1. INTRODUCTION
Food-borne infections caused by Salmonella enterica subsp. enterica (salmonellosis) represent a major public health challenge worldwide. Salmonellosis causes substantial morbidity and mortality throughout the world. Approximately 200,000 people die of typhoid globally each year, and most of these mortalities occur in the developing countries [1,2]. In addition, there has been a dramatic increase in the number of reported cases of salmonellosis in the developed countries, due in part to travelers visiting endemic regions [3]. More than 2,500 S. enterica strains have been reported worldwide based on their antigenic properties, and most of these strains cause either typhoidal or non-typhoidal salmonellosis [4,5]. Typhoidal salmonellosis is caused by S. enterica serotypes Typhi and Paratyphi A, B, and C, causing enteric fever, gastroenteritis, and bacteraemia, whereas non-typhoidal salmonellosis is caused by S. enterica serotypes Enteritidis and Typhimurium, causing a variety of diseases ranging from mild/severe gastroenteritis to systemic infection in humans and animals [6–8].
The wide range of Salmonella infections is attributed to the presence of numerous virulence factors that play essential roles in Salmonella pathogenesis, including in host cell adhesion, invasion, and intracellular replication [9]. Most of the genes encoding virulence factors are located in the regions known as Salmonella pathogenicity islands (SPIs) in the Salmonella chromosome, two of which (SPI-1 and SPI-2) encode a type three secretion system (T3SS) that is involved in the invasion of host intestinal cells and intercellular survival. The two main regulatory proteins for T3SS function are HilA and InvA, which play important roles in the invasion of epithelial cells and macrophage apoptosis [10]. In addition, fimbriae, encoded by several putative fimbrial operons, such as agf, fim, and sef, play a significant role in Salmonella pathogenicity by binding to specific receptors on the host cells, leading to bacterial colonization, and/or toxin delivery [11,12]. In addition, Salmonella is one of the causative agents of diarrhea due to its ability to produce enterotoxin via a process mediated by the stn gene [2,12]. Additionally, the misL gene that encodes MisL, an auto-transporter protein, is an extracellular matrix adhesin that is involved in the invasion process and intestinal colonization, leading to systemic infection [2,12].
In the recent years, there has been a continuous emergence of multidrug-resistant Salmonella strains; therefore, the development of an effective vaccine for systemic Salmonella infections has become of a great importance for the control of these global food-borne pathogens [13,14]. Vaccination is one of the most effective approaches for infectious disease control and can successfully protect humans against various diseases. Currently, there are two approved vaccines against typhoid Salmonella, namely, the Vi polysaccharide vaccine and the live attenuated Salmonella Typhi Ty21a strain vaccine. However, there is no licensed vaccine against non-typhoid Salmonella [15,16]. Among the components of Salmonella that may be possible candidates for vaccine development are the cell outer membrane proteins (OMPs), which play an essential role in adaptation to environmental conditions, motility, adherence, and host cell colonization. Furthermore, OMPs play a significant role in the injection of toxins and cellular proteases, as well as the formation of channels for the removal of antibiotics (efflux pumps) [5,14,17]. The aim of the current work included determination of various virulence genes in S. enterica strains (n = 33) isolated from patients in Saudi Arabia [18] and investigation of the immunogenicity of the OMPs of S. enterica serovar Typhimurium as a vaccine candidate against typhoid in experimental animals.
2. MATERIALS AND METHODS
2.1. Microorganisms and Growth Culture
The various S. enterica strains studied (n = 33) were previously isolated from clinical and environmental sources (Riyadh, Saudi Arabia) and assigned the accession numbers KU843835–KU843866 [18]. Bacterial growth was carried out on the media recommended for Salmonella spp. including xylose lysine deoxycholate agar (Oxoid, UK) and deoxycholate citrate agar (Oxoid) for 24 hours at 37°C.
2.2. PCR-Based Detection of Virulence Genes
The isolates of S. enterica (n = 33) were screened for seven virulence genes by Polymerase Chain Reaction (PCR) according to the methods reported previously with some modification using the set of specific primers shown in Table 1 [2]. The tested virulence genes included aggregative fimbriae (agfA), fimbrial proteins (sefC, fimA), Salmonella pathogenicity island-3 (misL), Salmonella pathogenicity island-1 (hilA), and Salmonella specific invasive (invA) and enterotoxic (stn) genes. The genomic DNA of the S. enterica isolates (n = 33) was extracted from overnight cultures using the DNeasy Blood and Tissue Kit (Qiagen, UK) following the manufacturer’s instructions. The PCRs were performed in a final volume of 25 μl, containing 12.5 µl of GoTaq® Green Master Mix (Promega), 2 µl of DNA (50 µg/ml), 2 μl of forward and reverse primers, and 6.5 µl of nuclease-free water (Promega, UK). The PCR was carried out under the following conditions: initial denaturation for 5 minutes at 95°C; 35 cycles of denaturation at 95°C for 30 seconds, annealing at a specific annealing temperature (Table 1) for 30 seconds, and extension at 72°C for 1.5 minutes; and a final extension step at 72°C for 5 minutes. The PCR products were analyzed by 1.5% (w/v) agarose gel electrophoresis in 1× TAE (40 mM Tris, 20 mM acetic acid, and 1 mM Ethylenediaminetetraacetic acid (EDTA), pH 8.0) at 90 V for 45 minutes using a 100-bp DNA ladder (Qiagen, UK) as the molecular size standard. DNA from Escherichia coli was used as a negative control.
2.3. Preparation of Outer Membrane Proteins
2.3.1. Extraction of outer membrane proteins
The OMPs of S. enterica serovar Typhimurium strain SA3 (accession no. KU843837) were extracted according to a previously reported method with some modifications [19]. Briefly, 1 g (wet weight) of Salmonella Typhimurium cell biomass was re-suspended in 20 ml of extraction buffer [10 mM Tris-HCl (pH 7.5), 10 mM EDTA disodium salt and 6 M urea], and stirred for 1 hour at 4°C. The cell extract was dialyzed against distilled water for 48 hours at 4°C with frequent changing of the water (every 6 hours). The dialyzed extract was centrifuged at 6,000 rpm for 1 hour at 4°C, and the supernatant was collected and lyophilized (LABCONCO, Czech Republic). The crude OMPs were stored at −75°C until use.
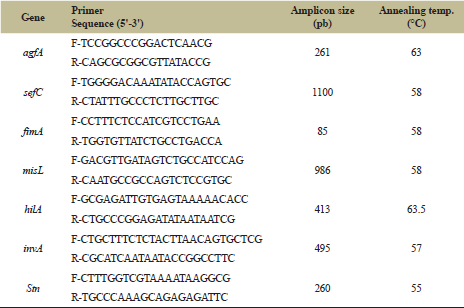 | Table 1: Primers sequnces for various virulence genes of Salmonella enterica isolates [2]. [Click here to view] |
2.4. Estimation of Protein Concentration
The protein concentration was determined according to the method described by Bradford [20]. One milliliter of Bradford reagent was added to 50 μl of sample, and the optical density was measured after 5 minutes at 595 nm. Bovine serum albumin (50–1,000 µg/ml) was used as the protein standard.
2.5. Polyacrylamide Gel Electrophoresis
The extracted OMPs were separated by sodium dodecyl sulfate (SDS)-Polyacrylamide Gel Electrophoresis (PAGE) according to the Laemmli [21] method using a mini gel electrophoresis system (Biometra, UK). The protein samples were solubilized in sample loading buffer [0.625 M Tris-HCl (pH 6.8), 2% SDS, 10% glycerol, and 5% β-mercaptoethanol] at 100°C for 5 minutes. SDS-PAGE was carried out on a 12% separating gel with a discontinuous buffer system. Thereafter, the gels were stained by Coomassie blue staining solution for 1–2 hours with continuous shaking. The gel was transferred to destaining solution and subjected to shaking with continuous changing of the solution until distinct protein bands were visible.
2.6. Purification of OMPs
Protein-acrylamide (PA) complexes were prepared according to the method of Hamid and Jain [19]. The major OMPs extracted from S. enterica Typhimurium strain SA3 were separated by 12% SDS-PAGE, and six dominant protein bands with molecular sizes of 27, 29, 38, 48, 64, and 70 kDa were selected and used as eliciting antigens. The bands corresponding to the selected proteins were excised from the gels using a sterile blade, transferred to a 50-ml centrifuge tube, washed twice with distilled water, and pulverized using a small mortar and pestle. The pulverized materials containing proteins and acrylamide were lyophilized to form a fine powder and kept at −75°C until further use.
2.7. Mouse Husbandry and Immunization
BALB/c female mice, 6–8 weeks old (20–25 g), were obtained from the Central Animal House at the College of Pharmacy (King Saud University). The mice were housed in clean, well-aerated polypropylene cages at 25°C and fed a pellet diet and water. The mice immunization schedule was performed according to Hamid and Jain [19] and Yang et al. [16]. The experiment was carried out using a total of 56 mice that were divided into 14 groups (n = 4 mice/group). Twelve groups were immunized subcutaneously with the six selected proteins (two mouse groups for each OMP), and two groups of animals served as controls (Supplementary Data, S1). Prior to immunization, the mice were bled from the tail veins to obtain pre-immune sera. The prepared PA complex powder was suspended in phosphate-buffered saline (PBS) and emulsified with an equal volume of complete Freund’s adjuvant to obtain a final protein concentration of 25 µg protein/0.5 ml of emulsion. Two mice from each group received four subcutaneous injections of the PA complex (25 µg of protein/dose) on days 0, 7, 21, and 28 and a final booster dose on day 40. Two mice in each group acted as controls and received lyophilized polyacrylamide (without protein) dissolved in PBS under the same immunization schedule. In addition, two mouse groups (n = 8) did not receive any injection and acted as healthy control B. Thereafter, the mice were bled on day 50 to obtain immune sera and kept at −75°C until use.
2.8. Determination of Bacterial Lethal Dose (LD50)
The 50% lethal dose (LD50) of S. enterica serovar Typhimurium strain SA3 was determined according to the previously reported methods [22,23]. Serial dilutions of overnight cultures of strain SA3 were prepared, and cell suspensions with cell counts ranging from 2 × l04 to 2 × l09 CFU/ml were orally administered to six groups of mice (8 mice/group). The animals were monitored for 14 days, and the lethal effect was recorded. The LD50, which was defined as the bacterial dose that led to the death of 50% of the mice, was estimated based on the number of surviving mice on day 14.
2.9. Animal Challenge with Salmonella Typhimurium SA-3
After 4 weeks of immunization with the PA complexes, the immunized and control mice were injected intraperitoneally with two different doses of S. enterica Typhimurium SA3, including 50× and 100× LD50. In addition, five mice were injected with PBS as a control. After the bacterial challenge, the animals were monitored for 14 days, and their clinical symptoms and mortality rate were recorded. Bacterial infection was confirmed by re-isolation of S. enterica Typhimurium SA3 from the infected mice. Briefly, livers and spleens were collected from the infected and control mice under aseptic conditions on day 2 post-infection. Then, the livers and spleens were homogenized in sterile phosphate buffer, and aliquots were inoculated into selenite F broth and incubated for 18 hours at 37°C. Thereafter, a loopful of each culture was transferred to nutrient agar and incubated under the same conditions. The obtained bacterial colonies were identified by a Vitek® 2-C15 automated system (BioMerieux Inc., France) and 16S rDNA sequencing analysis as previously described [18].
2.10. Histopathological Examination
Histopathological changes in the mouse organs caused by Salmonella Typhimurium SA3 were investigated. The spleens and livers were removed from the infected and control mice, fixed in formalin solution (10%), and kept in paraffin blocks. Transverse sections (5 µm) were prepared using a microtome, stained with eosin and haematoxylin (H/E), and examined by microscopy [16].
2.11. Detection of OMP-Specific Antibodies
The humoral immune response elicited by OMPs in the immunized animals was detected using immunodiffusion [24]. Briefly, 1% (w/v) melted agarose solution, prepared in sterile PBS, was poured into sterile Petri dishes (≈2-mm thick). After gel solidification, wells were punched in the gels using a sterile cork borer. Then, 50 µl of the crude extracts containing Salmonella Typhimurium SA3 OMPs were loaded in the central wells, while the surrounding wells were loaded with 50 µl of different sera obtained from mice immunized with different OMPs (27, 29, 38, 48, 64, and 70 kDa OMP). Thereafter, the plates were incubated at 4°C for 24–48 hours and monitored for agglutination.
3. RESULTS AND DISCUSSION
The emergence of multidrug resistance in S. enterica, the causative agent of salmonellosis, has led to further investigation of the virulence factors of this organism and to the search for an effective control method [25,26]. Consequently, this study aimed to identify the various virulence factors of S. enterica strains isolated in Saudi Arabia, in addition to preliminary identification of vaccine candidates against S. enterica serovar Typhimurium based on the OMP subunits.
3.1. Detection of Virulence Genes of S. enterica Isolates
The degree of pathogenicity of S. enterica depends on several virulence factors that are encoded by various genes, including the fimbriae operon, virulence plasmid, SPIs, and enterotoxin gene. However, the association of some of these factors with pathogenicity remains controversial [27]. In the present study, the presence of putative virulence genes (n = 7) responsible for the pathogenicity of S. enterica (n = 33) isolated from patients in Saudi Arabia was detected by PCR using a set of primers specific to various virulence genes. The aggregative fimbriae (agfA) gene was detected, with the expected PCR product size of approximately 261 bp (Fig. 1), in most S. enterica strains (n = 32/33), being absent in only S. enterica serovar Paratyphi C strain SA7. As shown in Figure 2, the sefC (fimbrial protein) gene was amplified with the correct amplicon size of 1,100 bp in 81.8% of the isolates (27/33). Fimbriae play a role in Salmonella pathogenicity by binding to specific receptors of host cells, leading to bacterial colonization, and/or toxin delivery, which is reflected in the appearance of infection [28,29]. The high frequency of agfA was similar to the results of previous studies on different Salmonella serovars [25,28]. In addition, fimbrial protein SEF14, which encodes the sef operon, was detected in only Salmonella Enteritidis and Salmonella Paratyphi B isolates. This finding is consistent with the results of previous studies, providing a useful marker to discriminate S. Enteritidis from Salmonella Typhi as well as Salmonella Paratyphi B from Salmonella Typhimurium isolates [30]. Salmonella strains express type-1 fimbriae that enable the bacteria to bind to eukaryotic cells. Fimbrial proteins are encoded by the fim gene. In this study, all the isolates (n = 33) were found to harbor the fimA gene (S1), which consistent with previous reports that studied the prevalence of the fimA operon in various Enterobacteriaceae strains [28,30].
 | Figure 1: Agraose gel electrophoresis of the amplified agf gene. Fifteen microliter of the PCR product was separated in 1% agarose containing ethidium bromide solution (1 µg/ml), and visualized using Gel documentation system. M: 100 bp ladder; Neg: negative control; Other lanes: S. enterica strains (n = 33). [Click here to view] |
On the other hand, all the tested S. enterica isolates (n = 33) were found to harbor hilA (Salmonella pathogenicity island-1), invA (Salmonella specific invasive), stn (enterotoxic), and misL (Salmonella pathogenicity island-3), regardless of serovar and the source of the isolate, yielding PCR amplicons of the expected sizes of 413, 495, 260, and 986 bp, respectively (Supplementary Data S2–S5). All these genes exhibited a high degree of conservation among Salmonella strains, but are absent among other closely related enteric bacteria [2,6,11] (Fabián et al., 2012).
3.2. Immunogenicity of OMPs
The OMPs of Gram-negative bacteria can elicit a significant antibody response from the host defense system. However, there is relatively limited information regarding the potential of OMPs to confer protection against Salmonella infections [14,31]. Moreover, while there are two approved vaccines against typhoid Salmonella, including the Vi polysaccharide vaccine and a live attenuated Salmonella Typhi Ty21a strain vaccine, there is no licensed vaccine against non-typhoid Salmonella [15,30].
 | Figure 2: Agraose gel electrophoresis of the amplified sefC gene. Fifteen microliter of the PCR product was separated in 1% agarose containing ethidium bromide solution (1 µg/ml), and visualized using Gel documentation system. M: 100 bp ladder; Neg: negative control; Other lanes: S. enterica strains (n = 33). [Click here to view] |
3.3. Preparation of OMPs of Salmonella Typhimurium SA3
The OMPs of Salmonella Typhimurium SA3 were extracted using the urea-EDTA extraction method. The urea extraction method has been proven to be effective for solubilization of the surface proteins of a number of bacterial species and has yielded favorable results [19]. The SDS-PAGE analysis pattern showed approximately 20 proteins with molecular masses ranging from approximately 15–95 kDa (Fig. 3). Six of the most dominant OMPs of S. enterica Typhimurium SA3 were selected for the immunization studies, including proteins with molecular masses of 27, 29, 38, 48, 64, and 70 kDa.
3.4. Mouse Protection Studies
The roles of the selected OMPs in providing protection against Salmonella infection in experimental mice were investigated. The selected OMP bands were excised from the PAGE gel, processed to obtain pulverized PA complexes, and then used to immunize mice. Thereafter, the mice immunized with various OMPs were challenged with two different doses of S. enterica Typhimurium SA3, including 50× and 100× LD50, and monitored for the appearance of infection symptoms and/or mortality in comparison with the control mice. For confirmation of infection of the mice with Salmonella Typhimurium SA3, the bacteria were re-isolated from livers and spleens collected from infected and control mice. The obtained bacteria were identified by biochemical tests and 16S rDNA sequencing.
 | Figure 3: SDS-PAGE (12%) analysis of urea-extracted OMPs from S. enterica seroval Typhimurium (isolate SA-3). Lane M, molecular weight markers. The sizes of the marker bands are shown at the left. Other lanes: OMPs fractions. [Click here to view] |
The results revealed the isolation of Salmonella Typhimurium SA3 from the infected and control mice, indicating the success of the challenge experiment. The animals were monitored for up to 14 days post-infection, and their clinical symptoms and mortality rate were recorded. The results indicated that after 1 day of infection, the control mice (n = 4) showed symptoms of salmonellosis, including skin pustules, sluggishness, ruffled hair, slow or no responsiveness, and loss of appetite, and these mice died within 3–5 days (Supplementary Data S7). However, immunization of the mice with various OMPs led to the protection of the mice to varying degrees (Table 2). Immunization of the mice with the 38-kDa OMP resulted in survival of 100% and 75% of the mice challenged with doses of 50× LD50 and 100× LD50 of Salmonella Typhimurium strain SA3, respectively. In addition, immunization with both the 64- and 70-kDa OMPs resulted in the protection of 66.7% of the mice challenged with a dose of 50× LD50. The 48-kDa protein provided 75% and 50% protection under similar conditions, whereas the 27- and 29-kDa proteins exhibited 75% and 66.7% protection, of mice injected with doses of 50× and 100× LD50, respectively. The lowest protection (33.3% of the mice) was observed in animals immunized with the 64-kDa OMP.
This strong protection of the immunized mice by the 38-kDa OMP, in comparison to the control, against Salmonella Typhimurium infection can be attributed to elicitation of specific and/or nonspecific immune responses [14]. In comparison to the results of other studies, the 38-kDa OMP provided effective and superior protection against Salmonella Typhimurium infection. A 43-kDa truncated OmpC pore protein showed 50%–75% protection against Salmonella Typhimurium infection in birds [32]. Recently, Toobak et al. [33] identified three porin OMPs from S. enterica serovar Typhi with similar molecular sizes of approximately 37.62 kDa (OmpA), 40.7 kDa (OmpC), and 39.05 kDa (OmpF); however, immunization of experimental mice with these proteins did not lead to protection against Salmonella infection [33]. However, a non-porin 19-kDa OMP investigated by Hamid and Jain [19] showed the increased protection (100%) of mice challenged with 100× LD50 of Salmonella Typhimurium. Immunization with the OMPs of Salmonella is considered to be a potential approach for conferring protection against typhoid. The OMPs of Gram-negative bacteria include a family of pore-forming proteins that form water-filled channels that allow small hydrophilic solutes to pass through the pore, in addition to other non-porin proteins that play significant roles in the pathogenicity of the organism [34,35].
 | Table 2: Results of mice immunization with various outer membrane proteins of S. enterica serovar Typhimurium SA3. [Click here to view] |
3.5. Detection of OMP-Specific Antibodies
The sera collected prior to and after the mice immunizations were evaluated for the humoral immune response elicited by OMPs using immunodiffusion (Fig. 4). The results indicated that the OMP with a molecular weight of 38 kDa stimulated a significant humoral response in mice. Sera from animals immunized with the 38-kDa OMP could be precipitated with the OMPs of S. enterica Typhimurium SA3. In addition, the antiserum of the OMP with a molecular weight of 29 kDa exhibited a faint precipitation line, whereas no reaction was observed with antisera obtained from other OMP groups. The serum from unimmunized animals did not show any precipitation with the extracted Salmonella Typhimurium OMPs.
3.6. Histopathological Studies
The protection efficiency of the 38-kDa OMP against Salmonella Typhimurium challenge in immunized mice was further evaluated based on the morphological analysis of the histopathological changes. The livers and spleens that were collected from the unimmunized infected control showed clear hepatomegaly and splenomegaly compared with those of uninfected mice (Fig. 5). The pathological study indicated that the H/E-stained liver and spleen sections of the control mice showed clinical signs of Salmonella infection, including focal necrosis of the liver, distinct splenorrhagia, and vacuolar degeneration of splenic reticular cells. In addition, the H/E-stained sections showed the presence of inflammatory cells in both the liver and spleen as well as abnormal structures such as vacuoles and hydropic degeneration (Figs. 6 and 7). On the other hand, the immunized mice showed standard healthy structures [16].
 | Figure 4: Gel immune-diffusion precipitation test showed precipitate line with OMP-38 antisera against OMPs mixture. [Click here to view] |
 | Figure 5: The spleen and Splenomegaly (A); and Liver and Hepatomegaly (B) of infected and uninfected mice. [Click here to view] |
 | Figure 6: Spleen section of healthy (control) (A) and infected (B) spleen section with inflammatory cells. [Click here to view] |
 | Figure 7: Liver section of healthy (control) (A) and infected mice (B). FN = focal necrosis, HD = hydropic degeneration, ICs = inflammatory cells around the blood vessels, V = blood vessel. [Click here to view] |
4. CONCLUSION
PCR-based detection of the virulence genes in S. enterica strains (n = 33) isolated from patients in Saudi Arabia revealed slight variations in gene distribution among the isolates, particularly for the aggregative fimbriae gene (agfA) and the fimbrial protein-encoding gene sefC. Investigation of the immunogenicity of OMPs of Salmonella Typhimurium SA3 revealed that several OMPs conferred varying degrees of protection against infection. Mice immunized with the 38-kDa OMP exhibited 100% protection when challenged with 50× LD50 of Salmonella Typhimurium SA3 and 75% protection against an even higher bacterial challenge of 100× LD50. This 38-kDa OMP is a promising potential vaccine against S. enterica serovar Typhimurium. Further research on the protein structure is in progress.
ACKNOWLEDGMENTS
The authors extend their appreciation to the Research Center at College of Science, Deanship of Scientific Research at King Saud University for funding this work.
REFERENCES
1. Lamas A, Miranda MJ, Regal P, Vazquez B, Franco MC, Cepeda A. A comprehensive review of non enterica subspecies of Salmonella enterica. Microbiol Res 2018;206:60. CrossRef
2. Mezal EH, Stefanova R, Khan AA. Isolation and molecular characterization of Salmonella enterica serovar Javiana from food, environmental and clinical samples. Int J Food Microbiol 2013;164(1):113–8. CrossRef
3. Xiong K, Chen Z, Zhu C, Li J, Hu X, Rao X, et al. Safety and immunogenicity of an attenuated Salmonella enterica serovar Paratyphi A vaccine candidate. Int J Med Microbiol 2015;305(6):563–71. CrossRef
4. Heredia N, García S. Animals as sources of food-borne pathogens: a review. Anim Nutr 2018;3:250–5. CrossRef
5. Liu Q, Liu Q, Yi J, Liang K, Liu T, Roland LK, et al. Outer membrane vesicles derived from Salmonella Typhimurium mutants with truncated LPS induce cross-protective immune responses against infection of Salmonella enterica serovars in the mouse model. Intern J Med Microbiol 2016;306(8):697–706. CrossRef
6. Abakpa GO, Umoha JV, Ameha BJ, Yakubua ES, Kwagab PKJ, Kamaruzamanc S. Diversity and antimicrobial resistance of Salmonella enterica isolated from fresh produce and environmental samples. Environ Nanotech Monit Manag 2015;3:38–46. CrossRef
7. Ceyssens P-J, Matthews W, Vanhoof R, Bertrand S. Trends in serotype distribution and antimicrobial susceptibility in Salmonella enterica isolates from humans in Belgium, 2009 to 2013. Antimicrob Agents Chemoth 2015;59:544–52. CrossRef
8. Tanner JR, Kingsley RA. Evolution of Salmonella within host. Trends Microbiol 2018;26(12):986–98. CrossRef
9. Herhaus L, Dikic I. Regulation of Salmonella-host cell interactions via the ubiquitin system. Inter J Med Microbiol 2018;308(1):176–84. CrossRef
10. Zhang K, Griffiths G, Repnik U, Horner M. Seeing is understanding: Salmonella’s way to penetrate the intestinal epithelium. Inter J Med Microbiol 2018;308(1):97–106. CrossRef
11. Fowler CC, Galán JE. Decoding a Salmonella typhi regulatory network that controls typhoid toxin expression within human cells. Cell Host Microbe 2018;23(1):65–76. CrossRef
12. Kurtz JR, Gagging JA, McLachlan JB. Salmonella infection: interplay between the bacteria and host immune system. Immunol Lett 2017;190:42–50. CrossRef
13. Kuang D, Zhang J, Xu X, Shi W, Chen S, Yang X, et al. Emerging high-level ciprofloxacin resistance and molecular basis of resistance in Salmonella enterica from humans, food and animals. Inter J Food Microbiol 2018;280:1–9. CrossRef
14. Tennant SM, MacLennan CA, Simon R, Martin LB, Khan MI. Nontyphoidal salmonella disease: current status of vaccine research and development. Vaccine 2016;34(26):2907–10. CrossRef
15. Han Y, Liu Q, Williams S, Liang K, Li P, Cheng A, et al. A bivalent vaccine derived from attenuated Salmonella expressing O-antigen polysaccharide provides protection against avian pathogenic Escherichia coli O1 and O2. Infection 2018;36(8):1038–46.
16. Yang Y, Wan C, Xu H, Aguilar ZP, Tan Q, Xu F, et al. Identification of an outer membrane protein of Salmonella enterica serovar Typhimurium as a potential vaccine candidate for Salmonellosis in mice. Microbes Infect 2013;5:388–98.
17. Dougan G, John V, Palmer S, Mastroeni P. Immunity to salmonellosis. Immunol Rev 2011;240:196–210. CrossRef
18. El-Tayeb AM, Ibrahim AS, Al-Salamah1 AA, Almaary SK, Elbadawi BY. Prevalence, serotyping and antimicrobials resistance mechanism of Salmonella enterica isolated from clinical and environmental samples in Saudi Arabia. Brazil J Microbiol 2017;48:499–508. CrossRef
19. Hamid N, Jain S. Characterization of an outer membrane protein of Salmonella enterica serovar Typhimurium that confers protection against typhoid. Clin Vaccine Immunol 2008;15(9):1461–71. CrossRef
20. Bradford M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 1976;72(1):248–54. CrossRef
21. Laemmli UK. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 1970;227(5259):680–5. CrossRef
22. Cawthraw S, Pennings JLA, Hodemaekers HM, de Jonge R, Havelaar AH, Hoebee B, et al. Gene expression profiles induced by Salmonella infection in resistant and susceptible mice. Microbes Infect 2011;13(4):383–93. CrossRef
23. Dehghani B, Rasooli I, Gargari SL, Nadooshan MR, Owlia P, Nazarian S. Immunogenicity of Salmonella enterica serovar Enteritidis virulence protein, InvH, and cross-reactivity of its antisera with Salmonella strains. Microbiol Res 2013;168(2):84–90. CrossRef
24. Adamkiewic V, Peković D. Experimental pulpal Arthus allergy. Oral Surg Oral Med Oral Pathol 1980;50(5):450–6. CrossRef
25. Fardsanei F, Dallas SM, Douraghi M, Salehi ZT, Mahmoodi M, Memariani H, et al. Genetic diversity and virulence genes of Salmonella enterica subspecies enterica serotype Enteritidis isolated from meats and eggs. Microb Pathogen 2017;107:451–6. CrossRef
26. Fardsanei F, Dallas SM, Memariani HDM, Bakhshi B, Salehi ZT, Nikkhahi F. Antimicrobial resistance, virulence genes and genetic relatedness of Salmonella enterica serotype Enteritidis isolates recovered from human gastroenteritis in Tehran, Iran. J Global Antimicrob Resist 2018;12:220–6. CrossRef
27. Lopez EF, Pescaretti MM, Moreno R, Delgado AM. Salmonella Typhimurium general virulence factors: a battle of David against Goliath? Food Res Inter 2012;45:842–51. CrossRef
28. CrăciunaÅŸ C, Keul AL, Flonta M, Cristea M. DNA-based diagnostic tests for Salmonella strains targeting hilA, agfA, spvC and sef genes. J Environ Manage 2012;95 Suppl:S15–8. CrossRef
29. Turki Y, Ouzari H, Mehri I, Aissa BR, Hassan A. Biofilm formation, virulence gene and multi-drug resistance in Salmonella Kentucky isolated in Tunisia. Food Res Intern 2012;45:940–6. CrossRef
30. Andrews JR, Ryan ET. Diagnostics for invasive Salmonella infections: current challenges and future directions. Vaccine 2015;33:8–15. CrossRef
31. Karami A, Ahmad Z, Safiri Z, Plural F. Detection of Salmonella strain by rapid-cycle multiplex PCR. JJM 2011;4(2):91–8.
32. Prejit, Agarwal RK, Porteen K, Dubal ZB, Asha K, Shweta S, et al. Evaluation of recombinant outer membrane protein based vaccine against Salmonella Typhimurium in birds. Biologicals 2013;41(3):162–8. CrossRef
33. Toobak H, Rasooli I, Talei D, Jahangiri A, Owlia P, Darvish Alipour Astaneh S. Immune response variations to Salmonella enterica serovar Typhi recombinant porin proteins in mice. Biologicals 2013;41(4):224–30. CrossRef
34. Cho Y, Park S, Barate AK, Truong QL, Han JH, Jung CH, et al. Proteomic analysis of outer membrane proteins in Salmonella enterica Enteritidis. J Microbiol Biotechnol 2015;25(2):288–95. CrossRef
35. Jha R, Kumar A, Sabena A, Tamuly S, Saxena KM. Cloning, sequencing and in silico analysis of Omp C of Salmonella typhimurium. ISRN Vet Sci 2012;2012:512848. CrossRef
SUPPLEMENTARY MATERIAL
 | Supplementary Figure 1: Agraose gel electrophoresis of the amplified fimA gene. 15 µl of the PCR product was separated in 1% agarose containing ethidium bromide solution (1 µg/ml), and visualized using Gel documentation system. M: 100bp ladder; Neg: negative control; Other lanes: S. enterica strains (n=33). [Click here to view] |
 | Supplementary Figure 2: Agraose gel electrophoresis of the amplified misL gene. 15 µl of the PCR product was separated in 1.5% agarose containing ethidium bromide solution (1 µg/ml), and visualized using Gel documentation system. M: 100bp ladder; Neg: negative control; Other lanes: S. enterica strains (n=33). [Click here to view] |
 | Supplementary Figure 3: Agraose gel electrophoresis of the amplified hilA gene. 15 µl of the PCR product was separated in 2% agarose containing ethidium bromide solution (1 µg/ml), and visualized using Gel documentation system. M: 100bp ladder; Neg: negative control; Other lanes: S. enterica strains (n=33). [Click here to view] |
 | Supplementary Figure 4: Agraose gel electrophoresis of the amplified invA gene. 15 µl of the PCR product was separated in 2% agarose containing ethidium bromide solution (1 µg/ml), and visualized using Gel documentation system. M: 100bp ladder; Neg: negative control; Other lanes: S. enterica strains (n=33). [Click here to view] |
 | Supplementary Figure 5: Agraose gel electrophoresis of the amplified stn gene. 15 µl of the PCR product was separated in 2% agarose containing ethidium bromide solution (1 µg/ml), and visualized using Gel documentation system. M: 100bp ladder; Neg: negative control; Other lanes: S. enterica strains (n=33). [Click here to view] |
 | Supplementary Figure 6: Immunization of experimental mice with OMPs extracted from S. enterica Typhimurium SA-3. The experiment was carried out using a total of 56 mice that were divided into 14 groups (n=4 mice/group). Twelve groups were immunized subcutaneously with the selected six proteins, (two mice groups for each OMP), and two groups of animals were served as control. The prepared proteins-acrylamide (PA) complex powder was suspended in phosphate-buffered saline (PBS), emulsified with an equal volume of complete Freund’s adjuvant to give final protein concentration of 25 µg protein/0.5 ml emulsion. Two mice from each group received four subcutaneous injections of the PA complex (25 µg protein/dose) on days 0, 7, 21, 28 and a final booster dose on day 40. The control mice (n=2) in each group received lyophilized polyacrylamide (without protein) dissolved into PBS at the same immunization schedule. Control B mice didn't receive any injection. [Click here to view] |
 | Supplementary Figure 7: Shows the immunization experiments of Salmonella vaccine attempt (control) (A); infected mice with salmonellosis symptoms such as ruffled hair and Skin pustules (B, C & D); Dead mice (E & F). [Click here to view] |