1. INTRODUCTION
Paddy (Oryza sativa L.) is one of the most important cereal crops worldwide and serves as a staple food for nearly half of the global population. Major rice-producing countries, such as China, India, Indonesia, Bangladesh, and Vietnam play a critical role in sustaining global food security, with India alone producing more than 135 million metric tons annually [1]. Despite high production levels, post-harvest losses caused by fungal contamination remain a major constraint affecting grain quality and food safety. Stored paddy grains are particularly vulnerable to infestation by fungal pathogens, such as Aspergillus niger and Aspergillus flavus, which are responsible for grain deterioration and mycotoxin contamination [2-5]. These fungi pose serious health risks, including nephrotoxicity, immunosuppression, respiratory disorders, and carcinogenic effects, especially under prolonged exposure [6]. Increasing resistance to chemical fungicides and concerns regarding residual toxicity have intensified the need for sustainable and environmentally safe alternatives for post-harvest disease management. Plant growth-promoting rhizobacteria (PGPR) have been widely investigated as eco-friendly biological agents for improving crop productivity and suppressing phytopathogens. Among them, fluorescent Pseudomonas species have gained particular attention due to their ability to produce a diverse array of antifungal metabolites, including siderophores, hydrolytic enzymes, volatile organic compounds, phenazines, pyoverdine, pyrrolnitrin, and 2,4-diacetylphloroglucinol (DAPG) [7,8,26]. These metabolites inhibit fungal pathogens through multiple mechanisms, such as iron sequestration, disruption of fungal membrane integrity, interference with respiratory metabolism, and induction of oxidative stress [31,34].
Bio-priming, defined as the treatment of seeds with beneficial microorganisms before sowing or storage, is widely recognized as a sustainable strategy for improving seed germination, seedling vigor, and disease resistance [15,26]. Numerous studies have demonstrated that bio-priming with live PGPR, such as Pseudomonas fluorescens, Bacillus subtilis, and Trichoderma harzianum, enhances early plant establishment and reduces seed- and soil-borne fungal infections [12,15]. Bio-priming with Pseudomonas spp. has been reported to suppress fungal pathogens, including Aspergillus and Fusarium, through effective seed surface colonization and in situ production of antifungal metabolites, such as DAPG, phenazines, and siderophores [4,5,24]. Similarly, Bacillus- and Trichoderma-based bio-priming approaches have been shown to improve disease resistance through competitive exclusion, production of lytic enzymes, and induction of host defense responses [6-8]. However, the efficacy of conventional bio-priming strategies relying on live microbial inocula is strongly influenced by environmental factors, microbial survival during storage, and inconsistent colonization of the seed surface or rhizosphere [9,26]. These limitations are particularly pronounced in post-harvest and storage environments, where conditions may not favor microbial persistence or metabolic activity. As a result, despite extensive research on live-cell bio-priming, the direct application of cell-free, metabolite-based formulations remains relatively underexplored, especially for managing storage-associated fungal deterioration of paddy grains.
Extracellular antifungal metabolites produced by PGPR, particularly DAPG and phenazines from Pseudomonas spp., are well documented for their broad-spectrum antifungal activity against phytopathogens, such as Fusarium, Rhizoctonia, and Aspergillus [24,30,34]. Nevertheless, most studies have focused on metabolite identification or rhizosphere-mediated disease suppression during early plant growth stages, rather than evaluating the functional efficacy of metabolite-rich supernatants as standalone bio-priming agents. The comparative effectiveness of whole-cell bacterial formulations versus extracellular metabolite-based treatments under post-harvest or storage-stage conditions, therefore, remains insufficiently addressed. In this context, the present study aims to bridge this knowledge gap by isolating and characterizing a locally adapted P. fluorescens strain and functionally comparing the antifungal efficacy of its bacterial broth and cell-free, metabolite-rich supernatant against storage-associated fungal pathogens of paddy. By focusing on a supernatant-mediated, metabolite-based bio-priming strategy, this work evaluates an alternative approach that may offer greater consistency and applicability for post-harvest fungal management, thereby contributing to the development of sustainable and eco-friendly alternatives to chemical fungicides.
2. MATERIALS AND METHODS
2.1. Sampling Area and Cultivar
The experimental fields were located in the Gangoh Block of Saharanpur District, Uttar Pradesh, India. Throughout the study, a commonly cultivated paddy variety, “Pusa Basmati-1509”, was used.
2.2. Isolation and Characterization of Rhizobacteria and their Plant Growth Promoting Traits
Soil samples were collected from the rhizospheric zone of paddy crops. Bacteria were isolated from loosely attached soil particles using the serial dilution method, and dilutions ranging from 10-1 to 10-6 were prepared [9,10]. From each dilution, 100 μL was spread onto selective fluorescent Pseudomonas medium (HiMedia) plates in triplicate and incubated at 29 °C for 48 h. Seven bacterial strains were isolated and purified using the streak plate method. All isolates were examined for fluorescence under UV light. Further, kit-based biochemical examination was completed using the KB-002 biochemical test kit. Each well of the KB-002 kit was inoculated with a standardized 0.5 McFarland suspension and incubated at 28 ± 2 °C for 24–48 h. Results were interpreted based on colour changes according to the manufacturer’s instructions. Pseudomonas fluorescens MTCC 2421 served as the positive control, while uninoculated wells served as negative controls. All isolates were screened for siderophore production using the chrome azurol S (CAS) assay [11]. Additionally, indole-3-acetic acid (IAA) production was analyzed as described by [12]. Biochemical traits were further compared with the reference strain P. fluorescens MTCC 2421 to confirm identification. Among the isolated strains, the most potent strain, MS-05, was further analyzed by 16S rRNA gene sequencing for species-level identification following the method described in [13]. The fluorescent Pseudomonas strain was submitted to the NCBI database to obtain an accession number.
2.3. Fungal Deteriorating Agent
The fungal deteriorating agent for Aspergillus niger was isolated from 1-year-old stored paddy grains following the method of Chitranshi and Arora [14]. Briefly, infected rice grains were placed on freshly prepared PDA plates and incubated for 7–10 days at 25–29°C. Further, the fungal strain was preserved on potato dextrose agar plates at 4°c in the departmental laboratory.
2.4. Assessment of the Biocontrol Potential of P. fluorescens against Fungal Pathogens
2.4.1. Dual culture assay
The dual culture assay was performed in triplicate to assess the antifungal activity of potent bacterial strains against selected pathogenic fungi, with minor modifications from Sultana and Hossain [15]. Briefly, a 5 mm disc from a 1-week-old fungal culture was placed 1.5 cm from the left edge of a PDA-containing Petri dish. The bacterial culture was then inoculated 1.5 cm from the right edge of the same plate. A control plate containing only the fungal inoculum was also prepared. In addition, a positive control was maintained using 0.2% Mancozeb, a standard fungicide with broad-spectrum antifungal activity, while the negative control consisted of plates with only the fungal culture without bacterial inoculation. All plates were incubated at 29°C for 1 week in separate incubators to prevent potential cross-contamination, ensuring independent and sterile growth conditions for each treatment. The antagonistic activity of the bacterial strains was evaluated by observing the inhibition of fungal growth.
2.4.2. Supernatant spray assay
The collected solvent layer was evaporated under sterile airflow and the dried extract was reconstituted in sterile distilled water (10 mg/mL). The extract was then passed through a 0.22 μm syringe filter to ensure complete removal of bacterial cells. For the spray assay, 1 mL of the filtered supernatant was uniformly applied to each 90 mm PDA plate using a sterile atomizer. All plates received equal volume, and the treated plates were air-dried for 10 min in a laminar flow hood before fungal inoculation.
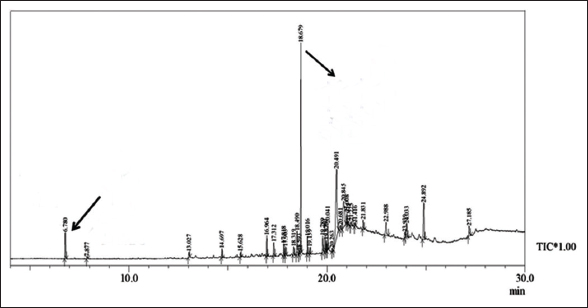
2.4.3. GC-MS analysis for bacterial metabolite identification
For the detection of metabolites produced during the dual culture assay, P. fluorescens MS-05 was analyzed further by Gas Chromatography and mass spectrometry (GC-MS) technique as described by Abed et al. [16]. The GC–MS analysis was carried out using an Agilent 7890A Gas Chromatograph coupled with a 5975C Mass Selective Detector. Separation of metabolites was achieved on an HP-5MS capillary column (30 m × 0.25 mm × 0.25 μm film thickness). Helium was used as the carrier gas at a constant flow rate of 1.0 mL/min. The injector temperature was set at 250°C, the injection volume was 1 μL, and the split ratio was 10:1. The oven temperature program started at 60°C (2 min hold), increased to 280°C at 10°C/min, and was held for 10 min. The ionization mode was electron impact (EI) at 70 eV, and mass spectra were recorded over the m/z range 50–600. Compound identification was performed by comparing retention times and mass spectra with those in the NIST 14 library and with authentic standards of DAPG and phenazine-1-carboxylic acid (PCA). Briefly, P. fluorescens MS-05 was cultured in nutrient broth medium (Hi-media) in Erlenmeyer flasks, the flasks were placed on a rotary shaker incubator at room temperature (approximately 25–30°C) with 110 rpm for 7 days. Further, the culture broth was filtered, and the filtered substances were mixed separately with chloroform and ethanol absolute (research grade) in a 1:1 ratio. The addition of chloroform resulted in the formation of two layers in the culture broth; the solvent layer was separated using a separating funnel and stored in sterile vials. The ethanol-added culture broth was kept as an aqueous extract.
2.5. Statistical Analysis
All experiments were conducted in triplicate (n = 3). Data presented as mean ± SD; significance determined by one-way ANOVA (P < 0.05).
3. RESULTS AND DISCUSSION
3.1. Morphological and Biochemical Identification of Rhizobacteria
The morphological identification of seven bacterial isolates through a compound microscope (Olympus 14C0504) shows that all isolates MS-01, MS-02, MS-03, MS-04, MS-05, MS-06, and MS-07, were Gram-negative and rod-shaped. Moreover, these isolates were further analyzed for various bio-chemical activities [Table 1]. All the strains were further checked for IAA and siderophore activity test and found that bacterial strain MS-05 shows the best IAA production, including positive siderophore production [Table 2]. Bacterial strain MS-05 shows the best results in all the kit-based bio-chemical test specifically for P. fluorescens. Similar observation in IAA activity was reported earlier by the researchers [17,18]. Further, it was observed that within a short period of time, considerable enhancement in the germination was observed in case of the crop treated with IAA [19]. Siderophore production by beneficial microorganisms is known to play an essential role in natural biocontrol by restricting iron availability to fungal pathogens [20-22].
Table 1: Biochemical identification of isolates by KB-002 biochemical test kit Hi-Media (Gram-positive Rods) (+) denotes positive results and (–) denotes negative results for biochemical characterization.
| Biochemical tests | MS-01 | MS-02 | MS-03 | MS-04 | MS-05 | MS-06 | MS-07 |
|---|---|---|---|---|---|---|---|
| Oxidase | + | + | + | + | + | + | + |
| Catalase | + | + | + | + | + | + | + |
| H2S production | -- | -- | -- | -- | -- | -- | -- |
| Nitrate reduction | + | + | -- | -- | + | -- | --- |
Table 2: Qualitative analysis of isolates for PGP activity (+) week (++) moderate and (+++) high and (--) negative results, IAA (indole-3-acetic acid).
| Qualitative | Quantitative | ||
|---|---|---|---|
| Sample ID | IAA | Siderophore | IAA |
| MS-01 | + | -- | 0.746±0.29 |
| MS-02 | + | -- | 01.164±0.42 |
| MS-03 | + | -- | 0.918±0.38 |
| MS-04 | + | -- | 2.465±0.90 |
| MS-05 | +++ | +++ | 10.925±3.69 |
| MS-06 | + | -- | 1.14±0.43 |
| MS-07 | ++ | -- | 4.668±1.90 |
IAA: Indole-3-acetic acid
3.2. Molecular Characterization
Antagonistically active bacterial strain MS-05 was screened further on various basic tests among all other seven strains. The strain was Gram-negative and rod-shaped, which shows positive oxidase and catalase tests with fluorescent property under UV light. Further, in 16s rRNA sequencing (Sanger method) strain was showing 99% similarities with Pseudomonas [23]. Moreover, a comparative study with the available database on NCBI and the phylogenetic analysis [Figure 1] of that strain MS-05 shows the direct relation with P. fluorescens. Further, the sequence was submitted to the NCBI database with Accession No: PP501522.1.
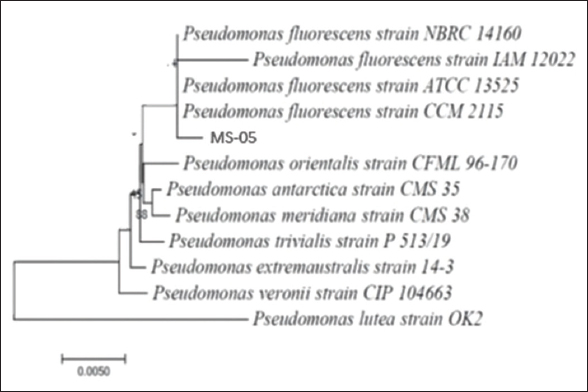 | Figure 1: Phylogenetic trees of bacteria MS-05 (MEGA-X). [Click here to view] |
3.3. Anti-Fungal Assay
The antagonistic potential of P. fluorescens MS-05 was evaluated against the fungal deteriorating agent AN-03 (Aspergillus niger, NCBI Accession No. OR083359) using a dual-culture antifungal assay. The assay was performed in triplicate (n = 3), and the inhibition zone was measured after 7 days of incubation. The mean zone of inhibition recorded for P. fluorescens MS-05 was 21.3 ± 1.2 mm, whereas the control plate (fungus only) showed unrestricted mycelial growth. Observations taken at 10, 20, and 30 days revealed complete growth suppression of A. niger in plates treated with the bacterial supernatant, while minimal growth was observed in plates treated with bacterial broth [Table 3]. These results demonstrate that the cell-free supernatant of P. fluorescens MS-05 exhibits superior antifungal efficacy compared with the bacterial suspension. The efficacy observed in this study aligns with earlier reports of Pseudomonas spp. producing DAPG, phenazines, pyoluteorin, and pyrrolnitrin that inhibit fungal growth and promote plant health [24-26]. Comparable inhibition was reported for P. protegens ML15 [27,28] and P. chlororaphis KNU17Pc1 [36]. P. fluorescens is widely recognized as an effective biocontrol agent, and the present findings further support its ability to suppress storage-associated fungal pathogens. A representative image of the dual-culture assay showing inhibition of A. niger by P. fluorescens MS-05 is presented in Figure 2.
Table 3: Dual culture assay on PDA for detection of antagonistic activity of bacteria against deteriorating fungal pathogen in Mm, ± standard deviation (SD) from three independent replicates (n=3).
| Treatment | 2nd day (mm) | 5th day (mm) | 7th day (mm) | Mean zone of inhibition (mm±SD, n=3) |
|---|---|---|---|---|
| Plate A | 15.3 | 26.4 | 56.3 | 21.3±1.2 |
| Plate B | 12.5 | 27.2 | 48.2 | 20.8±1.4 |
| Plate C | 14.2 | 29.3 | 52.6 | 21.5±1.0 |
| Control (fungus only) | – | – | – | 0.0 |
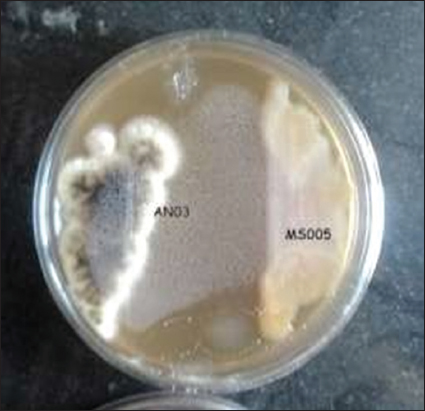 | Figure 2: Dual culture assay Aspergillus niger versus Pseudomonas fluorescens MS-05. [Click here to view] |