Since the onset of industrialization, the natural water balance in many agroclimatic zones has been disrupted by numerous anthropogenic activities. These activities interfere with the natural dispersion of salt across regions and degrade natural and agricultural habitats. Soil salinization is a major threat to global food safety due to changing environmental conditions. Alternative strategies are required to maintain agricultural output in saline-stressed soil and enhance crop production in an environmentally friendly way. The application of beneficial salt-tolerant microbes may improve plant survival under stressed conditions by altering numerous biochemical, physical, and molecular pathways. These microbes employ several strategies to thrive and reproduce under such conditions, including the production of growth hormones, lowering stress hormone ethylene levels, nitrogen (N) fixation, and improved nutrient and water uptake. Alterations in genes linked to the salt overly sensitive (SOS) pathway, together with increased expression of diverse antioxidant enzymes, are among the defined mechanisms used by stress-tolerant microbes to overcome the detrimental effects of salinity stress. Omics and meta-omics studies are powerful tools for uncovering the pathways of unknown genes. However, more precise omics investigations require a thorough understanding of the known molecular mechanisms underlying salinity stress tolerance in plants mediated by growth-promoting microbes.
Salinization of freshwater resources and soil by natural or anthropogenic activities has become a growing problem, disrupting ecological services and socioeconomic connections. In addition, salinity stress threatens agricultural ecosystems, affecting the majority of cultivated crops. Increasing salt stress conditions directly affect plant growth and the quality and quantity of food production worldwide [1]. Plants are generally categorized into halophytes (that can survive under salt stress) and the glycophytes (that cannot survive under salt stress and ultimately die). Most plant species fall under the second category. Therefore, this is the most severe stress that impedes crop production globally [2]. Salt stress influences plants at biochemical, physiological, morphological, and molecular levels. This stress results in reduced chlorophyll content, plant biomass, plant necrosis, reduced carbon fixation, membrane damage, stomatal closure, reduced photosynthesis, and reduced phytohormone production [3,4].
Higher soil salt concentrations affect plant resilience by disrupting cellular homeostasis and impairing key physiological and biochemical processes. Excess Na+ and Cl− ions damage plant cells through osmotic and oxidative stress [5]. At early stages of salinity, osmotic stress results in a range of physiological alterations, including reduced photosynthetic activity, nutrient imbalance, membrane disruption, diminished potential to detoxify reactive oxygen species (ROS), and reduced stomatal aperture [6]. In addition, reduced photosynthetic rate due to salt stress also exerts a harmful impact on plants’ vegetative and reproductive growth [7]. Ion toxicity restricts the entry of nutrients, including Ca2+ and K+, resulting in a nutritional imbalance in plants under salt stress. The photosynthetic apparatus is also damaged by ion toxicity, which blocks photosystem II reaction center, the electron transport chain, and the oxygen-evolving complex [8].
Numerous physiological and molecular mechanisms have evolved in plants to deal with salt stress conditions. Recent information from salt stress tolerance mechanisms emphasizes osmolytes that are crucial for osmotic regulation, free radical quenching, and antioxidant machinery. Polyamines (PAs) also play an advantageous role in stress tolerance and adaptation strategies, generally by acid neutralization, membrane stabilization, and suppression of ROS synthesis. The association of nitric oxide and PAs biosynthesis also raises expectations for improved plant stress tolerance [9]. The SOS pathway is a vital mechanism for ion homeostasis in plants that transports surplus Na+ ions for removal or sequestration. This is an essential defense mechanism and signaling pathway for plant growth under salinity stress conditions. Nevertheless, the mechanism by which components of the SOS pathways are built into plants to enable plant stress tolerance remains unclear [10].
In addition to the plants, many plant growth-promoting (PGP) microbes also have specific mechanisms that play a crucial role in salinity stress tolerance and plant development under stressed conditions. These PGP microbes activate the production of various growth hormones in plants, such as cytokinin (CKX2), gibberellin, and auxin. They are also known to produce important volatile organic compounds and growth regulators for nitrogen (N) fixation, solubilization of essential nutrients, and siderophore production [11]. The nitrogenase enzyme is responsible for N fixation, which is common in the majority of N-fixing bacteria. It is a complex enzyme with two metal components, including dinitrogenase reductase and dinitrogenase MoFe. The nifH gene coding for Fe dinitrogenase reductase and the nifD and K genes coding for MoFe dinitrogenase [12]. In an investigation, Muchhadiya et al. [13] reported the identification of stress-tolerant Rhizobium species, containing nif and nod genes responsible for N-fixation.
The stress-tolerant microbes are also able to solubilize phosphorus (P) by releasing H+ to the outer surface in exchange for cation intake. The phosphates are mostly solubilized by the acidification of soil by organic acid production [14]. Gluconic acid (GA) and 2-ketogluconic acid are among the most common organic acids produced by microbes for P-solubilization. Glucose-1-dehydrogenase produces GA, and pyrroloquinoline quinine acts as a cofactor [15]. SOS signaling pathways are used by the plants to regulate ion homeostasis and lessen Na+ influx. Stress-tolerant microbes also have the ability to upregulate the genes involved in the SOS pathway, H+-pumping pyrophosphatase (H+-ppase), and Na+/H+ antiporter to alleviate ion toxicity in plants [16]. The prevention of water loss is crucial to reducing osmotic stress in plant cells. Under salinity stress conditions, plants produce various osmoprotectants (most common are sugars, ammonium compounds, and some amino acids) that help them thrive under such conditions [17]. Stress adaptive PGP microbes mitigate oxidative stress either by the regulation of plant genes involved in the production of ROS-scavengers or by the synthesis of antioxidative enzymes [18].
Many stress adaptive microbes are reported to produce phytohormones, exopolysaccharides (EPS), and 1-aminocyclopropane-1-carboxylate (ACC) deaminase enzyme that helps plants to thrive under stressed conditions [Figure 1]. In a study, Ayaz et al. [19] demonstrated that plants inoculated with a salt-tolerant Bacillus strain under stress conditions considerably overexpressed genes linked with auxin (ARF), CKX2, and ethylene encoded genes. The ACC deaminase enzymes play an important role in stress mitigation and thus increase plant growth. The detailed analysis of the AcdS gene encoding for ACC deaminase reveals its occurrence in a range of beneficial microbes [20]. EPS produced by the microbes connect Na+ cations, which decreases their amount around the plant root zone, thus serving in ionic balance maintenance. They also help in water retention, soil aggregation, and metal ion chelation in plants under salt stress [21]. The presence of the complete eps gene cluster has been investigated in the chromosomal DNA of Streptococcus thermophilus. Among all eps gene clusters, epsA, B, C, and D genes were extremely conserved and were accountable for EPS regulation and polymerization [22].
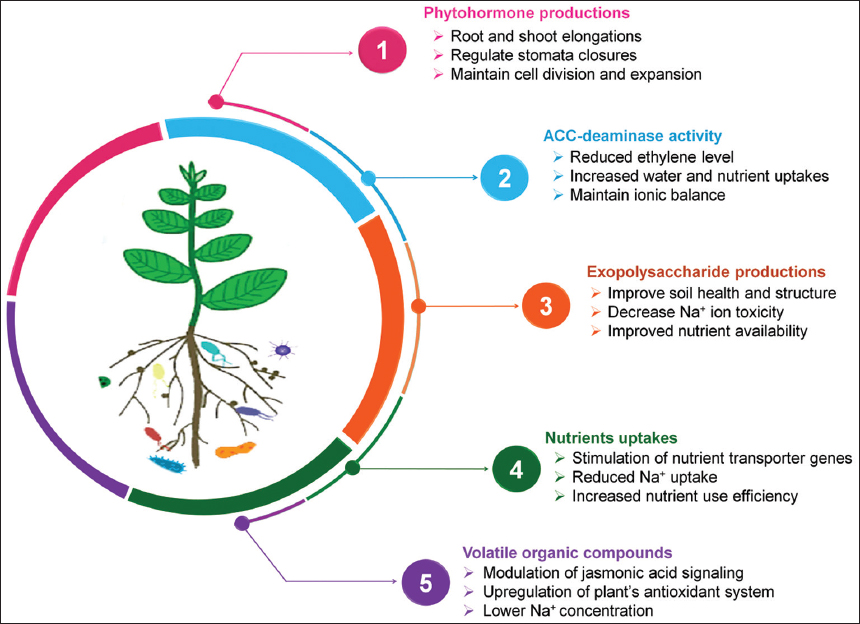 | Figure 1: Mechanistic insights of microbes mediated alleviation of salt stress. Adapted with permission from Sharma et al. [23]. [Click here to view] |
The identification of novel stress-adaptive microbial inoculants, perseverance of microbial inoculants in soil, and examining population density of functional genes and microbial gene sequences is an endless process to attain a desirable impact on crop growth and production [24]. The omic technologies are used for the identification of diverse protein expression, transcription factors, gene expression, and plant-microbial interaction in response to salinity stress tolerance [25]. The omic approaches, including meta-transcriptomics and metagenomics, offer enormous potential to recognize multifarious microbial networks concerned with stress signaling. The high-throughput screening of important microbes is facilitated by genomics technologies, exploiting genetic engineering and gene modification techniques for addressing salinity stress tolerance in plants. The plant microbiome engineering could be enormously advantageous in the expansion of approaches to improve crop productivity, stress resistance, and plant health, and to accomplish sustainable agriculture in an environmentally friendly manner [26].
The utilization of stress-adaptive microbes for plant stress tolerance is among the most important strategies used for the mitigation of adverse effects of salt stress in various crops. However, several reports suggest that superior results are obtained only under laboratory and greenhouse conditions, as opposed to the field conditions, where outcomes fluctuate. Therefore, the most important problem is to discover the difficulty and anticipate the present investigations in a way that all potential functional features of stress-tolerant microbes could be proficiently used for the formation of efficient bioinoculants for crops growing under saline agricultural ecosystems. The bottom-up approaches are essential to overcome these difficulties: screening for the best stress-adaptive microbial strains, examining their stress tolerance, preparing effective formulations, and, in the end, utilizing them under field conditions. The “Next Generation Agriculture” consists of the use of gene-editing tools, such as CRISPR, to engineer exclusive plant microbiomes and to engineer the metabolic pathways of stress-adaptive microbes to increase the utmost plant resistance.