 2023;13(18):1989-2001. [CrossRef]
43. Fan B, Chopp M, Zhang ZG, Liu XS. Treatment of diabetic peripheral neuropathy with engineered mesenchymal stromal cell-derived exosomes enriched with microRNA-146a provide amplified therapeutic efficacy. Exp Neurol. 2021;341:113694. [CrossRef]
44. Zeng H, Guo S, Ren X, Wu Z, Liu S, Yao X. Current strategies for exosome cargo loading and targeting delivery. Cells. 2023;12(10):1416. [CrossRef]
45. Fu S, Wang Y, Xia X, Zheng JC. Exosome engineering:Current progress in cargo loading and targeted delivery. NanoImpact. 2020;20:100261. [CrossRef]
46. Xu M, Yang Q, Sun X, Wang Y. Recent advancements in the loading and modification of therapeutic exosomes. Front Bioeng Biotechnol. 2020;8:586130. [CrossRef]
47. Sato YT, Umezaki K, Sawada S, Mukai SA, Sasaki Y, Harada N, et al. Engineering hybrid exosomes by membrane fusion with liposomes. Sci Rep. 2016;6(1):21933. [CrossRef]
48. Zhang S, Dong Y, Wang Y, Sun W, Wei M, Yuan L, et al. Selective encapsulation of therapeutic mRNA in engineered extracellular vesicles by DNA aptamer. Nano Lett. 2021;21(20):8563-70. [CrossRef]
49. Iqbal Z, Rehman K, Mahmood A, Shabbir M, Liang Y, Duan L, et al. Exosome for mRNA delivery:Strategies and therapeutic applications. J Nanobiotechnology. 2024;22(1):395. [CrossRef]
50. Soltanmohammadi F, Gharehbaba AM, Zangi AR, Adibkia K, Javadzadeh Y. Current knowledge of hybrid nanoplatforms composed of exosomes and organic/inorganic nanoparticles for disease treatment and cell/tissue imaging. Biomed Pharmacother. 2024;178:117248. [CrossRef]
51. Amiri A, Bagherifar R, Ansari Dezfouli E, Kiaie SH, Jafari R, Ramezani R. Exosomes as bio-inspired nanocarriers for RNA delivery: Preparation and applications. J Transl Med. 2022;20(1):125. [CrossRef]
52. Munagala R, Aqil F, Jeyabalan J, Kandimalla R, Wallen M, Tyagi N, et al. Exosome-mediated delivery of RNA and DNA for gene therapy. Cancer Lett. 2021;505:58-72. [CrossRef]
53. Zhang Y, Liu Q, Zhang X, Huang H, Tang S, Chai Y, et al. Recent advances in exosome-mediated nucleic acid delivery for cancer therapy. J Nanobiotechnol. 2022;20(1):279. [CrossRef]
54. Zhou Y, Zhou G, Tian C, Jiang W, Jin L, Zhang C, et al. Exosome-mediated small RNA delivery for gene therapy. Wiley Interdiscip Rev RNA. 2016;7(6):758-71. [CrossRef]
55. Song Z, Tao Y, Liu Y, Li J. Advances in delivery systems for CRISPR/Cas-mediated cancer treatment:A focus on viral vectors and extracellular vesicles. Front Immunol. 2024;15:1444437. [CrossRef]
56. Balaraman AK, Babu MA, Moglad E, Mandaliya V, Rekha MM, Gupta S, et al. Exosome-mediated delivery of CRISPR-Cas9:A revolutionary approach to cancer gene editing. Pathol Res Pract. 2025;266:155785. [CrossRef]
57. Wan T, Zhong J, Pan Q, Zhou T, Ping Y, Liu X. Exosome-mediated delivery of Cas9 ribonucleoprotein complexes for tissue-specific gene therapy of liver diseases. Sci Adv. 2022;8(37):eabp9435. [CrossRef]
58. Kim H, Jang H, Cho H, Choi JS, Hwang KY, Choi Y, et al. Recent advances in exosome-based drug delivery for cancer therapy. Cancers (Basel). 2021;13(17):4435. [CrossRef]
59. Rezaie J, Ajezi S, Avc?ÇB, Karimipour M, Geranmayeh MH, Nourazarian A, et al. Exosomes and their application in biomedical field:Difficulties and advantages. Mol Neurobiol. 2018;55:3372-93. [CrossRef]
60. Xu S, Liu B, Fan J, Xue C, Lu Y, Li C, et al. Engineered mesenchymal stem cell-derived exosomes with high CXCR4 levels for targeted siRNA gene therapy against cancer. Nanoscale. 2022;14(11): 4098-113. [CrossRef]
61. Sarkami SA, Molavipordanjani S, Abediankenari S, Akhtari J, Gill P, Ghalehnoei H, et al. Engineering HEK293T cell line by lentivirus to produce miR34a-loaded exosomes. Mol Biol Rep. 2023;50:8827-37. [CrossRef]
62. Mei R, Wan Z, Li Z, Wei M, Qin W, Yuan L, et al. “All-in-one”exosome engineering strategy for effective therapy of familial hypercholesterolemia. ACS Appl Mater Interfaces. 2022;45(14):50626-36. [CrossRef]
63. Gul R, Bashir H, Sarfraz M, Shaikh AJ, Bin Jardan YA, Hussain Z, et al. Human plasma derived exosomes:Impact of active and passive drug loading approaches on drug delivery. Saudi Pharm J. 2024;32(6):102096. [CrossRef]
64. Pan Q, Deng X, Gao W, Chang J, Pu Y, He B. Small molecules-PEG amphiphilic conjugates as carriers for drug delivery:1. The effect of molecular structures on drug encapsulation. J Drug Deliv Sci Technol. 2020;60:101997.[CrossRef]
65. Abusara OH, Agha ASAA, Bardaweel SK. Advancements and innovations in liquid biopsy through microfluidic technology for cancer diagnosis. Analyst. 2025;150(9):1711-25. [CrossRef]
66. Xi XM, Xia SJ, Lu R. Drug loading techniques for exosome-based drug delivery systems. Pharmazie. 2021;76(2):61-7.
67. Semcheddine F, Guissi NEI, Liu W, Tayyaba, Gang L, Jiang H, et al. Rapid and label-free cancer theranostics via in situ bio-self-assembled DNA-gold nanostructures loaded exosomes. Mater Horiz. 2021;8(10):2771-84. [CrossRef]
68. Mondal J, Pillarisetti S, Junnuthula V, Saha M, Hwang SR, Park I, et al. Hybrid exosomes, exosome-like nanovesicles and engineered exosomes for therapeutic applications. J Control Release. 2022;353:1127-49. [CrossRef]
69. Agha ASA, Alshaer W, Aburjai T. Advancements and challenges in aptamer-based therapeutics and diagnostics across diverse medical domains:A comprehensive review. Jordan J Pharm Sci. 2024;17(2):344-61. [CrossRef]
70. Li S, Lin ZX, Jiang XY, Yu X. Exosomal cargo-loading and synthetic exosome-mimics as potential therapeutic tools. Acta Pharmacol Sin. 2018;39:542-51. [CrossRef]
71. Hao R, Yu Z, Du J, Hu SY, Yuan C, Guo H, et al. A high-throughput nanofluidic device for exosome nanoporation to develop cargo delivery vehicles. Small. 2021;17(35):e2102150. [CrossRef]
72. Gao X, Ran N, Dong X, Zuo B, Yang R, Zhou Q, et al. Anchor peptide captures, targets, and loads exosomes of diverse origins for diagnostics and therapy. Sci Transl Med. 2018;10(444):eaat0195. [CrossRef]
73. Hosseini NF, Amini R, Ramezani M, Saidijam M, Hashemi SM, Najafi R. AS1411 aptamer-functionalized exosomes in the targeted delivery of doxorubicin in fighting colorectal cancer. Biomed Pharmacother. 2022;155:113690. [CrossRef]
74. Smyth TJ, Petrova KB, Payton NM, Persaud I, Redzic JS, Graner MW, et al. Surface functionalization of exosomes using click chemistry. Bioconjug Chem. 2014;25:1777-84. [CrossRef]
75. Hung ME, Leonard JN. Stabilization of exosome-targeting peptides via engineered glycosylation. J Biol Chem. 2015;290:8166-72. [CrossRef]
76. Agha ASA, Al-Samydai A, Aburjai T. New frontiers in CRISPR:Addressing antimicrobial resistance with Cas9, Cas12, Cas13, and Cas14. Heliyon. 2025;11(2):e42013. [CrossRef]
77. Gupta N, Polkoff KM, Qiao L, Cheng K, Piedrahita JA. 200 Developing exosomes as a mediator for CRISPR/Cas-9 delivery. Reprod Fertil Dev. 2019;31(1):225. [CrossRef]
78. Ye Y, Zhang X, Xie F, Xu B, Xie P, Yang T, et al. An engineered exosome for delivering sgRNA:Cas9 ribonucleoprotein complex and genome editing in recipient cells. Biomater Sci. 2020;8(10):2966-76. [CrossRef]
79. Dubey S, Chen Z, Talis A, Molotkov A, AlìA, Mintz A, et al. An Exosome-Based Gene Delivery Platform for Cell-Specific CRISPR/Cas9 Genome Editing. BioRxiv;2023. [CrossRef]
80. Kim SM, Yang Y, Oh SJ, Hong Y, Seo M, Jang M. Cancer-derived exosomes as a delivery platform of CRISPR/Cas9 confer cancer cell tropism-dependent targeting. J Control Release. 2017;266:8-16. [CrossRef]
81. McAndrews KM, Xiao F, Chronopoulos A, LeBleu VS, Kugeratski FG, Kalluri R. Exosome-mediated delivery of CRISPR/Cas9 for targeting of oncogenic KrasG12D in pancreatic cancer. Life Sci Alliance. 2021;4(9):e202000875. [CrossRef]
82. Chen R, Huang H, Liu H, Xi J, Ning J, Zeng W, et al. Friend or foe?Evidence indicates endogenous exosomes can deliver functional gRNA and Cas9 protein. Small. 2019;15:e1902686. [CrossRef]
83. Ren X, Kang C, Garcia-Contreras L, Kim D. Understanding of ovarian cancer cell-derived exosome tropism for future therapeutic applications. Int J Mol Sci. 2023;24(9):8166. [CrossRef]
84. Javdani-Mallak A, Mowla SJ, Alibolandi M. Tumor-derived exosomes and their application in cancer treatment. J Transl Med. 2025;23(1):751. [CrossRef]
85. Abbas MA, Al-Kabariti AY, El-Rayyes R, Obeidat R, Al-Rayyes AA. TRPC1 and TRPC5 in colorectal cancer:Mechanisms, prognostic markers, and therapeutic targets. Drug Dev Res. 2025;86(5):e70123. [CrossRef]
86. Vakhshiteh F, Rahmani S, Ostad SN, Madjd Z, Dinarvand R, Atyabi F. Exosomes derived from miR-34a-overexpressing mesenchymal stem cells inhibit in vitro tumor growth:A new approach for drug delivery. Life Sci. 2021;266:118871. [CrossRef]
87. Shirley JL, de Jong YP, Terhorst C, Herzog RW. Immune responses to viral gene therapy vectors. Mol Ther. 2020;28(3):709-22. [CrossRef]
88. Almotiri A, Abogosh A, Abdelfattah A, Alowaisy D, Rodrigues NP. Treating genetic blood disorders in the era of CRISPR-mediated genome editing. Mol Ther. 2025;33(6):2645-62. [CrossRef]
89. Demirci S, Leonard A, Tisdale JF. Genome editing strategies for fetal hemoglobin induction in beta-hemoglobinopathies. Hum Mol Genet. 2020;29(R1):R100-6. [CrossRef]
90. Soltanmohammadi F, Maghsoodi M, Alizadeh E, Adibkia K, Azarmi Y, Mahmoudi Gharehbaba A, et al. Bio fluid exosomes:Promises, challenges, and future directions in translational medicine. J Transl Med. 2025;23(1):993. [CrossRef]
91. Nash LA, McFall ER, Perozzo AM, Turner M, Poulin KL, De Repentigny Y, et al. Survival motor neuron protein is released from cells in exosomes:A potential biomarker for spinal muscular atrophy. Sci Rep. 2017;7(1):13859. [CrossRef]
92. Sosero YL, Gan-Or Z. LRRK2 and Parkinson's disease:From genetics to targeted therapy. Ann Clin Transl Neurol. 2023;10(6): 850-64. [CrossRef]
93. Sanadgol N, Abedi M, Hashemzaei M, Kamran Z, Khalseh R, Beyer C, et al. Exosomes as nanocarriers for brain-targeted delivery of therapeutic nucleic acids:Advances and challenges. J Nanobiotechnology. 2025;23(1):453. [CrossRef]
94. Lerussi G, Villagrasa-Araya V, Molto-Abad M, Del Toro M, Pintos-Morell G, Seras-Franzoso J, et al. Extracellular vesicles as tools for crossing the blood-brain barrier to treat lysosomal storage diseases. Life (Basel). 2025;15(1):70. [CrossRef]
95. Chen FM, Liu X. Advancing biomaterials of human origin for tissue engineering. Prog Polym Sci. 2016;53:86-168. [CrossRef]
96. Gu A, Chen JY. GETgene-AI:A framework for prioritizing actionable cancer drug targets. Front Syst Biol. 2025;5:1649758. [CrossRef]
97. Holubowicz R, Du SW, Felgner J, Smidak R, Choi EH, Palczewska G, et al. Safer and efficient base editing and prime editing via ribonucleoproteins delivered through optimized lipid-nanoparticle formulations. Nat Biomed Eng. 2025;9(1):57-78. [CrossRef]
98. Sadeghi S, Tehrani FR, Tahmasebi S, Shafiee A, Hashemi SM. Exosome engineering in cell therapy and drug delivery. Inflammopharmacology. 2023;31(1):145-69. [CrossRef]
99. Capelli C, Cuofano C, Pavoni C, Frigerio S, Lisini D, Nava S, et al. Potency assays and biomarkers for cell-based advanced therapy medicinal products. Front Immunol. 2023;14:1186224. [CrossRef]
100. Azimi S. Intelligent nanoparticle design:Unlocking the potential of AI for transformative drug delivery. Curr Opin Biomed Eng. 2025;36:100625. [CrossRef]
101. Chou WC, Canchola A, Zhang F, Lin Z. Machine learning and artificial intelligence in nanomedicine. Wiley Interdiscip Rev Nanomed Nanobiotechnol. 2025;17(4):e70027. [CrossRef]
102. Agrahari V, Choonara YE, Mosharraf M, Patel SK, Zhang F. The role of artificial intelligence and machine learning in accelerating the discovery and development of nanomedicine. Pharm Res. 2024;41(12):2289-97. [CrossRef]
103. Welsh JA, Goberdhan DCI, O'Driscoll L, Buzas EI, Blenkiron C, Bussolati B, et al. Minimal information for studies of extracellular vesicles (MISEV2023):From basic to advanced approaches. J Extracell Vesicles. 2024;13(2):e12404. [CrossRef]
104. Shekari F, Alibhai FJ, Baharvand H, Borger V, Bruno S, Davies O, et al. Cell culture-derived extracellular vesicles:Considerations for reporting cell culturing parameters. J Extracell Biol. 2023;2(10):e115. [CrossRef]
105. Huang S, Chen L, Sun C, Zhang J, Chen T, Cao Y, et al. Electric Fields Trigger Ceramide-Dependent Budding of Exosome Vesicles into Multivesicular Endosomes and Boost the Generation of Exosomes. bioRxiv;2025. [CrossRef]
106. Zhou L, Kodidela S, Godse S, Thomas-Gooch S, Kumar A, Raji B, et al. Targeted drug delivery to the central nervous system using extracellular vesicles. Pharmaceuticals (Basel). 2022;15(3):358. [CrossRef]
107. Bracken MB. Why animal studies are often poor predictors of human reactions to exposure. J R Soc Med. 2009;102(3):120-2. [CrossRef]
108. Papakonstantinou E, Dragoumani K, Chrousos GP, Vlachakis D. Exosomal epigenetics. EMBnet J. 2024;29:e1049. [CrossRef]
109. Seijas A, Cora D, Novo M, Al-Soufi W, Sanchez L, Arana AJ. CRISPR/Cas9 delivery systems to enhance gene editing efficiency. Int J Mol Sci. 2025;26(9:4420. [CrossRef]
110. Tran HL, Zheng W, Issadore DA, Im H, Cho YK, Zhang Y, et al. Extracellular vesicles for clinical diagnostics:From bulk measurements to single-vesicle analysis. ACS Nano. 2025;19(31):28021-109. [CrossRef]
111. Kuang L, Wu L, Li Y. Extracellular vesicles in tumor immunity:Mechanisms and novel insights. Mol Cancer. 2025;24(1):45. [CrossRef]
112. Asmamaw Mengstie M, Teshome Azezew M, Asmamaw Dejenie T, Teshome AA, Tadele Admasu F, Behaile Teklemariam A, et al. Recent advancements in reducing the off-target effect of CRISPR-Cas9 genome editing. Biologics. 2024;18:21-8. [CrossRef]
113. Verma N, Arora S. Navigating the global regulatory landscape for exosome-based therapeutics:Challenges, strategies, and future directions. Pharmaceutics. 2025;17(8):990. [CrossRef]
114. Rodriguez-Gomez FD, Monferrer D, Penon O, Rivera-Gil P. Regulatory pathways and guidelines for nanotechnology-enabled health products:A comparative review of EU and US frameworks. Front Med (Lausanne). 2025;12:1544393. [CrossRef]
115. Lin M, Guo J, Gu Z, Tang W, Tao H, You S, et al. Machine learning and multi-omics integration:Advancing cardiovascular translational research and clinical practice. J Transl Med. 2025;23(1):388. [CrossRef]
116. Thomas KS, Edpuganti S, Puthooran DM, Thomas A, Joy A, Latheef S. Artificial intelligence in modern clinical practice (Review). Med Int (Lond). 2026;6(1):5. [CrossRef]