1. INTRODUCTION
Antimicrobial peptides (AMPs) are small peptides typically consisting of 12–50 amino acids that possess a wide range of antibacterial and antifungal properties, making them valuable as functional ingredients in pharmacology and food products [1]. Bacterial resistance to AMP is very rare [2]. AMPs can penetrate microbial cell membranes through electrostatic interactions. This mechanism is detrimental to microorganisms [3]. AMPs can be used as preservatives and to enhance food safety [4-6].
One of the directions in food biotechnology is the use of peptides [7] to create food products with antimicrobial properties, including individual products of preventive personalized medicine. Personalized AMP prevention and treatment methods include profiles of microorganisms or their genetic variations used in the therapy of different diseases, using an individualized approach to patients [8,9].
Therefore, bioengineering improvement of AMP properties is a relevant area of research in biotechnology and genetic engineering. This can be achieved by the method of incorporation of biologically active target sequences – Molecular Peptide Transplantation (MPT) – “isolation/synthesis of a bioactive fragment of a peptide/protein, its subsequent transfer into a target protein/peptide to create a new protein product with given unique technological and biological properties” [10,11].
Advances in bioengineering have made it possible to create modified peptides with unique effects. In particular, there are known in silico methods and models that are suitable for developing modified peptides with antibacterial effects. Agrawal and Raghava determined that chemical modifications increase the antibacterial effect [12]. Using structural analysis methods, Agrawal and Raghava proved that antibacterial modified peptides contain not only amino acid residues, but also various halogens, phosphoric acid residues, etc. [12]. Using chemical modifications, Wang created a range of AMPs [13]. Thus, the validity and necessity of applying computational methods for predicting and modeling various biologically active substances, pharmaceutical substances, and nutrients are emphasized once again [14].
The use of modeling allows scientists to reduce screening time and costs. Using models, structural modifications of AMPs with high antibacterial activity can be designed to increase their efficacy and predict their possible toxicity, since some peptides can be cytotoxic, destroying body cells. The side effects of AMPs are predicted using the quantitative structure-activity relationship (QSAR) model, which affects the design of effective and safe peptides [15]. Peptide design utilizes cyclic peptide scaffolds that are resistant to proteolysis and temperature effects. Such peptides are referred to as next-generation peptides [16,17].
The study aimed to perform molecular dynamics modeling (MDM), synthesis by MPT, and evaluation of the in vitro efficacy of the obtained new-generation AMP.
2. MATERIALS AND METHODS
2.1. Research and Analytical Equipment
Unless otherwise specified, analytical equipment from the following manufacturers was used in this research.
A high-performance liquid chromatograph, SHIMADZU LC20AD (SHIMADZU, Japan), was used to purify the synthesized peptide. A Matrix-Assisted Laser Desorption/Ionization Time-of-Flight Mass Spectrometry (MALDI-TOF MS) BRUKER Autoflex Speed (Bruker, Germany) ultra-flexible mass spectrometer was used for the molecular distribution of amino acid sequences.
A Tuttnauer autoclave-steam sterilizer 3850 ML (Tuttnauer, Israel) was used to sterilize laboratory ware and culture media. The following equipment was used to cultivate test strains and test antimicrobial activity using the disc diffusion method: BAVp-01 Laminar-S-1.2 microbiological safety cabinet (Laminar Systems, Russia), TS-1/20 air thermostat (Smolensk SKTB SPU, Russia). A KFK-3-01 spectrophotometer (Zagorsk Optical-Mechanical Plant, Russia) was used to study the concentration of the microbial suspension.
For internal water purification, a Millipore Elix Essential 3 air conditioning and ultraviolet water purification system (resistance 15 MΩ/cm, Merck Millipore, USA) was used.
2.2. Selection and Characterization of Proteolysis-resistant Peptide
The Cybase cyclic peptide database (https://www.cybase.org.au/) was used to search for proteolysis-resistant gastrointestinal (GI) and heat-treatment-resistant cyclic peptides suitable for MPT antimicrobial active sequences.
It has become widely used because it does not contain amino acids capable of forming disulfide bridges and contains only a small amount of amino acid residues. This allows a new short AMP to be synthesized on its basis, which is resistant to proteolysis and thermal treatment.
The PLP-5 peptide belongs to the orbitides, which are homodimeric cyclic peptides (i.e., cyclic peptides containing only standard amino acids) without disulfide bonds consisting of 5–12 amino acid residues: glycine, leucine, phenylalanine, valine, and aspartic acid (GLFVD); has three hydrophobic amino acids: valine, leucine, and phenylalanine; has a negative charge of 1; and has a molecular mass of 549.626 Da [18].
AMP activity was proved in silico using MDM [19,20]. According to [21-23], AMPs designed by machine learning include QL, SN, LS, YL, WW, and LW sequences and have antimicrobial properties. In this context, we used MPT to attach QL, SN, YL, and WW sequences to the PLP-5 scaffold and obtained a new peptide, QLSNGLFVDYLWW.
2.3. Production of an AMP
A novel AMP was prepared by MPT of antimicrobial sequences QL, SN, YL, and WW to the PLP-5 peptide scaffold and was synthesized at Pepmic Co., Ltd (Suzhou, China) by solid-phase synthesis followed by high-performance liquid chromatography purification on a SHIMADZU chromatography column (SHIMADZU, Japan). The molecular distribution of amino acid sequences was determined using a MALDI-TOF MS Ultraflex mass spectrometer (Bruker, Germany).
2.4. Identification of an AMP
The MDM and identification of the novel AMP were performed using the publicly available peptide database PeptideAtlas (USA). Mass spectrometer output files were collected for human, mouse, yeast, and several other organisms and reviewed using the latest search engines and protein sequences. All sequence and spectral library search results are then processed using the Trans-Proteomic Pipeline. Identification results are provided on the PeptideAtlas website (https://peptideatlas.org/#). MDM included evaluation (prediction) of antigenicity (allergenicity), biological activity (antimicrobial), and QSAR (structure, physicochemical properties – activity, metabolism, bioavailability, etc.).
2.5. Evaluation of Peptide Antigenicity
The antigenicity (allergenicity) of the peptide was evaluated using the program http://imed.med.ucm.es/Tools/antigenic.pl and the openly available online tool AllerTOP (https://www.ddg-pharmfac.net/). Using two resources enabled cross-checking and more reliable results.
2.6. Prediction of Antimicrobial Activity
The antimicrobial activity of the synthesized peptide was predicted using the Peptide Ranker platform (http://distilldeep.ucd.ie/PeptideRanker). QSAR was evaluated using the Antimicrobial Peptide Database (APD) on AMPs (https://aps.unmc.edu/home).
2.7. Prediction of Toxicity and Bioavailability of a New Peptide
The toxicity, biological activity, metabolism, and bioavailability of the new peptide were predicted using the absorption, distribution, metabolism, excretion, and toxicity (ADMET) platform (https://admetmesh.scbdd.com/).
2.8. Confirmation of Peptide Antimicrobial Activity
2.8.1. Justification for the selection of test strains of microorganisms
Four strains of microorganisms were used as research objects: Staphylococcus aureus, Pseudomonas aeruginosa, Saccharomyces cerevisiae, and Aspergillus niger (Kurchatov Institute Research Center, Russia).
The choice of test strains of microorganisms is based on the following. S. aureus is a Gram-positive bacterium that, under certain conditions, can cause various skin diseases, sepsis, pneumonia, etc. P. aeruginosa is a conditionally pathogenic Gram-negative bacterium. A weakened immune system can cause various diseases of the GI tract, skin, urinary tract, etc. S. cerevisiae is a single-celled fungus and is a model organism in science. When a person’s immunity is weakened, they can cause endocarditis, pneumonia, and fungemia. A. niger is a mold fungus that causes aspergillosis, an opportunistic fungal infection.
2.8.2. Study of the antimicrobial activity of a new peptide
The antimicrobial activity of the peptide was confirmed in the experiment using the disk diffusion method strictly as described in the papers [24,25].
Commercial Lysogeny Broth medium (Servicebio, China) was used to cultivate test strains of S. aureus, P. aeruginosa, and Saccharomyces cerevisiae. For the cultivation of A. niger test strains, commercial Czapek nutrient medium (Servicebio, China) was used. Test strains were cultivated on a nutrient medium in a TS-1/20 air thermostat (Smolensk SKTB SPU, Russia) at the appropriate temperature (for S. aureus, P. aeruginosa, and S. cerevisiae, the cultivation temperature was 37°C, and for A. niger, it was 25°C). Test strains were cultivated to a microbial suspension concentration of 1.5–108 colony-forming units/mL.
A microbial suspension was applied to Petri dishes containing the appropriate culture medium. Once the microbial suspension had been absorbed into the agarized culture medium, paper discs (5 mm in diameter) were applied. Next, the analyte peptide was applied to the disc at a concentration of 30 μg/disc. The analyte peptide was first suspended in a phosphate-saline buffer solution (pH = 7.4).
For comparison, kanamycin (Sintez, Russia) was used at a dose of 30 μg/disc for bacteria, fluconazole (Sintez, Russia) at a dose of 25 μg/disc for mold, and 10 μg/disc for yeast.
Next, the Petri dishes were covered and placed in a TS-1/20 air thermostat (Smolensk SKTB SPU, Russia) for cultivation. Cultivation was carried out for 48 h at the appropriate temperature (for S. aureus, P. aeruginosa, and S. cerevisiae, the cultivation temperature was 37°C, and for A. niger, it was 25°C). At the final stage, the diameter of the lysis zone was measured.
3. RESULTS
3.1. The Results of the Identification of the Peptide
When identifying the QLSNGLFVDYLWW peptide according to the PeptideAtlas database, no matches were found with the known peptides available in this database. Therefore, the QLSNGLFVDYLWW peptide can be considered unique and identified by its peptide sequence.
3.2. Predicting Allergenicity of the Peptide
In predicting allergenicity [Figure 1], the peptide was found to contain no allergenic determinants, indicating that it has no ability to cause allergy when ingested.
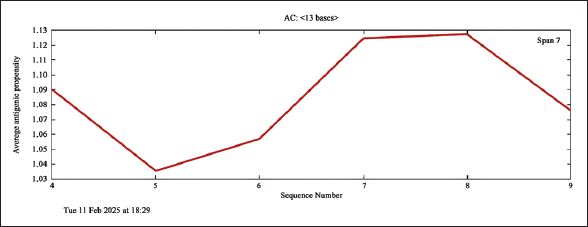 | Figure 1: The results of the prediction of the allergenicity of the peptide QLSNGLFVDYLWW. [Click here to view] |
3.3. Predicting the Biological Activity of the Peptide
When predicting the biological activity (antimicrobial) on the PeptideRanker platform, the activity coefficient of the peptide was found to be 0.82, which exceeds the threshold of 0.5 for biopeptides.
QSAR by the APD AMP database showed that the peptide QLSNGLFVDYLWW was hydrophobic (overall hydrophobicity ratio of 54%), had a weak negative charge (−1), molecular mass of 1640.861 Da, high protein binding potential (Boman index): −0.22 kcal/mol, can form alpha-helices, and has at least 5 residues on a hydrophobic surface.
Using the ADMET server, it was found that the developed peptide has a lipophilicity by logP of 1.043 units, the balance between lipophilicity and hydrophilicity by logD7.4 at 1.339 units, and an acid-alkaline dissociation constant (pKa) of 5. 641 units, the index of effective clinical use Fsp3 is more than 0.457 units, the index of distribution and metabolism in the body VDss is equal to 0.879 units, plasma clearance of CLplasma is 0.879 mL/min/kg, and half-life T1/2 averages from 4 to 8 h.
3.4. Peptides with Antibacterial Activity
To confirm the antibacterial activity of the investigated peptide, an in vitro experiment was performed. The results are presented in Table 1.
Table 1: Antimicrobial activity of the QLSNGLFVDYLWW peptide.
| Sample | Growth inhibition zone, mm | |||
|---|---|---|---|---|
| Staphylococcus aureus | Pseudomonas aeruginosa | Saccharomyces cerevisiae | Aspergillus niger | |
| QLSNGLFVDYLWW peptide | 28.5 | 19.5 | 45.0 | - |
| Control (fluconazole/kanamycin) | 27.5 | 23.0 | 16.5 | 15.0 |
Figure 2 shows the antimicrobial effect of the QLSNGLFVDYLWW peptide at an amount of 30 μg/disc against P. aeruginosa, S. cerevisiae, and S. aureus.
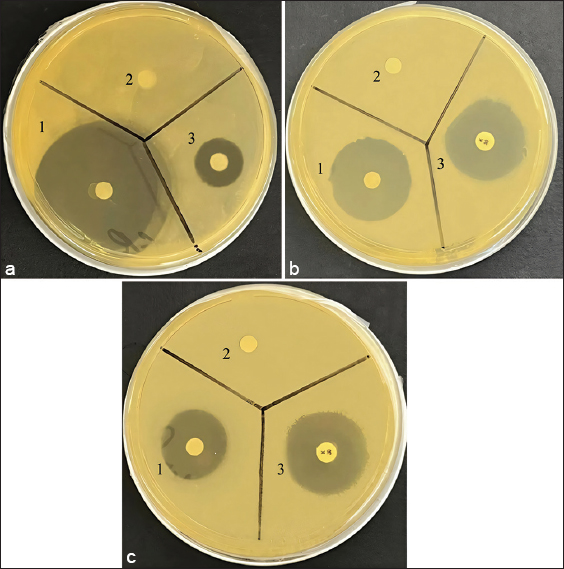 | Figure 2: Antimicrobial effect of the QLSNGLFVDYLWW peptide: (a) Saccharomyces cerevisiae; (b) Staphylococcus aureus; (c) Pseudomonas aeruginosa; 1 – a sample of the QLSNGLFVDYLWW peptide 30 μg/disc, 2 – negative control – phosphate-salt buffer; 3 – positive control (a – fluconazole 25 μg/disc; b – kanamycin 30 μg/disc). [Click here to view] |
The results clearly demonstrated the antimicrobial activity of the QLSNGLFVDYLWW peptide at a concentration of 30 μg/disc against Gram-positive and Gram-negative bacteria and yeast, which is comparable to the antibiotic kanamycin and the antifungal fluconazole.
The research focused on MDM, the synthesis of the QLSNGLFVDYLWW peptide, and the evaluation of its antimicrobial potential. Therefore, other biological activities were not studied, as this was beyond the scope of this research.
4. DISCUSSION
The results of these studies revealed that modern approaches can be used to construct a new low-molecular-weight peptide. Our conclusions and the correctness of our approach were also confirmed by our previously obtained results [26,27]. In particular, using the Enalos In Silico Nano platform, a unique peptide CTKSICTKKTLRTCPPIC [26] was designed at the preliminary stage, and the biological activity and properties of the unique peptide GRCTSICGPGPTCPPICFFD were predicted [27]. The results presented confirmed the reliability of our findings.
To use the peptide we developed and designed in the food industry, it was necessary to study its potential allergenicity. According to Hayes et al., the potential allergenicity of peptides can be studied not only in vitro and in vivo, but also using in silico methods [28]. The results of our studies, presented in Figure 1, indicate that the QLSNGLFVDYLWW peptide we synthesized does not contain amino acid sequences capable of causing an immunological response (allergic reactions). Using an in silico approach, it was studied and proven that four casein hydrolysate peptides are potential allergens [29]. The work of Lafarga et al. used this approach to show that out of 148 peptides, only four peptides were allergen-free [30]. These examples demonstrate the validity of the in silico approach for assessing allergenicity. Thus, we can conclude that our findings regarding the non-allergenicity of the new peptide QLSNGLFVDYLWW are reliable. Based on the in silico results, it is possible to decide on the need for additional in vitro and in vivo tests [28].
It should also be noted that computational and prediction methods are constantly evolving and allow scientists to select the composition of a biological molecule based on predictions of its biological activity [31,32]. This approach not only allows obtaining a new molecule with high speed and accuracy of prediction but also enables objective screening of therapeutic molecules [33]. It is worth noting that methods for designing new molecules and predicting therapeutic and other activities are based on analysis of the primary structure. Thus, Thomas et al. identified the antibacterial activity of the peptide based on its primary sequence, which was subsequently confirmed (qualitatively and quantitatively) using in vitro methods. It is also worth noting that when using virtual methods, statistical analysis is usually applied after forecasting, using regression methods to determine patterns that explain the biological activity of peptides [34]. In silico screening evaluates the reliability of the activity of each peptide, which is encoded by convenient variables that best reflect its functional characteristics. Classification or regression algorithms are used for analysis, allowing qualitative or quantitative differentiation of peptide activities [34].
Similar to the example above, using the in silico method, we were able to identify that the peptide QLSNGLFVDYLWW we designed had antimicrobial activity due to its ability to interact with components of the bacterial cell membrane. Using the APD database, we were able to identify that the constructed peptide QLSNGLFVDYLWW had a similarity of 41.18% to the AMP AP04690 and 38.89% to AP01225, whereas AP02378 had a similarity of 38.46%.
It should also be noted that the data obtained on the structure and properties of QLSNGLFVDYLWW AMP are consistent with the studies of [35], in which strategies, one of which was the creation of fully functional analogs of weakly anionic peptides capable of resisting proteases, were used. The P13#1 peptide mimics cathelicidins and has a high biological activity similar to the human cathelicidin LL-37 and shows no toxicity.
The antibacterial activity of the newly designed peptide QLSNGLFVDYLWW, predicted using in silico methods, was confirmed by the disc diffusion method [Figure 2 and Table 1]. It should be noted that our results are consistent with the literature data obtained by other scientists for antibacterial synthesized peptides. The synthesized peptides named DeNo1049, DeNo1057, and DeNo1051, with a concentration of 16 μg/mL, are active against S. aureus ATCC 29213 and with a concentration of 4–8 μg/mL against Escherichia coli ATCC 25922. Interestingly, the 30.0–50.0% similarity intervals with peptides from APD have a relatively high proportion of peptides with antimicrobial activity at 75.0%. Consequently, the QLSNGLFVDYLWW peptide falls within this interval, indicating the efficacy of the proposed strategy to design AMPs by MPT of antimicrobial sequences into a stable framework with preliminary MDM of peptide properties and scientific validation of antimicrobial properties on a structure-activity basis [23].
As for the mechanism of action of AMP QLSNGLFVDYLWW, in our opinion, it is not aimed at the destruction of the bacterial cell membrane, since the peptide is anionic and cannot interact with the entire surface of the negatively charged membrane, although it has an alpha-helix. It should be noted that most AMPs are mainly alpha-helical, cationic, and amphipathic, causing instability and degradation of cell membranes [36]. AMPs are more often active against Gram-negative bacteria; however, there are peptides (daptomycin) that have an increased effect against Gram-positive bacteria [37]. Glycopeptides inhibit peptidoglycan synthesis and are detrimental to Helicobacter pylori [38].
However, there are AMPs that do not act on the bacterial membrane but on the DNA and RNA of the cell [39].
Therefore, it can be assumed that the QLSNGLFVDYLWW peptide can utilize both mechanisms of antimicrobial action because its weak negative charge allows it to interact with certain positively charged regions of the membrane.
The antimicrobial mechanism of the peptide may be related to non-membrane targeting, which involves direct penetration of the peptide into the bacterial cell through the cell membrane by endocytosis. Thus, it affects the synthesis of RNA and DNA and affects important organelles of the bacterial cell. Such a mechanism of action of anionic AMPs has been described in [40].
By disrupting the spatial structure of nucleic acids and proteins, AMP QLSNGLFVDYLWW interferes with their synthesis. For example, the synthesis of AMPs of buforin II and indolicidin [40]. By forcibly targeting enzyme and protein synthesis along a particular pathway, nucleic acid and protein synthesis can be inhibited. Indolicidin inhibits double-stranded DNA by acting on type I DNA topoisomerase. Various AMPs affect DNA replication-participating enzymes, RNA polymerase, DNA gyrase, and other enzymes [40]. Some AMPs promote cell death and apoptosis through nucleic acid damage by disrupting signaling pathways. AMPs, by inhibiting protein synthesis, can affect ribosome translation [41]. The results have broader potential significance, as scientific articles on bacterial resistance to native peptide preparations (colistin or polymyxins) [42] and sometimes cross-resistance to multiple AMPs [43] are increasingly appearing, highlighting the need for new generation AMPs.
5. CONCLUSIONS
A new hydrophobic peptide (QLSNGLFVDYLWW) was developed using an in silico method. It was proven that the peptide does not cause allergies and has high antibacterial activity against both Gram-positive and Gram-negative bacteria, as well as yeast.
Based on the results obtained, it can be assumed that the synthesized peptide can be used as an antibacterial additive in the formulation of functional foods. However, synthesized peptides do not replace adequate nutrition but are only a supplement to the main diet.
6. AUTHORS’ CONTRIBUTIONS
All authors made substantial contributions to conception and design, acquisition of data, or analysis and interpretation of data; took part in drafting the article or revising it critically for important intellectual content; agreed to submit to the current journal; gave final approval of the version to be published; and agree to be accountable for all aspects of the work. All the authors are eligible to be author as per the International Committee of Medical Journal Editors (ICMJE) requirements/guidelines.
7. FUNDING
The study was supported by a grant from the Ministry of Science and Higher Education of the Russian Federation for major scientific projects in priority areas of scientific and technological development (No. 075-15-2024-483).
8. CONFLICT OF INTEREST
The authors report no financial or any other conflicts of interest in this work.
9. ETHICAL APPROVALS
This study does not involve experiments on animals or human subjects.
10. DATA AVAILABILITY
All supporting data are available through the corresponding author.
11. PUBLISHER’S NOTE
All claims expressed in this article are solely those of the authors and do not necessarily represent those of the publisher, the editors and the reviewers. This journal remains neutral with regard to jurisdictional claims in published institutional affiliation.
12. USE OF ARTIFICIAL INTELLIGENCE (AI)-ASSISTED TECHNOLOGY
The authors declare that they have not used artificial intelligence (AI)-tools for writing and editing of the manuscript, and no images were manipulated using AI.
REFERENCES
1. Wu Q, Patocka J, Kuca K. Insect antimicrobial peptides, a mini review. Toxins (Basel). 2018;10(11):461. [CrossRef]
2. Boparai J, Sharma P. Mini review on antimicrobial peptides, sources, mechanism and recent applications. Protein Pept Lett. 2020;27(1):4-16. [CrossRef]
3. Zhang LJ, Gallo R. Antimicrobial peptides. Curr Biol. 2016;26(1):R14-9. [CrossRef]
4. Jiang H, Liu J. Enhancing food safety with antimicrobial peptides in packaging materials. Food Control. 2023;148:108827.
5. Mann A, Nehra K, Rana JS, Dahiya T. Antibiotic resistance in agriculture:Perspectives on upcoming strategies to overcome upsurge in resistance. Curr Res Microb Sci. 2021;2:100030. [CrossRef]
6. Huan Y, Kong Q, Mou H, Yi H. Antimicrobial peptides:Classification, design, application and research progress in multiple fields. Front Microbiol. 2020;11:582779. [CrossRef]
7. Tikhonov SL, Chernukha IM. Cow milk polypeptide - a promising functional ingredient of specialized food products for the prevention of viral infections. Polzunov Bull. 2023;1:114-122. (Rus). [CrossRef]
8. Smith J, White P. Diagnostic applications of antimicrobial peptides:Current status and future directions. Diagn Microbiol Infect Dis. 2021;101(3):234-45.
9. Ulitina EA, Valieva SH, Tikhonov SL, Tikhonova NV. New antimicrobial food peptide:Characterization, properties and efficiency evaluation. Agrarn Nauka. 2024;4:132-7. (Rus). [CrossRef]
10. Camarero J, Campbell M. The potential of the cyclotide scaffold for drug development. Biomedicines. 2019;7(2):31. [CrossRef]
11. Wang CK, Craik DJ. Linking molecular evolution to molecular grafting. J Biol Chem. 2021;296:100425. [CrossRef]
12. Agrawal P, Raghava GPS. Prediction of antimicrobial potential of a chemically modified peptide from its tertiary structure. Front Microbiol. 2018;9:2551. [CrossRef]
13. Wang G. Post-translational modifications of natural antimicrobial peptides and strategies for peptide engineering. Curr Biotechnol. 2012;1:72-9. [CrossRef]
14. Dhanda SK, Usmani SS, Agrawal P, Nagpal G, Gautam A, Raghava GPS. Novel in silico tools for designing peptide-based subunit vaccines and immunotherapeutics. Brief Bioinform. 2017;18:467-78. [CrossRef]
15. Porto W, Pires A, Franco O. Computational tools for exploring sequence databases as a resource for antimicrobial peptides. Biotechnol Adv. 2018;36(1):221-33. [CrossRef]
16. Wang G, Li X, Wang Z. APD3:The antimicrobial peptide database as a tool for research and education. Nucleic Acids Res.2016;44:D1087-93. [CrossRef]
17. Qin L, Cui Z, Wu Y, Wang H, Zhang X, Guan J, et al. Challenges and strategies to enhance the systemic absorption of inhaled peptides and proteins. Pharm Res. 2022;40:1037-55. [CrossRef]
18. Arnison P, Bibb M, Bierbaum G, Bowers AA, Bugni TS, et al. Ribosomally synthesized and post-translationally modified peptide natural products:Overview and recommendations for a universal nomenclature. Nat Prod Rep. 2013;30:108-60. [CrossRef]
19. Beirigo PJ, Torquato HF, Santos CH, Carvalho MGD, Castro RN, Paredes-Gamero EJ, et al. [1-8-NaC]-Zanriorb A1, a proapoptotic orbitide from leaves of Zan-thoxylum riedelianum. J Nat Prod. 2016;79:1454-8. [CrossRef]
20. Richter A, Sutherland D, Ebrahimikondori H, Babcock A, Louie N, Li C, et al. Associating biological activity and predicted structure of antimicrobial peptides from amphibians and insects. Antibiotics (Basel). 2022;11(12):1710. [CrossRef]
21. Lin D, Sutherland D, Aninta SI, Louie N, Nip KM, Li C, et al. Mining amphibian and insect transcriptomes for antimicrobial pep-tide sequences with rAMPage. Antibiotics (Basel). 2022;11(7):952. [CrossRef]
22. Das P, Sercu T, Wadhawan K, Padhi I, Gehrmann S, Cipcigan F, et al. Accelerated antimicrobial discovery via deep generative models and molecular dynamics simulations. Nat Biomed Eng. 2021;5(6):613-23. [CrossRef]
23. Li C, Sutherland D, Richter A, Coombe L, Yanai A, Warren RL, et al. De novo synthetic antimicrobial peptide design with a recurrent neural network. Protein Sci. 2024;33(8):e5088. [CrossRef]
24. Babich O, Shevchenko M, Ivanova S, Pavsky V, Zimina M, Noskova S, et al. Antimicrobial potential of microorganisms isolated from the bottom sediments of lake Baikal. Antibiotics (Basel). 2021;10(8):927. [CrossRef]
25. Shevchenko M, Sukhikh S, Babich O, Noskova S, Ivanova S, Lisun V, et al. First insight into the diversity and antibacterial potential of psychrophilic and psychotrophic microbial communities of abandoned amber quarry. Microorganisms. 2021;9(7):1521. [CrossRef]
26. Tikhonov S, Tikhonova N, Timofeeva M, Shikhalev S. A new food peptide for the prevention of cell photoaging:Characterization and confirmation of effectiveness in vitro. Bull East Siberian State Univ Technol Manage. 2024;4(95):43-51. (Rus). [CrossRef]
27. Tikhonov S, Tikhonova N, Babich O, Sukhikh S, Sysuev E, Zvereva D. Development and prediction of the properties of a new biopeptide for dietary nutrition. Polzunovskiy Vestnik. 2025;3:31-8.
28. Hayes M, RougéP, Barre A, Herouet-Guicheney C, Roggen EL. In silico tools for exploring potential human allergy to proteins. Drug Discov Today Dis Models. 2015;17:3-11. [CrossRef]
29. Tu M, Liu H, Zhang R, Chen H, Fan F, Shi P, et al. Bioactive hydrolysates from casein:generation, identification, and in silico toxicity and allergenicity prediction of peptides. J Sci Food Agric. 2018;98(9):3416-26. [CrossRef]
30. Lafarga T, Wilm M, Wynne K, Hayes M. Bioactive hydrolysates from bovine blood globulins:Generation, characterisation, and in silico prediction of toxicity and allergenicity. J Funct Foods. 2016;24:142-55. [CrossRef]
31. Marks JR, Placone J, Hristova K, Wimley WC. Spontaneous membrane-translocating peptides by orthogonal high-throughput screening. J Am Chem Soc. 2011;133:8995-9004. [CrossRef]
32. Wold S, Jonsson J, Sjörström M, Sandberg M, Rännar S. DNA and peptide sequences and chemical processes multi variately modelled by principal component analysis and partial least-squares projections to latent structures. Anal Chim Acta. 1993;277:239-53. [CrossRef]
33. Wang P, Hu L, Liu G, Jiang N, Chen X, Xu J, et al. Prediction of antimicrobial peptides based on sequence alignment and feature selection methods. PLoS One. 2011;6(4):e18476. [CrossRef]
34. Thomas S, Karnik S, Barai RS, Jayaraman VK, Idicula-Thomas S. CAMP:A useful resource for research on antimicrobial peptides. Nucleic Acids Res. 2010;38:D774-80. [CrossRef]
35. Cafaro V, Bosso A, Di Nardo I, D'Amato A, Izzo I, De Riccardis F, et al. The antimicrobial, antibiofilm and anti-inflammatory activities of P13#1, a cathelicidin-like achiral peptoid. Pharmaceuticals (Basel). 2023;16.1386. [CrossRef]
36. Bin Hafeez A, Jiang X, Bergen PJ, Zhu Y. Antimicrobial peptides:An update on classifications and databases. Int J Mol Sci. 2021;22:11691. [CrossRef]
37. Zhang QY, Yan ZB, Meng YM, Hong XY, Shao G, Ma JJ, et al. Antimicrobial peptides:Mechanism of action, activity and clinical potential. Mil Med Res. 2021;8:48. [CrossRef]
38. Olleik H, Perrier J, Hijazi A, Baydoun E, Maresca M. Antimicrobial peptides and peptidomimetics as treatment option for Helicobacter pylori infection. In:Peptide and Protein Engineering for Biotechnological and Therapeutic Applications. Singapore:World Scientific;2023. 25-56. [CrossRef]
39. Kapil S, Sharma V. D-amino acids in antimicrobial peptides:A potential approach to treat and combat antimicrobial resistance. Can J Microbiol. 2020;67:119-37. [CrossRef]
40. Le C, Fang C, Sekaran S. Intracellular targeting mechanisms by antimicrobial peptides. Antimicrob Agents Chemother. 2017;61:e02340-16. [CrossRef]
41. Li X, Zuo S, Wang B, Zhang K, Wang Y. Antimicrobial mechanisms and clinical application prospects of antimicrobial peptides. Molecules. 2022;27:2675. [CrossRef]
42. Aghapour Z, Gholizadeh P, Ganbarov K, Bialvaei AZ, Mahmood SS, et al. Molecular mechanisms related to colistin resistance in Enterobacteriaceae. Infect Drug Resist. 2019;12:965-75. [CrossRef]
43. Fleitas O, Franco OL. Induced bacterial cross-resistance toward host antimicrobial peptides:A worrying phenomenon. Front Microbiol. 2016;7:381. [CrossRef]