1. INTRODUCTION
Soil is not merely a medium for plant anchorage and nutrient supply; it is a dynamic ecosystem rich with microbial life that underpins essential ecological functions such as organic matter decomposition, nutrient mineralization, biogeochemical cycling, and plant-microbe interactions. Among the microbial residents of the rhizosphere the narrow zone of soil influenced by plant roots bacteria play a critical role in supporting plant health and productivity. Within this community, plant growth-promoting rhizobacteria (PGPR) represent a functional group of microorganisms that directly or indirectly benefit plant growth through a variety of mechanisms. These include biological nitrogen fixation, solubilization of insoluble phosphate, synthesis of phytohormones such as indole-3-acetic acid, production of siderophores that chelate iron, and induction of systemic resistance against pathogens [1,2]. Recent metagenomic studies have shown that PGPR inoculation can lead to significant shifts in the taxonomic and functional composition of rhizospheric microbial communities. For instance, inoculation with Bacillus subtilis has been reported to increase the relative abundance of beneficial taxa such as Actinobacteria and Proteobacteria while reducing populations of known phytopathogens. Similarly, Azospirillum brasilense treatment was found to alter the structure of wheat root-associated microbiota, enhancing microbial richness and stability [3,4]. Their potential to reduce chemical fertilizer dependence and enhance stress tolerance makes PGPR a cornerstone of sustainable agricultural strategies. In the context of tomato (Solanum lycopersicum), a high-value vegetable crop cultivated worldwide, PGPR have been widely explored as bioinoculants to enhance crop vigor, yield, and disease resistance. Previous research has demonstrated the effectiveness of Pseudomonas spp. and Burkholderia spp. in stimulating seedling growth, improving nutrient acquisition, and suppressing pathogens such as Fusarium oxysporum, Ralstonia solanacearum, and Phytophthora capsici [5]. These strains are known for their rapid colonization of root surfaces, biofilm formation, and secretion of antimicrobial compounds that confer competitive advantages in the rhizosphere. Despite these known benefits, most studies have focused primarily on plant phenotypic responses and yield outcomes, with relatively little attention paid to how PGPR inoculation reshapes the broader microbial community in the rhizosphere. Emerging evidence suggests that PGPR not only act as microbial bioeffectors but also as ecological engineers capable of influencing the structure and dynamics of native microbial communities [6]. Their introduction into the soil ecosystem may lead to competitive interactions, niche displacement, or synergistic effects that modulate the abundance and activity of other soil microbes. These changes can, in turn, affect critical processes such as nutrient cycling, organic matter decomposition, and suppression of soil-borne diseases [7]. The extent and direction of such community-level shifts are likely to depend on a combination of factors, including the PGPR strain used, soil type, plant developmental stage, and environmental conditions. Understanding these microbial interactions is crucial for predicting the long-term effects of PGPR inoculation and for developing bioformulations that are compatible with existing soil microbial ecosystems. Traditionally, studies investigating soil microbial communities have relied on culture-dependent techniques that are limited by the inability to grow the vast majority of soil microbes under laboratory conditions. It is estimated that over 90% of soil bacterial taxa are unculturable using standard methods, leading to an incomplete and biased understanding of microbial diversity. The advent of culture-independent approaches such as next-generation sequencing and metagenomics has revolutionized soil microbiology by enabling high-throughput, in-depth characterization of microbial communities directly from environmental DNA samples [8,9]. Among these, 16S rRNA gene amplicon sequencing has emerged as a powerful tool for taxonomic profiling of bacterial communities at high resolution, allowing researchers to identify dominant taxa, detect rare species, and assess community composition and diversity metrics such as alpha and beta diversity. Recent metagenomic studies have shown that PGPR inoculation can lead to significant shifts in the taxonomic and functional composition of rhizospheric microbial communities. For instance, inoculation with Bacillus subtilis has been reported to increase the relative abundance of beneficial taxa such as Actinobacteria and Proteobacteria, while reducing populations of known phytopathogens. Similarly, A. brasilense treatment was found to alter the structure of wheat root-associated microbiota, enhancing microbial richness and stability [10,11]. Despite these insights, there remains a significant knowledge gap regarding how different PGPR strains differentially influence rhizospheric microbiomes, particularly in controlled settings that eliminate environmental variability. Most available studies have focused on single-strain inoculations or field trials, which, although valuable, make it difficult to isolate the specific effects of individual PGPR species on microbial community structure. This gap is particularly relevant during the early stages of plant development, a critical period for root microbiome assembly and plant-microbe signaling. Rhizosphere colonization by microbes during seedling establishment can have long-term consequences on plant growth trajectories, disease susceptibility, and stress resilience. Therefore, investigating how PGPR strains influence the initial microbial community dynamics can provide important clues for the rational design of microbial consortia tailored for specific crops or soil conditions. This study addresses this gap by applying 16S rRNA gene-based metagenomic analysis to compare the effects of two PGPR species Pseudomonas aeruginosa and Burkholderia dolosa—on the bacterial community structure in the rhizosphere of tomato under greenhouse conditions. These two strains represent taxonomically and functionally distinct bacterial groups with known plant-beneficial traits, yet their comparative impacts on native soil microbial communities remain largely uncharacterized. By conducting the experiment in a controlled greenhouse environment with sterilized sandy loam soil, we minimize environmental variability and focus on the microbial shifts induced specifically by PGPR inoculation. The novelty of this study lies in its integrative approach: we not only evaluate microbial diversity using both alpha and beta diversity indices but also identify taxonomic shifts at multiple phylogenetic levels (phylum, class, order, family, and genus) and determine the core and unique taxa associated with each treatment. Furthermore, we compare the microbial profiles of inoculated soils against an uninoculated control to assess how PGPR treatments restructure the rhizospheric microbiome. This approach allows for a mechanistic understanding of microbial community assembly in response to bioinoculant application.
2. MATERIALS AND METHODS
2.1. Greenhouse Experimental Design
The greenhouse experiment was conducted to evaluate the effect of PGPR inoculation on the soil bacterial community structure under tomato cultivation. The trial was set up using sterilized soil in earthen pots (20 cm diameter × 25 cm height), each filled with 5 kg of sandy loam soil [Figure 1]. Prior to transplanting, soil samples were analyzed for key physicochemical properties. Soil pH (6.8) was measured in a 1:2.5 soil-to-water suspension using a digital pH meter, and electrical conductivity (0.45 dS/m) was determined using a conductivity meter. Organic carbon content (0.48%) was estimated by the Walkley–Black method. Total nitrogen (210 mg/kg) was quantified using the Kjeldahl method, available phosphorus (18.6 mg/kg) was measured by the Olsen method, and potassium (165 mg/kg) was extracted using ammonium acetate and measured through flame photometry [12,13]. These baseline values helped characterize the initial soil environment for microbial and plant response assessments. Tomato seedlings (S. lycopersicum L.) were then transplanted into each pot. The experiment followed a completely randomized design with three treatments: uninoculated control, P. aeruginosa inoculated, and Burkholderia dolosa inoculated, each in triplicate [Table 1] [14,15].
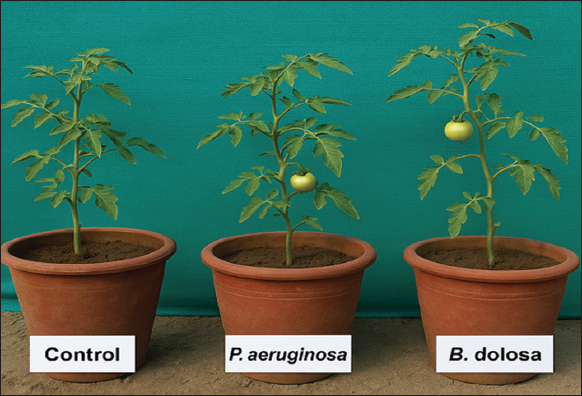 | Figure 1: Greenhouse pot experiment evaluating the effect of plant growth-promoting rhizobacteria inoculation on tomato growth. Three treatments were used: uninoculated control, Pseudomonas aeruginosa, and Burkholderia dolosa. [Click here to view] |
Table 1: Experimental design for PGPR inoculation in tomato plants under greenhouse conditions.
| Treatment | Plant height (cm) | No. of fruits/plant | Fresh biomass (g/plant) | Dry biomass (g/plant) | Soil bacterial count (CFU/g) | Shannon diversity index (H’) |
|---|---|---|---|---|---|---|
| T0: Control | 32.5±2.1c | 3.1±0.3c | 115.4±5.7c | 28.2±1.4c | 1.2×106c | 1.88±0.06c |
| T1: Pseudomonas aeruginosa | 46.2±1.8b | 6.7±0.5b | 162.7±6.1b | 42.5±2.0b | 3.9×106b | 2.51±0.09b |
| T2: Burkholderia dolosa | 50.3±2.4a | 7.5±0.6a | 178.9±7.4a | 48.3±2.3a | 4.5×106a | 2.68±0.07a |
Values are mean±standard deviation (n=3). Different superscript letters (a, b, c) within a column indicate significant differences at P<0.05 based on one-way analysis of variance followed by Tukey’s HSD test.
2.2. PGPR Strains and Inoculum Preparation
Two PGPR strains P. aeruginosa and B. dolosa previously isolated and submitted to NCBI with accession number of PV090851 and PQ059698 from tomato rhizosphere and characterized for plant growth promoting traits were used in this study. Bacterial cultures were grown in nutrient broth at 28 ± 2°C for 24 h, centrifuged at 6000 rpm, and resuspended in sterile distilled water to achieve a final cell concentration of ~108 CFU/mL. Each seedling received 10 mL of bacterial suspension applied directly to the root zone at the time of transplanting [16,17].
2.3. Soil Sampling and DNA Extraction
Rhizospheric soil samples were collected at two time points: pre-inoculation (Day 0) and post-inoculation (Day 30). Soil adhering to the root zone was carefully brushed into sterile bags, transported on ice, and stored at −80°C until DNA extraction. Total soil DNA was extracted using the DNeasy PowerSoil Kit (Qiagen, Germany) according to the manufacturer’s protocol. DNA concentration and purity were assessed using NanoDrop spectrophotometry and agarose gel electrophoresis [18,19].
2.4. 16S rRNA Amplicon Sequencing
The V3–V4 hypervariable regions of the bacterial 16S rRNA gene were amplified using universal primers 341F and 806R. The polymerase chain reaction (PCR) products were purified, quantified, and used to construct sequencing libraries with the Illumina MiSeq platform (2 × 300 bp paired-end reads). Sequencing was performed at Biokart India Pvt. Ltd. [20,21].
2.5. Bioinformatics and Data Analysis
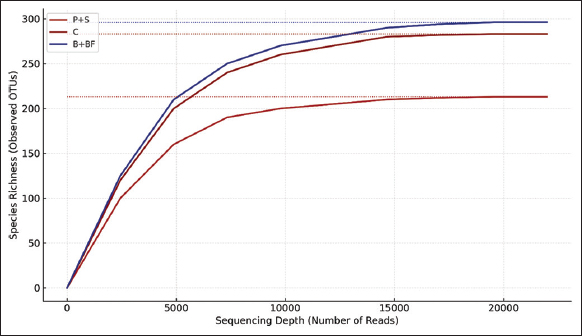
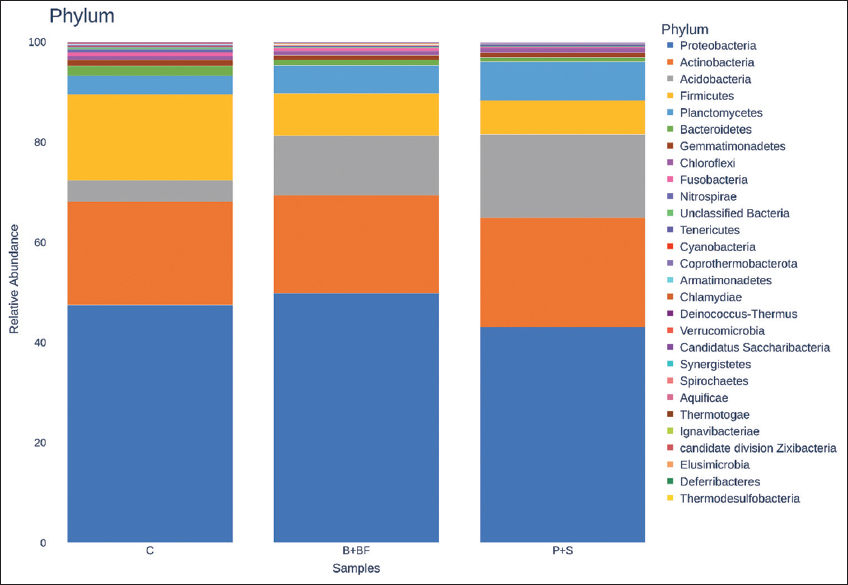
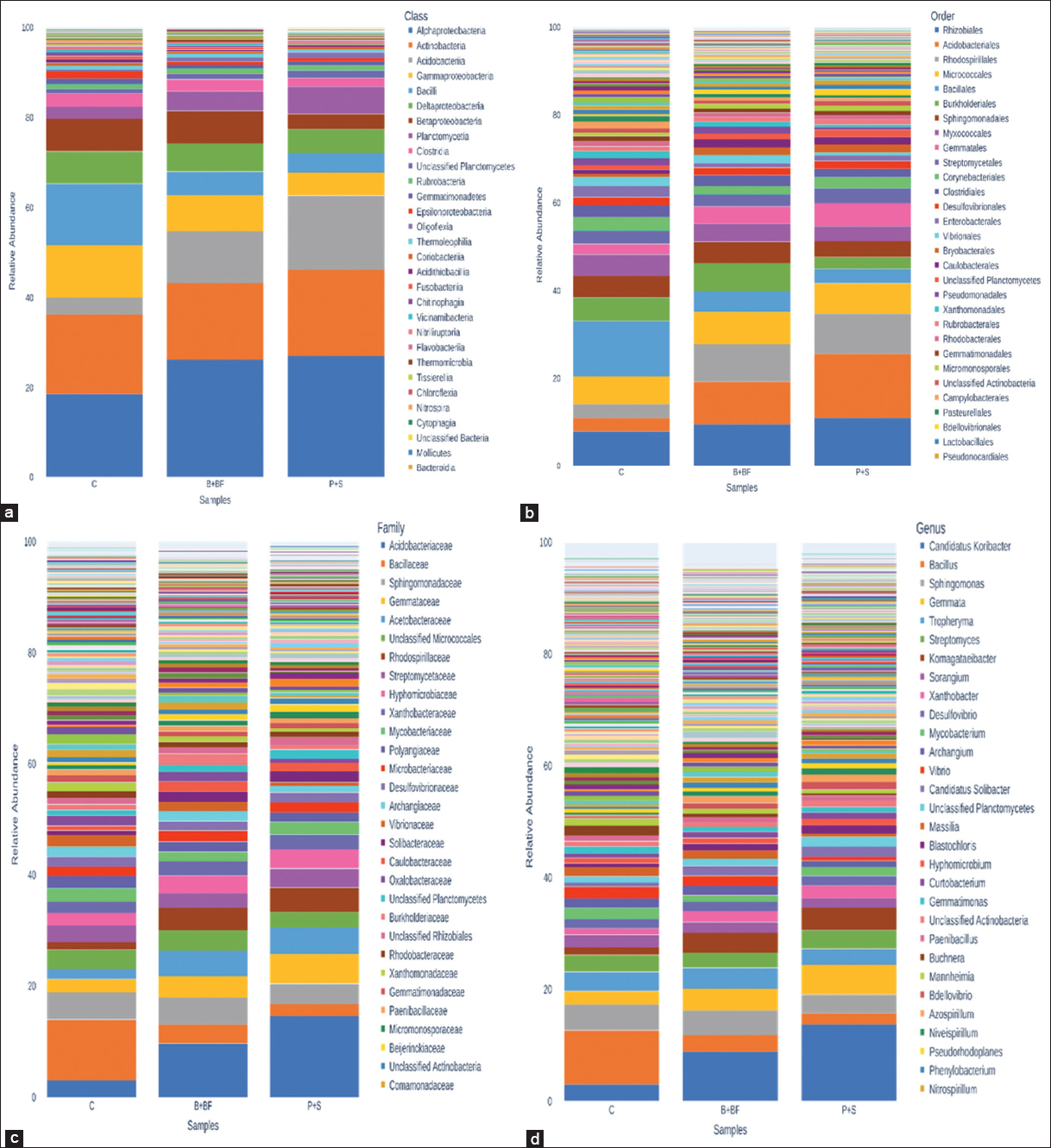
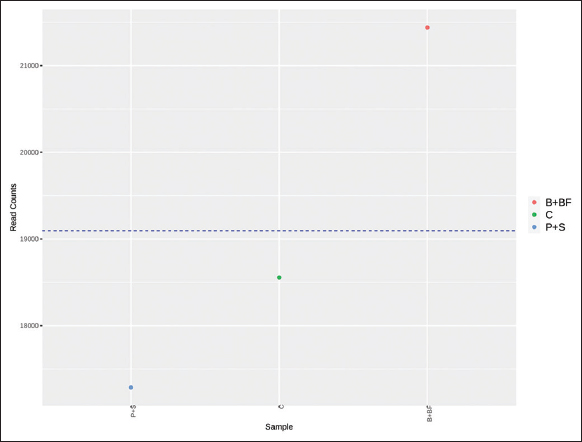
Raw sequence reads were quality-checked using FastQC (v0.11.9) and summarized using MultiQC. Sequence processing was carried out using QIIME2 (version 2023.2). DADA2 was used for denoising with truncation parameters set to 280 bp (forward) and 240 bp (reverse), using pooled error modeling and chimera removal by consensus. Operational taxonomic units (OTUs) were clustered at 97% similarity, and taxonomic classification was assigned using the SILVA 138 reference database. Since no biological replicates (per treatment) were sequenced with metagenomics, subsequent statistical testing (e.g. analysis of variance [ANOVA], permutational multivariate analysis of variance [PERMANOVA], differential abundance, etc.) was not conducted. They have been instead produced using alpha diversity indices (Shannon Index, Chao1) and beta diversity patterns (Bray Curtis, and unweighted UniFrac distances) and presented descriptively rather than implying any statistical inference. R packages phyloseq and vegan were used to visualize the differences in community composition by creating principal coordinate analysis (PCoA) plots, taxonomic bar charts, and heatmaps of the 50 most abundant genera. The missing biological replication precluded the use of any functional prediction (e.g. PICRUSt2) or ecological network analysis.