1. INTRODUCTION
Candida albicans is a leading human fungal pathogen, responsible for nearly 70% of hospital-acquired fungal infections and ranked as the fourth most common cause of central line-associated bloodstream infections [1,2]. While C. albicans is the most frequently isolated species, other non-albicans Candida spp. (Candida parapsilosis, Candida tropicalis, Candida krusei, Candida glabrata, Candida dubliniensis, Candida lusitaniae, and the emerging Candida auris) also contribute to clinical infections, with varying antifungal resistance profiles [3]. The clinical impact of C. albicans is particularly severe in immunocompromised individuals, including patients undergoing chemotherapy, transplantation, and those suffering from acquired immunodeficiency syndrome [4].
The pathogenic success of C. albicans is linked to its ability to undergo morphological transitions and to metabolize diverse host nutrients. The yeast-to-hypha switch promotes tissue invasion, dissemination, and biofilm formation [5,6], while metabolic plasticity enables survival across distinct host niches [7]. Among host nutrients, N-acetylglucosamine (GlcNAc) is crucial as it serves both as a carbon source and as a potent inducer of hyphal growth [8-10]. Further, amino acids provide nitrogen, modulate morphogenesis, and influence antifungal susceptibility [11-13].
Nutrient uptake in fungi is tightly regulated not only at the transcriptional level but also through post-translational mechanisms controlling transporter abundance at the plasma membrane. In Saccharomyces cerevisiae, this regulation is mediated by arrestin-related trafficking (ART) adaptor proteins, which recruit the ubiquitin ligase Rsp5 to target transporters for ubiquitination and endocytosis [14,15]. Similar mechanisms are emerging in C. albicans. For instance, the GlcNAc transporter Ngt1 undergoes ubiquitin-dependent endocytosis in response to sugar availability [16,17], while the broad-specificity amino acid permease CaGAP1 plays a dual role in nitrogen acquisition and virulence [12,13].
Despite the importance of carbon and nitrogen uptake, the ART proteins coordinating transporter turnover in C. albicans remain poorly characterized. In silico analysis of ORF19.4887, designated as CaECM21, revealed a conserved arrestin domain and PPxY motifs predicted to interact with Rsp5, suggesting a role in endocytosis. Recent studies in Cryptococcus neoformans and Aspergillus fumigatus have demonstrated that ART-domain proteins contribute to transporter regulation and antifungal tolerance [18,19]. These findings imply a conserved role of arrestin-like proteins in fungal adaptability and drug response.
Comparative sequence analyses further indicate that CaECM21 shares structural features with its orthologs Rod1 in S. cerevisiae, ArtA in A. fumigatus, and arrestin-like proteins in C. neoformans, all containing characteristic arrestin-C domains and PPxY motifs. This evolutionary conservation suggests that ART-mediated trafficking is an ancient and fundamental mechanism for maintaining plasma membrane homeostasis across fungal species.
Based on these insights, we hypothesized that CaECM21 regulates the endocytosis of nutrient transporters such as CaGAP1, thereby integrating carbon and nitrogen sensing in C. albicans. To investigate the endocytic process, we employed a combination of in silico, molecular, and cell biological approaches. These included generating homozygous CaECM21 gene deletion, C-terminal GFP-tagging, fluorescence microscopy under defined nutrient conditions, and antifungal susceptibility assays. This study provides the first experimental evidence establishing CaECM21 as a functional ART adaptor protein in C. albicans, linking nutrient transporter trafficking to stress adaptation and antifungal response.
2. MATERIALS AND METHODS
2.1. Strains and Culture Conditions
All experiments were performed using the C. albicans laboratory strain BWP17, which is auxotrophic for uridine, histidine, and arginine [Table 1]. Strains were maintained as glycerol stocks at −80°C and freshly streaked onto YPD agar plates (1% yeast extract, 2% peptone, 2% dextrose, 2% agar; HiMedia) before experimentation. Plates were incubated at 30°C for 48 h, and single colonies were used to initiate cultures. For routine propagation, cells were grown in YPD broth (1% yeast extract, 2% peptone, 2% glucose, w/v) at 30°C with shaking (200 rpm). Auxotrophic requirements were supplemented with uridine and histidine (20 μg/mL each) and arginine (30 μg/mL; Sigma-Aldrich) [20].
Table 1: List of strains and plasmids used to prepare Δ/ΔCaECM21.
| Strain | Genotype of constructed strains | Source/reference |
|---|
| Candida albicans strain | | |
| BWP17 | ura3Δ::limm434/ura3Δ::limm434 his1::hisG/his1::hisG arg4::hisG/arg4::hisG | Alistair J.P. Brown |
| HHR1 | ΔCaecm21::HIS1/CaECM21 ura3Δ:: Δ imm434/ura3Δ:: Δ imm434 his1::hisG/his1::hisG arg4::hisG/arg4::his | This study |
| HHR2 | ΔCaecm21::HIS1/ecmΔ:: ARG4 ura3Δ:: Δ imm434/ura3Δ:: Δ imm434 his1::hisG/his1::hisG arg4::hisG/arg4::hisG | This study |
| HHR3 | NGT1-GFPΔ:: URA3/CaECM21 ura3Δ::limm434/ura3Δ::limm434 his1::hisG/his1::hisG arg4::hisG/arg4::hisG | This study |
| HHR4 | pFA-GFPΔ-CaGap1::HIS1/CaGap1 ura3Δ::limm434/ura3Δ::limm434 his1::hisG/his1::hisG arg4::hisG/arg4::hisG | This study |
| HHR5 | pFA-GFPΔ-CaGap1-ecm2Δ::HIS1/Caecm21Δura3Δ:limm434/ura3Δ::limm434 his1::hisG/his1::hisG arg4::hisG/arg4::hisG | This study |
| Escherichia coli strains | | |
| pSN52 | C.d HIS1, KanR (pCR-Blunt II-TOPO vector (Invitrogen) carrying HIS1 gene of Candida dubliniensis) | Rao et al., 2020 |
| pSN69 | pCR-Blunt II-TOPO vector (Invitrogen) carrying the ARG4 gene of Candida dubliniensis | Rao et al., 2020 |
| pFa-γ- ARG4 | plasmid harboring a codon optimized γ, a photostable GFP variant along with the URA3 marker | Rao et al., 2020 |
For experimental assays, overnight cultures were diluted into synthetic defined medium and allowed to reach mid-logarithmic phase (optical density [OD]600 ≈ 0.6). Cells were washed twice with YNB basal medium and resuspended in YNB complete and YNB lacking amino acids (W/O aa; Difco), supplemented with glucose or GlcNAc as the sole carbon source [21,22]. Stock solutions of GlcNAc and amino acids were filter-sterilized (0.45 μm) and added aseptically after autoclaving. Growth was routinely monitored by OD600 readings, and all experiments were performed in three independent biological replicates, each with three technical replicates. Wild-type, Δ/ΔCaecm21, and complemented strains (Δ/ΔCaecm21+CaECM21) were analyzed in parallel to confirm that observed phenotypes were attributable to CaECM21 deletion.
2.2. DNA Sequence Analysis
The C. albicans genome (~14.7 Mbp) encodes approximately 6,000 genes [23]. The ART-domain containing gene CaECM21 was selected for functional analysis and targeted for deletion in the BWP17 background. Homology searches using the basic local alignment search tool (BLAST) [24] revealed ~42% identity and ~58% similarity with S. cerevisiae Rod1, an ART adaptor protein [Figure 4]. Multiple sequence alignment with CLUSTALW [25] confirmed conservation of the ART domain and identified two PY motifs (PPxY) at residues 215–218 and 459–462, which may serve as binding sites for the ubiquitin ligase Rsp5. Further analysis of the CaECM21 open reading frame using Restriction Mapper 2.0 identified unique restriction sites suitable for cloning and mutant construction. Conserved domain analysis using the National Center for Biotechnology Information conserved domain database [26] predicted an N-terminal arrestin domain and several low-complexity regions, consistent with a role in adaptor-mediated protein trafficking [27]. All annotations were cross-checked against the Candida genome database (CGD) (http://www.candidagenome.org). These analyses guided the selection of CaECM21 as a candidate ART adaptor potentially involved in the endosomal transport of CaGap1.
2.3. Preparation of CaECM21 Deletion Mutants in C. albicans
In this study, deletion of CaECM21 in the BWP17 strain (gift from Dr. Swagata Ghosh, University of Kalyani) was performed using polymerase chain reaction (PCR)-mediated homologous recombination [20,28]. The two alleles were sequentially replaced using HIS1 and ARG4 markers, amplified from plasmids pSN52 and pSN69, respectively. Disruption cassettes were generated with primers containing 81 bp homology to the upstream and downstream flanking regions of CaECM21 [Table 2]. Purified cassettes were introduced into electrocompetent BWP17 cells through electroporation. Transformants were recovered in YPD medium containing 1 M sorbitol at 30°C for 2 h and selected on SD agar lacking histidine, producing heterozygous mutants (ΔCaecm21:HIS1, designated HHR1). A second round of transformation with the ARG4 cassette, followed by selection on SD agar lacking arginine, generated homozygous knockouts (Δ/ΔCaecm21:HIS1/ARG4, designated HHR2). Allele replacement was confirmed by diagnostic PCR using primers flanking the targeted locus and internal primers spanning the deleted region. Screening of ~40 transformants yielded ~65% correct replacement for the first allele and ~50% for the second. Two independent Δ/ΔCaECM21 mutants (HHR2-1 and HHR2-2) were retained for downstream phenotypic analyses.
Table 2: Primers used for CaECM21 deletion, confirmation, and GFP tagging in Candida albicans.
| Primer Name | Sequence (5’–3’) | Number of base pairs | Use of Primer | Genotype/construct |
|---|
| Caecm21-Del-F | GAACCATCAATCTTGGAA ATCGCAGCCACCAATTCTCAAA ATAAATCGAGAAGGCTA AGTAGCTCGGATCCACTAGTAACG | 81 | Amplification of the upstream flank for Caecm21 deletion | ΔCaecm21 (single knockout mutant) |
| Caecm21-Del-R | ATTACCACAAAATTTCATTA AAAATACAAAATTAAATATA AATTACAAGATAAATTAATAG CCAGTGTGATGGATATCTGC | 81 | Amplification of the downstream flank for Caecm21 deletion | ΔCaecm21 (single mutant) |
| Caecm21-check-F | TTATTTCCTTGTGGATGTCC | 20 | Confirmation of Caecm21 deletion (forward) | ΔCaecm21 single mutant (confirmation) |
| Caecm21-check -R | TATCGTGACTATTCTGATGG | 20 | Confirmation of Caecm21 deletion (reverse) | ΔCaecm21 single mutant (confirmation) |
| His-check.R | ATCAGATGGGTTATCTCGTC | 20 | Confirmation of HIS1 marker insertion | ΔCaecm21::HIS1 mutant |
| ARG4-check-R | TGGTTCAGGTAGATATTCCT | 20 | Confirmation of ARG4 marker insertion | Δ/ΔCaecm21::HIS1/ARG4 double mutant |
| Ngt1-GFP-F | ACCCCCATCGTTTTACTT AGTTCAAAATGTCAAGAAG ATTCTCCTCCTTATAGTTCT GTAGGTGCTGGCGCAGGTGCTTC | 80 | C-terminal tagging of Ngt1 using plasmid pFA-γ-URA in both BWP17 and Δ/ΔCaecm21 (forward primer) | Ngt1- γ (BWP17) and Ngt1- γ (Δ/ΔCaecm21) tagged strain |
| Ngt1-GFP-R | AAAATTACCACAAAATTT CATTAAAAATACAAATTAAAT ATAAATTACAAGATAAATTA ATATCTGATATCATCGATGAATTCGAG | 86 | C-terminal tagging of Ngt1 using plasmid pFA-γ-URA in both BWP17 and Δ/ΔCaecm21 (reverse primer) | Ngt1- γ (BWP17) and Ngt1- γ (Δ/ΔCaecm21) tagged strain |
| CaGap1-GFP-F | AGATTTGTTACAACAAGAAA TAGCTGAAGAAAAAGCTC AATTAGCTGAAAAACCATTC TATATGGTGCTGGCGCAGGTGCTTC | 83 | C-terminal tagging of CaGap1 using plasmid pFA-γ-URA in both BWP17 Δ/ΔCaecm21 (forward primer) | CaGap1-γ (BWP17) and CaGap1- γ (Δ/ΔCaecm21) tagged strain |
| CaGap1-GFP-R | ACTATATATATTAAGTATCA AAATTTGTCTATCTATTATTTTTAGCAC CAAAATCTATAAATTTCTGATAT CATCGATGAATTCGAG | 87 | C-terminal tagging CaGap1 using plasmid pFA-γ-URA in both BWP17 Δ/ΔCaecm21 (reverse primer) | CaGap1-γ (BWP17) and CaGap1- γ (Δ/ΔCaecm21) tagged strain |
For complementation, a wild-type CaECM21 fragment, including ~500 bp upstream promoter and ~300 bp downstream terminator, was cloned into a URA3-marked integration vector. The construct was linearized and introduced into Δ/ΔCaECM21 strains by electroporation. Uracil prototrophs were selected on SD-URA lacking plates, and correct integration was verified by PCR. Complemented strains (Δ/ΔCaECM21+CaECM21) were analyzed alongside wild-type and knockout strains to confirm that observed phenotypes were attributable to CaECM21 deletion. A schematic representation of the deletion and complementation strategy is depicted with a schematic diagram [Figure 1].
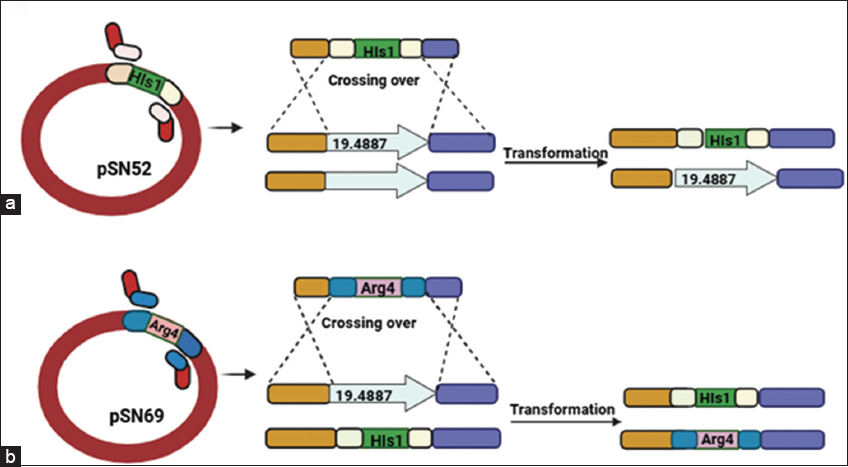 | Figure 1: Schematic representation of CaECM21 (ORF19.4887) gene deletion and generation of the homozygous knockout strain in Candida albicans. (a) The first allele of CaECM21 was disrupted using a deletion cassette derived from plasmid pSN52 carrying the HIS1 marker. Homologous recombination in the BWP17 strain replaced one copy of the CaECM21 locus with HIS1, and transformants were selected on SD medium lacking histidine and verified by diagnostic polymerase chain reaction (PCR). (b) The second allele was deleted using a cassette from plasmid pSN69 containing the ARG4 marker, transformed into the heterozygous strain. Transformants were selected on SD medium lacking arginine, and successful homozygous deletion mutants were confirmed by PCR analysis.
[Click here to view] |
2.4. Spot-Dilution Assay of GlcNAc-Induced Δ/ΔCaecm21 Cells
To assess the nutrient-specific growth, wild-type (BWP17), double mutant (Δ/ΔCaecm21), and complemented strains (ΔCaecm21) were grown overnight in YPD, washed with YNB basal medium, and adjusted to OD600 = 1. Serial dilutions (10-1 to 10-4) were spotted (10 μL) onto YNB-glucose and YNB-GlcNAc plates and incubated at 30°C for 24 h. Growth patterns were documented, and colony area was quantified using ImageJ [Figure 2]. All data represent mean±SD of triplicate biological repeats. Statistical significance was determined using one-way analysis of variance (ANOVA) followed by Tukey’s post hoc test (P < 0.05) [21].
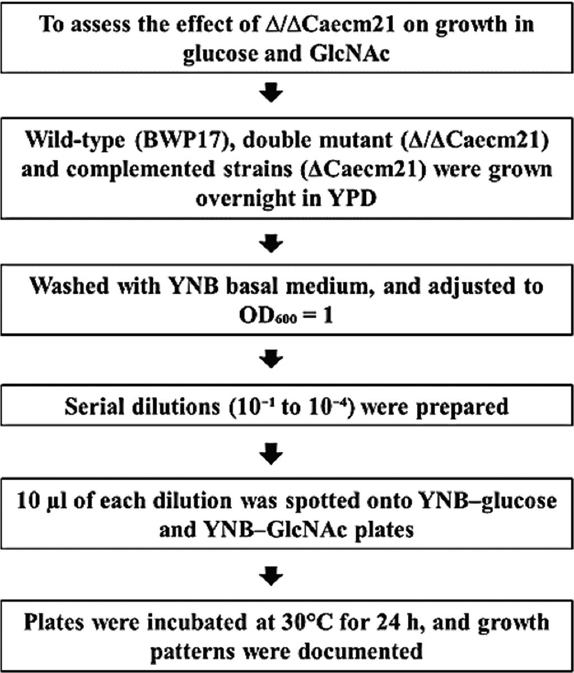 | Figure 2: Experimental workflow to assess the effect of glucose and GlcNAc on growth in Candida albicans strains. Spot assay of C. albicans strains on glucose and GlcNAc. Serial dilutions of wild-type, mutant, and complemented strains were spotted on YNB plates with glucose or GlcNAc and incubated at 30 °C for 24 h.
[Click here to view] |
2.5. Microscopic Study of GlcNAc-Induced Δ/ΔCaecm21 Cells
C. albicans strains were freshly revived on YPD agar, and a single colony was inoculated into 5 mL YPD broth. Cultures were incubated overnight at 30°C. The following day, 1% of the overnight culture was transferred into fresh YPD broth and grown to mid-logarithmic phase (OD600 ≈ 0.4). Cells were harvested by centrifugation, washed twice with YNB basal medium, and resuspended in YNB supplemented with glucose and GlcNAc. Cultures were incubated at 30°C for 2 h to induce morphological changes [Figure 5b] [29].
2.6. Epitope Tagging
C-terminal GFPγ tagging of Ngt1 and CaGAP1 proteins using a PCR-based strategy was performed in both BWP17 and Δ/ΔCaecm21 backgrounds using plasmid pFA-GFPγ-URA [30,31]. PCR cassettes with ~80 bp homology were electroporated into competent cells. Transformants were selected on SD-URA lacking plates, and integration was verified by PCR [Figure 3]. For bright-field microscopy, 5 μL of each cell suspension was mounted on a glass slide, covered with a coverslip, and observed using a Nikon 80i inverted microscope equipped with a Nikon DXM1200C digital camera [Figure 6a]. For fluorescence microscopy of Ngt1-GFPγ and CaGap1-GFPγ expressing cells, imaging was performed using an Olympus confocal microscope with a C-Apochromat ×60/1.2 numerical aperture objective. GFP was excited at 488 nm (range 440–475 nm) and emission was detected at 509 nm (range 500–540 nm) using the appropriate GFP/FITC filter set. The focal plane was positioned at the midsection of the cells to ensure accurate intracellular localization. Untagged controls were used as negative controls for autofluorescence and background signal. GFP-tagged strains in a wild-type background served as positive controls to ensure that fluorescence reflected specific transporter localization rather than artifacts of tagging [Figure 6b].
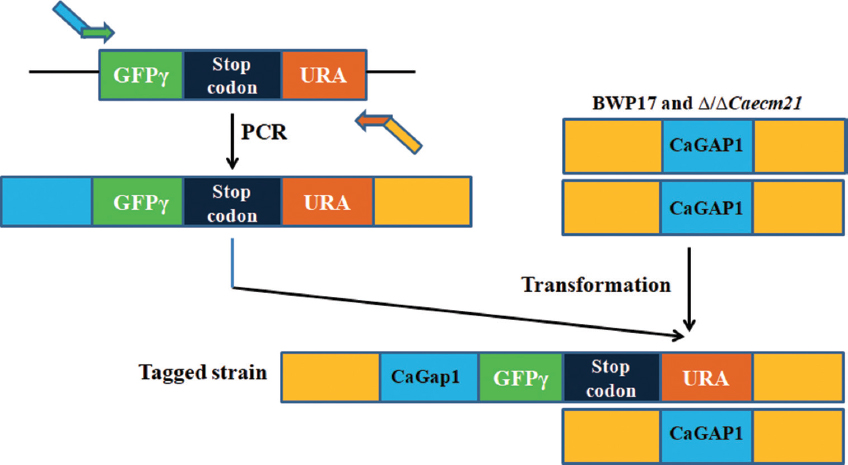 | Figure 3: Schematic overview of C-terminal GFPγ tagging of CaGAP1 in Candida albicans wild-type and Δ/ΔCaecm21 strains. A tagging cassette containing GFPγ, a stop codon, and the URA3 marker was amplified from plasmid pFA-GFPγ-URA and integrated at the 3′ end of the CaGAP1 ORF by homologous recombination. Transformants were selected on URA-lacking medium and confirmed by diagnostic polymerase chain reaction. The resulting CaGap1-GFPγ strains were used for fluorescence microscopy to study protein localization under different carbon source conditions.
[Click here to view] |
2.7. Impact of Glucose and GlcNAc on CaGap1 and Functional Assessment of CaECM21
To investigate the effect of nutrient conditions on CaGap1 localization and to assess the role of CaECM21, wild-type (BWP17) and Δ/ΔCaecm21 strains carrying CaGap1-GFPγ were cultured overnight in YPD at 30°C. Cells were washed twice with YNB basal medium and resuspended in YNB supplemented with both 2% glucose and 2% GlcNAc, with or W/O aa [32]. Untagged strains were used as negative controls to exclude autofluorescence, and images were processed using ImageJ [Figure 7].
2.8. Localization of CaGap1 under Nutrient Conditions
CaGap1 is a broad-spectrum aa transporter in C. albicans, essential for nitrogen acquisition and nutrient sensing [32]. To assess its expression and trafficking, wild-type and Δ/ΔCaecm21 strains expressing CaGap1-GFPγ were grown overnight in YPD. Cells were washed twice and induced under four conditions: YNB + glucose, YNB + GlcNAc, YNB W/O aa + glucose, and YNB W/O aa + GlcNAc. Samples were collected at 60 min and 120 min for fluorescence microscopy. Localization patterns (membrane, cytoplasmic, vacuolar) were quantified using Image J [Figure 8] [33].
2.9. Amphotericin B Sensitivity Assay
As CaEcm21 is observed to be involved in endosomal trafficking of membrane proteins like CaGap1, we wanted to check the sensitivity of mutant cells against the Amp B. Amp B is reported to bind to ergosterol moieties of the cell membrane, affecting the endocytic pathway [33]. Amphotericin B (Amp B) is widely recognized as a highly effective antifungal agent used to treat severe fungal infections [34]. Numerous investigations have highlighted the remarkable potency of Amp B in combating fungal pathogens [35,36]. We further investigated whether CaECM21 contributes to antifungal stress tolerance by testing sensitivity to Amp B. In particular, we assessed sensitivity to Amp B, a clinically relevant polyene that disrupts ergosterol-containing membranes. The susceptibility of C. albicans strains to amphotericin B (Amp B) was assessed by spot assay under four conditions: YNB + dimethyl sulfoxide (DMSO) (WC), YNB culture, 0.7 μg/mL Amp B, and 12 μg/mL Amp B. On control plates (YNB + DMSO, WC) and on YNB culture plates, wild-type, Δ/ΔCaecm21 and the complemented strain displayed comparable growth, indicating that neither the vehicle (DMSO) nor the culture filtrate affected basal viability [37]. Wild-type, Δ/ΔCaecm21, and complemented strains were grown overnight in YPD at 30°C, washed twice with Milli-Q water. Cell suspensions were adjusted to OD600 = 0.4 in sterile water, and ten-fold serial dilutions were prepared. 5 μL of each dilution was spotted onto YNB-GlcNAc agar plates supplemented with either 0.7 μg/mL or 12 μg/mL Amp B. YNB medium without drug was used to monitor basal growth, while YNB + DMSO was included as a vehicle control. Plates were incubated at 30°C for 48–72 h, and growth differences were documented. Spot intensity and colony size were quantified using ImageJ. Data represent averages from three independent biological replicates, and statistical analysis was performed by one-way ANOVA (Tukey’s post hoc, P < 0.05).
2.10. Fluorescence Quantification and Statistical Analysis
Quantitative analysis was performed using ImageJ (version 1.53; NIH, USA). Individual cells were segmented automatically using threshold-based region of interest selection, and mean fluorescence intensity (MFI) [Table 3] was calculated after subtracting background from cell-free regions. Each field contained ≥30 cells, and at least 3 fields were analyzed per replicate. Fluorescence data were represented as averages from three independent biological replicates as mean ± standard deviation (SD). For each comparison, 95% confidence intervals (CIs) and exact P-values were calculated to ensure statistical transparency. Brightness and contrast adjustments were applied uniformly to representative images using Adobe Photoshop version 5.5 (Adobe Systems, San Jose, CA). Statistical analyses were performed using GraphPad Prism 10, which was used for one-way or two-way ANOVA with Tukey’s post hoc tests. Results were considered significant thresholds as P < 0.05 [Figure 6 bi].
Table 3: Data are mean±SD of three independent replicates, significance determined by two-way analysis of variance with Tukey’s post hoc test.
| Time (min) | YNB complete (mean±SD) | YNB W/O-aa (mean±SD) | Significance (Tukey HSD) |
|---|
| 0 | 0.0±0.0 | 0.0±0.0 | NS |
| 30 | 132.7±0.7 | 144.5±0.8 | *** |
| 60 | 138.2±0.4 | 159.5±0.6 | *** |
| 90 | 141.0±0.7 | 166.2±0.9 | *** |
| 120 | 145.2±0.3 | 173.4±0.8 | *** |
3. RESULTS AND DISCUSSION
3.1. In silico Identification of ART Domain-Containing Proteins in C. albicans
To identify ART domain-containing proteins potentially involved in ubiquitin-mediated endocytosis, a BLASTp search was performed in CGD using the α-arrestin Rod1 (C2_01970C_A) as the query. Rod1 mediates internalization of plasma membrane transporters through interaction with the ubiquitin ligase Rsp5. Sequence analysis of C. albicans SC5314 Assembly 22 revealed a previously uncharacterized paralog, CaECM21 (C1_10180C_A), sharing 24.63% sequence identity with Rod1 across the conserved arrestin domain (E-value = 2e-10) [Figure 4]. Conserved domain analysis revealed two PPxY motifs (residues 213–218 and 459–462) predicted to interact with the WW domain of the ubiquitin ligase Rsp5. Such motifs are essential for ubiquitin-mediated endocytosis in S. cerevisiae and other fungi [38]. In silico predictions suggest that CaECM21 could function as a putative ART adaptor facilitating the internalization of nutrient transporters such as Ngt1 and CaGAP1.
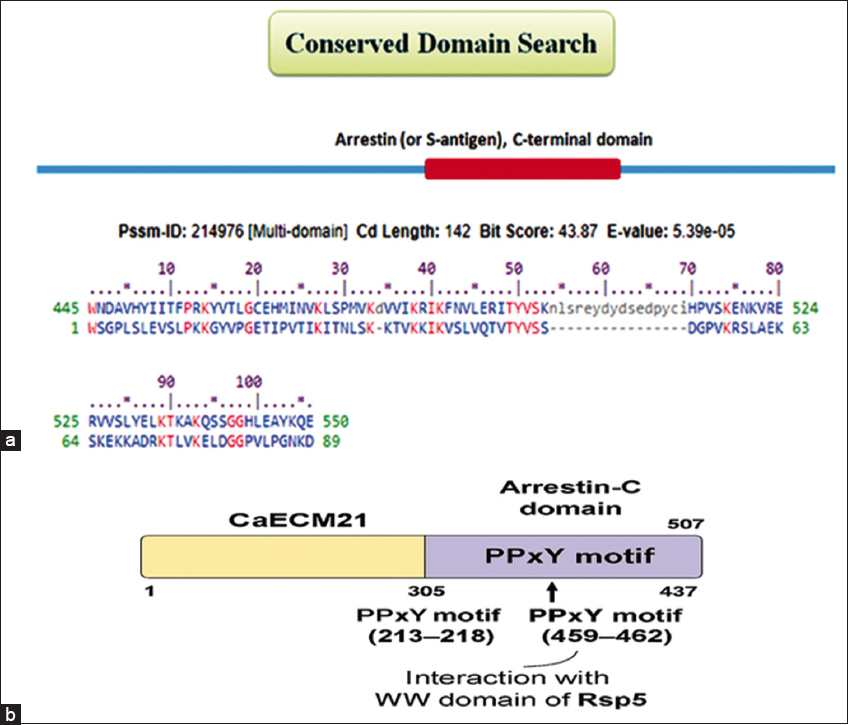 | Figure 4: Structural and functional representation of CaECM21 in ubiquitin-mediated endocytosis. (a) A conserved domain search revealed that CaECM21 in Candida albicans contains an Arrestin-C domain, similar to Rod1 (ORF19.1509), which also harbors a PPxY motif essential for Rsp5 interaction. BLASTP analysis identified CaECM21 as structurally related to Rod1, suggesting its potential role in regulating plasma membrane transporter internalization, possibly targeting Ngt1 and CaGap1 for endocytic turnover. (b) Schematic diagram of the C. albicans, CaEcm21 protein, highlighting key domains and motifs. The protein contains an Arrestin-C domain and two PPxY motifs located at amino acid positions 213–218 and 459–462, which mediate interaction with the WW domain of the E3 ubiquitin ligase Rsp5.
[Click here to view] |
3.2. Characterization of the Δ/ΔCaecm21 Deletion Mutant
The mutant was characterized for growth on different carbon sources, including glucose and GlcNAc, in the presence and absence of aa using spot assays. Additionally, cell morphology was assessed under glucose and GlcNAc-inducing conditions. Localization studies of CaGap1 were performed to investigate the role of CaEcm21 in endosomal trafficking. The Δ/ΔCaecm21 mutant was also tested for sensitivity to the antifungal agent amphotericin B. These analyses provide the first evidence that CaECM21, an ART-family protein, contributes to the endosomal trafficking of CaGap1. Notably, CaGap1-GFPγ expression is sensitive to YNB-GlcNAc and displays membrane localization in the presence of aa, supporting the role of CaEcm21 in regulating nutrient transporter trafficking.
3.3. Growth and Morphogenesis of CaECM21 Mutant Strains
To assess the role of CaEcm21 in C. albicans growth on glucose and GlcNAc media, spot assays were performed using the wild-type strain BWP17, double mutant (Δ/ΔCaecm21), and complemented strains (ΔCaecm21) [Figure 5a]. All strains were able to grow under both conditions, while Δ/ΔCaecm21 strains showed reduced growth both on glucose and GlcNAc plates, as shown in the bar diagram [Figure 5ai] and the statistical analysis [Table 4, P < 0.0001]. This indicates a condition-specific contribution of CaECM21 to nutrient adaptation.
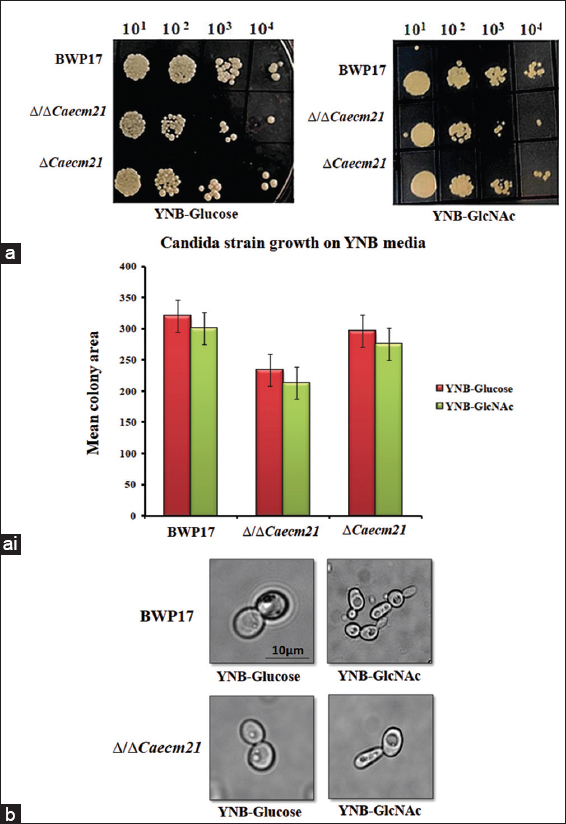 | Figure 5: Growth and morphological responses of wild-type, Δ/ΔCaecm21, and complemented strains under glucose and GlcNAc conditions. (a) Spot assay of BWP17, double mutant (Δ/ΔCaecm21) and complementary strains (Δ???????Caecm21) on YNB-glucose and YNB-GlcNAc. (a i) Quantitative growth analysis using integrated density measurements from four biological replicates (n = biological replicates [4], analysis of variance, P < 0.0001), Image J tool. (b) Microscopic examination of yeast-to-hyphal transition under glucose and GlcNAc after 2 h of induction. Scale bar: 10 μm.
[Click here to view] |
Table 4: Mean colony area of Candida albicans strains grown on YNB-glucose and YNB-GlcNAc media.
| Strain | YNB-glucose (mean±SD) | 95% CI | P versus WT | YNB-GlcNAc (mean±SD) | 95% CI | P versus WT |
|---|
| BWP17 | 320.4±8.1 | (307.5, 333.3) | Reference | 300.3±6.0 | (290.7, 309.9) | Reference |
| Δ/ΔCaecm21 | 234.2±5.5 | (225.5, 242.9) | 0.0007 | 213.3±5.4 | (204.7, 221.9) | 0.0011 |
| ΔCaecm21 | 296.3±5.9 | (286.9, 305.7) | 0.039 | 275.1±6.2 | (265.2, 285.0) | 0.041 |
To further investigate the role of CaECM21 in morphogenetic transition, cells were induced in YNB-glucose and YNB-GlcNAc media for 2 h, and morphological changes were examined microscopically. Cells grown in glucose predominantly retained a yeast-like morphology, whereas GlcNAc-induced cells formed mycelia in both wild-type and mutant strains, with no notable differences in morphological changes [Figure 5b]. These findings suggest that CaECM21 contributes to efficient growth and GlcNAc-induced morphogenesis, although its absence reduces the overall response.
3.4. Plasma Membrane Localization of GlcNAc-Induced Δ/ΔCaecm21
To determine whether membrane transporters’ localization correlates with CaECM21 predicted role in endosomal trafficking of nutrient transporters, the fusion protein was examined under GlcNAc-inducing conditions (YNB-GlcNAc) using fluorescence microscopy. Under these conditions, Ngt1-GFPγ localized predominantly to the plasma membrane [Figure 6a]. These observations provide the first experimental evidence for GlcNAc-dependent plasma membrane localization of Ngt1 in C. albicans and are consistent with previous studies employing similar GFP-tagging approaches [34].
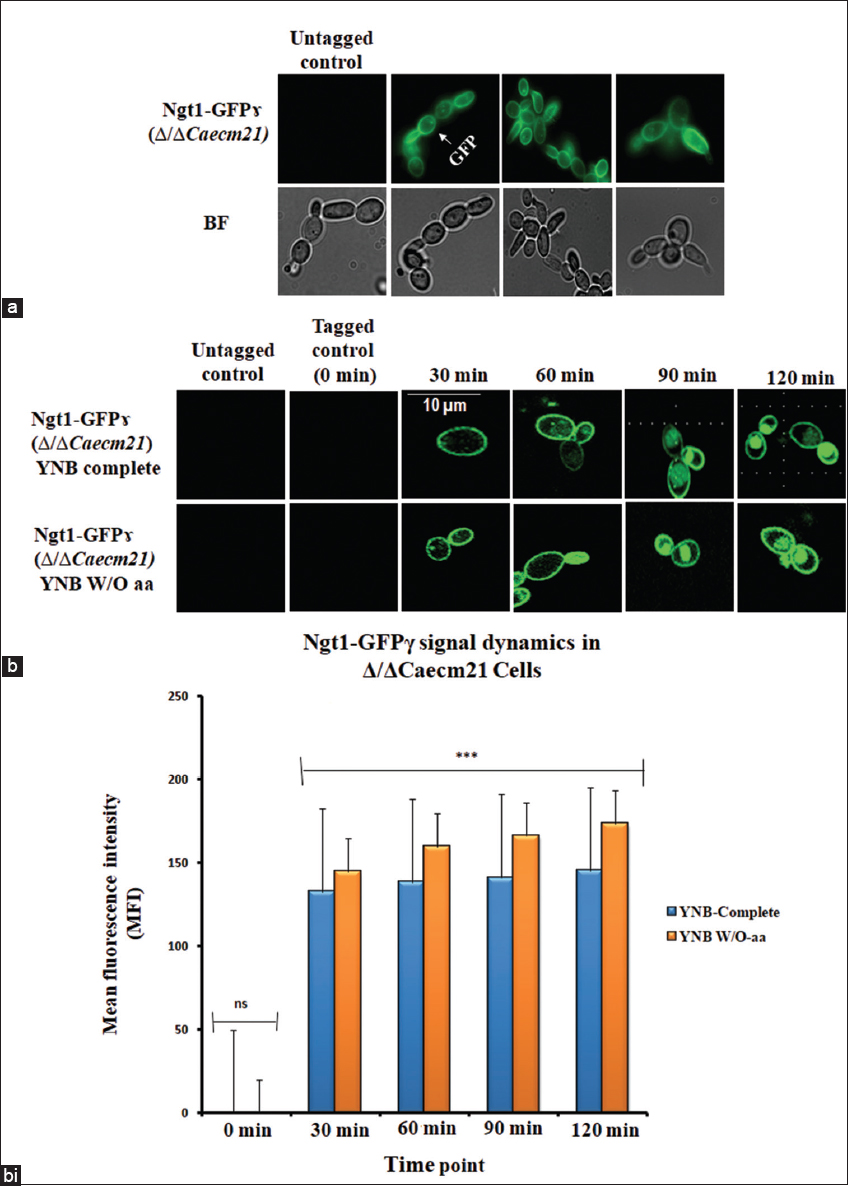 | Figure 6: GlcNAc-induced localization dynamics of Ngt1-GFPγ in Δ/ΔCaecm21 cells. (a) Fluorescence microscopy of Ngt1-GFPγ in ΔΔCaecm21 cells under GlcNAc induction. BF = bright field. (b) Time-course panels showing subcellular localization at 0, 30, 60, 90, and 120 min, Scale bar: 10 μm. (b i) Bar graph showing mean fluorescence intensity (MFI) of Ngt1-GFPγ under two nutrient conditions: YNB complete (blue) and YNB W/O-aa (orange). Fluorescence was undetectable at 0 min in both conditions (ns). From 30 min onward, MFI increased progressively in both media, with consistently higher values observed in YNB W/O-aa. Error bars represent standard deviation (SD) from three biological replicates (n = 3). Statistical significance was determined using Tukey’s post hoc test following two-way analysis of variance (P < 0.001, ns = not significant).
[Click here to view] |
3.5. Ngt1-GFPγ Localization under Amino Acid Limitation
To investigate the time-dependent relocalization of Ngt1 in Δ/ΔCaecm21 cells, cultures were grown overnight and incubated with YNB-GlcNAc and YNB W/O aa-GlcNAc conditions. All fluorescence pictures were observed under the epifluorescence microscope at different time intervals. Importantly, no detectable fluorescence was observed in untagged strains, confirming that signals originated specifically from GFP-tagged transporters. Tagged mutant strains at 0 min served as negative controls and showed no signal. Interestingly, between 30 and 60 min, fluorescence appeared at the plasma membrane, indicating transporter activation. At 90 min, internal puncta were visible, and at 120 min, fluorescence was predominantly vacuolar, suggesting endocytic trafficking [Figure 6b].
Quantitative analysis of MFI confirmed these localization dynamics across three biological replicates [39]. Cells grown in YNB W/O-aa consistently exhibited higher MFI than those in YNB complete medium at all time points beyond 0 min. The observation was compared across conditions using two-way ANOVA followed by Tukey’s post hoc test [P < 0.001, n = 3, Table 3]. The statistical analysis validated the observed differences in transporter localization, confirming that trends were not due to random variation. These results support the hypothesis that CaECM21 acts as an ART adaptor promoting nutrient-dependent transporter internalization [Figure 6bi].
3.6. Impact of Glucose and GlcNAc on CaGap1 and Functional Assessment of CaECM21
Under YNB + glucose or GlcNAc, both strains exhibited weak CaGap1-GFP signals, indicating low CaGap1 expression. In contrast, amino acid starvation (YNB W/O aa) enhanced CaGap1 internalization, with pronounced vacuolar accumulation under GlcNAc, while glucose induced weaker internalization [Figure 7]. These observations indicate that CaGap1 trafficking is strongly nutrient-dependent, with GlcNAc under amino acid starvation robustly promoting internalization, independent of CaECM21. Although the present study did not assess protein-protein interactions, future co-immunoprecipitation and ubiquitination assays will be instrumental in confirming the direct role of CaEcm21 as an adaptor for Rsp5-mediated transporter endocytosis.
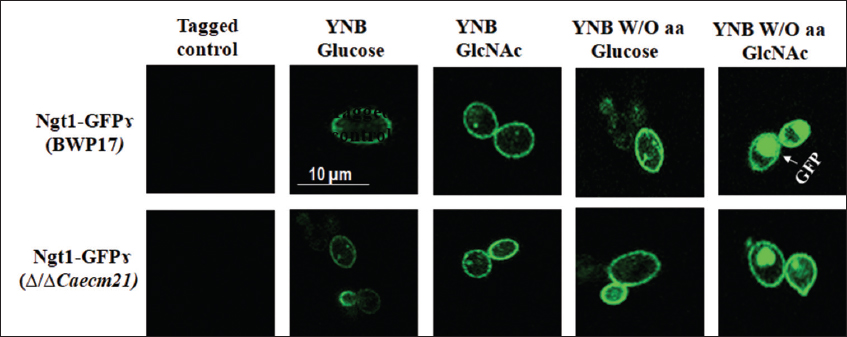 | Figure 7: Nutrient-dependent localization of CaGap1-GFPγ in Candida albicans. Fluorescence microscopy of wild-type (BWP17) and Δ/ΔCaecm21 strains grown in YNB with glucose or GlcNAc, with or without amino acids. Amino acid starvation enhanced CaGap1 internalization with a strong vacuolar signal in GlcNAc-grown wild-type cells, whereas the mutant retained membrane-associated fluorescence. Scale bar = 10 μm.
[Click here to view] |
3.7. Effect of GlcNAc-Induced Expression of CaGap1
In YNB-GlcNAc medium, in both wild-type and mutant strains, CaGap1 is mainly localized in the cell membrane, as in YNB W/O aa GlcNAc medium, pFA-GFPΔ (BWP17) majorly shows its localization signal at the vacuole. These results indicate that CaGap1 localizes to the plasma membrane in the presence of amino acids (YNB-GlcNAc medium), consistent with its role in aa uptake. To observe the internalization expression of CaGAP1, both wild-type and mutant strains were induced in YNB complete and YNB W/O aa. Fluorescence images were captured at 60 min and 120 min, respectively. Both wild type and mutant induced with YNB-GlcNAc, indicating low signal. Impressively, YNB W/O aa-GlcNAc induced wild-type cells show more internalization signal at the vacuole compared to the mutant [Figure 8].
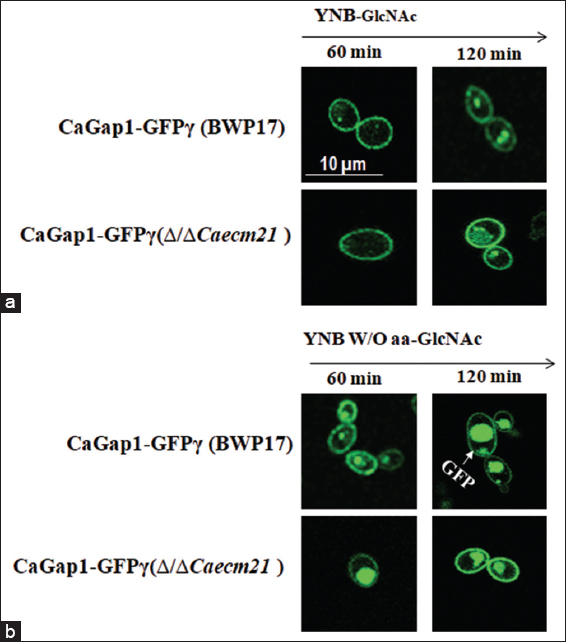 | Figure 8: Time-dependent localization of CaGap1-GFP in Candida albicans under GlcNAc conditions. (a) Fluorescence microscopy images of wild-type (BWP17) and Δ/ΔCaecm21 strains expressing CaGap1-GFPγ were cultured in YNB medium supplemented with GlcNAc and examined after 60 and 120 min. Both strains showed peripheral GFP fluorescence at both time points. (b) Cells grown in YNB W/O aa plus GlcNAc were analyzed at 60 and 120 min. Under amino acid starvation, GFP signal increased over time, with pronounced vacuolar accumulation in BWP17 cells at 120 min (arrow), while the Δ/ΔCaecm21 strain displayed altered localization.
[Click here to view] |
3.8. Sensitivity to Amp B
In contrast, exposure to Amp B produced a clear, concentration-dependent growth inhibition pattern [Figure 9]. While wild-type and complemented strains exhibited normal tolerance, Δ/ΔCaecm21 displayed hypersensitivity at both 0.7 μg/mL and 12 μg/mL [P < 0.05, Table 5]. This increased sensitivity likely results from altered membrane composition or trafficking defects affecting stress adaptation. The observation links CaECM21-mediated endocytosis to the maintenance of plasma membrane integrity under antifungal stress, although the direct molecular mechanism remains to be established.
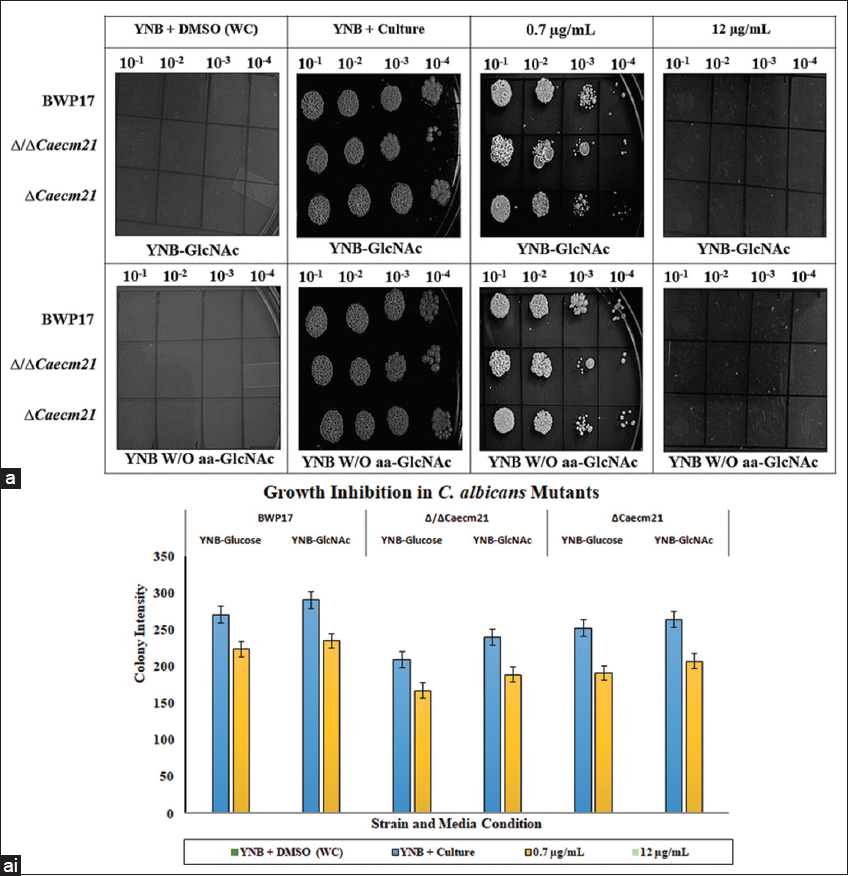 | Figure 9: Antifungal spot assay of amphotericin B (Amp B) against Candida albicans strains. (a) Wild-type, Δ/ΔCaecm21, and complemented strains were spotted in ten-fold serial dilutions (10-1 to 10-5) onto YNB-GlcNAc agar plates under four conditions: YNB + DMSO (without-drug control, WC), YNB culture, 0.7 μg/mL Amp B, and 12 μg/mL Amp B. Plates were incubated at 30°C for 48–72 h, and growth patterns were documented. Representative images are shown (upper panel). (a i) Quantitative analysis of colony size and spot intensity (integrated density) was performed using ImageJ from three independent biological replicates (lower panel). Data are presented as mean±standard deviation (SD), and statistical significance was assessed using one-way analysis of variance with Tukey’s post hoc test (P < 0.05).
[Click here to view] |
Table 5: Colony intensity of Candida albicans strains under different media conditions.
| Strain+media | YNB+DMSO (WC) mean±SD (95% CI) | YNB+culture mean±SD (95% CI) | 0.7 μg/mL mean±SD (95% CI) | 12 μg/mL mean±SD (95% CI) |
|---|
| BWP17-Glucose | 0±0 (0, 0) | 270.8±54.7 (232.1, 309.5) | 224.0±38.9 (199.3, 248.7) | 0±0 (0, 0) |
| BWP17-GlcNAc | 0.75±1.5 (−1.7, 3.2) | 290.8±59.0 (249.6, 332.0) | 235.3±38.6 (210.7, 259.9) | 0±0 (0, 0) |
| Δ/ΔCaecm21-Glucose | 0±0 (0, 0) | 209.8±79.0 (153.6, 266.0) | 167.5±60.0 (125.9, 209.1) | 0±0 (0, 0) |
| Δ/ΔCaecm21-GlcNAc | 0.75±1.5 (−1.7, 3.2) | 240.3±72.0 (190.4, 290.2) | 189.3±73.0 (138.4, 240.2) | 0±0 (0, 0) |
| ΔCaecm21-Glucose | 0±0 (0, 0) | 252.8±49.0 (218.7, 286.9) | 191.5±49.0 (157.4, 225.6) | 0±0 (0, 0) |
| ΔCaecm21-GlcNAc | 0.75±1.5 (−1.7, 3.2) | 264.3±49.0 (230.2, 298.4) | 207.5±45.0 (176.7, 238.3) | 0±0 (0, 0) |
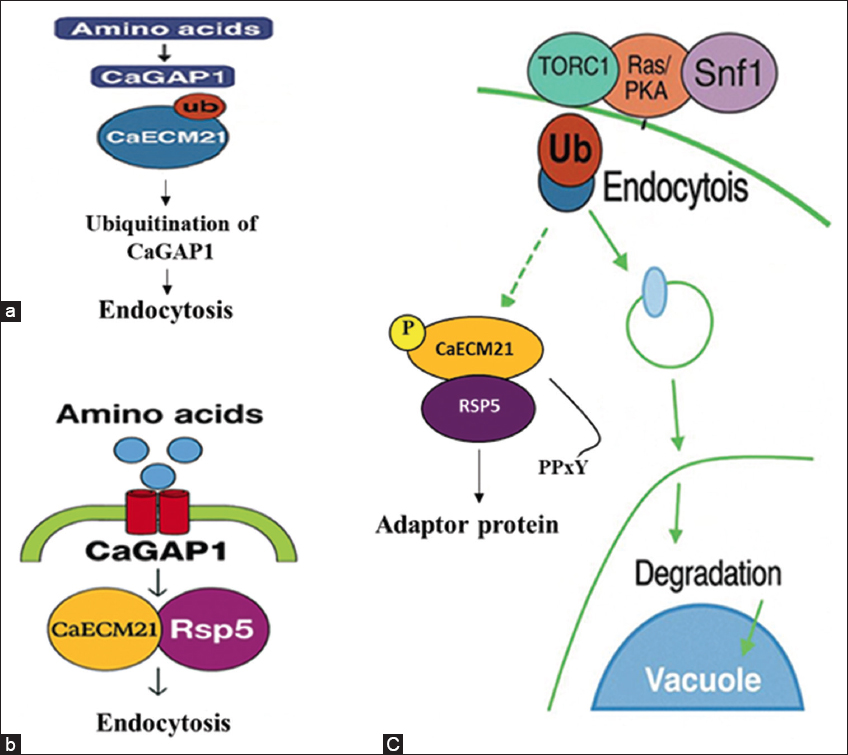 | Figure 10: Model Figure representing CaECM21 in the regulation of CaGAP1 endocytosis under nutrient-rich and nutrient-limited conditions in Candida albicans. (a) Under nutrient-rich conditions, CaGAP1 remains stable at the plasma membrane and mediates amino acid uptake, with minimal ubiquitination and limited endocytosis. (b) During nutrient limitation, CaECM21 contains two conserved PPxY motifs (213–218 and 459–462) that recruit the E3 ubiquitin ligase Rsp5, leading to ubiquitination of CaGAP1 and its endocytosis and triggering its internalization. (c) Integrated schematic showing nutrient-signaling pathways (TORC1, Ras/PKA, and Snf1) that modulate CaECM21 activity. Activated CaECM21 facilitates Rsp5-dependent ubiquitination of CaGAP1, leading to transporter internalization and vacuolar degradation. Rsp5-CaECM21 complex ubiquitinates CaGAP1, promoting endocytosis and transcription of ART genes, highlighting CaECM21 as a central checkpoint integrating nutrient signaling, endocytic regulation, and antifungal stress adaptation.
[Click here to view] |
4. CONCLUSION
This study focused on nutrient adaptation and revealed that CaECM21 (ORF19.4887) functions as a novel ART adaptor protein in Candida albicans. Integrated in silico, and molecular analysis show that CaECM21 regulates the endocytic turnover of nutrient transporter, CaGAP1. The Δ/ΔCaECM21 mutant exhibited reproducible growth defects under nutrient-limited conditions and increased sensitivity to amphotericin B, linking ART-mediated endocytosis to stress adaptation.
Functional characterization revealed that CaECM21 is required for proper localization of transporter, CaGAP1 and internalization under YNB–GlcNAc and W/O-aa conditions, whereas no major defect was observed in YNB–glucose. These findings provide the first experimental evidence connecting CaGAP1 internalization with CaECM21 function in C. albicans, thereby expanding the understanding of ART-mediated endosomal trafficking in this pathogen. Notably, CaECM21 deletion did not affect hyphal morphogenesis under the tested conditions, indicating a specific role in transporter trafficking and stress tolerance rather than global morphogenetic regulation.
The Δ/ΔCaECM21 displayed enhanced susceptibility to Amp B in a concentration-dependent manner, supporting a functional link between endocytic trafficking and membrane stress response. Although the molecular mechanism remains to be elucidated, future studies involving co-immunoprecipitation and ubiquitination assays will probably clarify the interaction network underlying CaECM21-mediated regulation.
Collectively, these results establish CaECM21 as a regulatory component of the fungal endocytic network that coordinates adaptive trafficking under nutrient limitation. By modulating the turnover of CaGAP1, CaECM21 provides mechanistic insight into the integration of carbon and nitrogen metabolism, a process critical for survival and potential virulence in fluctuating host environments. The presence of conserved PPxY motifs further supports its function as an ART-like adaptor. The relatively modest phenotypes observed may reflect functional redundancy among paralogous ART adaptors that partially compensate for CaECM21 deletion. This study mainly focused on CaECM21 as a regulatory component of the endocytic network that integrates nutrient sensing with transporter turnover and antifungal stress adaptation in C. albicans, as illustrated in Figure 10.
4.1. Future Perspective
Future investigations should aim to define the molecular basis of CaECM21–Rsp5 interactions and identify the specific cargo proteins regulated through this pathway. Expanding this analysis to additional ART-domain proteins may uncover a broader regulatory network governing plasma membrane transporter composition and nutrient adaptation in Candida albicans. Although, this study primarily characterized CaECM21-mediated regulation of CaGAP1, it opens avenues for testing whether other nutrient transporters are similarly controlled by this adaptor, providing a framework for understanding transporter specificity versus generality within the ART family.
Further studies using infection models are necessary to evaluate the contribution of CaECM21 to virulence and host–pathogen interactions. Comprehensive genetic analyses will be required to dissect redundancy within the ART network. From a therapeutic perspective, ART adaptors such as CaECM21 represent potential antifungal targets. Interfering with ART-mediated endocytosis may impair nutrient acquisition and stress adaptation, thereby enhancing antifungal susceptibility. Detailed biochemical and structural characterization will be essential to define CaECM21 interaction interfaces and assess its suitability as a selective antifungal target.
5. ACKNOWLEDGMENTS
The authors thank the Department of Science and Technology/SERB (CRG/2022/008507), Government of India, for providing financial support. KHR acknowledges a GITAM: Research Seed Grant, Gandhi Institute of Technology and Management (GITAM), Visakhapatnam, India. The author thanks the members of his laboratory at GITAM and the members of Dr. Swagata Ghosh’s laboratory at the University of Kalyani, India, for their helpful suggestions.
6. CONFLICTS OF INTEREST
The authors report no financial or any other conflicts of interest in this work.
7. AUTHOR’S CONTRIBUTIONS
All authors made substantial contributions to conception and design, acquisition of data, or analysis and interpretation of data; took part in drafting the article or revising it critically for important intellectual content; agreed to submit to the current journal; gave final approval of the version to be published; and agree to be accountable for all aspects of the work. All the authors are eligible to be author as per the International Committee of Medical Journal Editors (ICMJE) requirements/guidelines.
8. ETHICAL APPROVAL
Candida experiments were conducted as per guidelines. The study protocol was approved by the Institutional Biosafety Committee, GITAM School of Sciences, GITAM University, Visakhapatnam, India (Approval No. IAC/GU-1287-1287/HR-F/4; Date: 09.12.2023).
9. DATA AVAILABILITY
All the data is available with the authors and shall be provided upon request.
10. PUBLISHER’S NOTE
All claims expressed in this article are solely those of the authors and do not necessarily represent those of the publisher, the editors and the reviewers. This journal remains neutral with regard to jurisdictional claims in published institutional affiliation.
11. USE OF ARTIFICIAL INTELLIGENCE (AI)-ASSISTED TECHNOLOGY
The authors declare that they have not used artificial intelligence (AI)-tools for writing and editing of the manuscript, and no images were manipulated using AI.
REFERENCES
1. Wisplinghoff H, Bischoff T, Tallent SM, Seifert H, Wenzel RP, Edmond MB. Nosocomial bloodstream infections in US hospitals:Analysis of 24,179 cases from a prospective nationwide surveillance study. Clin Infect Dis. 2004;39(3):309-17. [CrossRef]
2. Pfaller MA, Diekema DJ. Epidemiology of invasive candidiasis:A persistent public health problem. Clin Microbiol Rev. 2007;20(1):133-63. [CrossRef]
3. Lockhart SR, Chowdhary A, Gold JA. The rapid emergence of antifungal-resistant human-pathogenic fungi. Nat Rev Microbiol. 2023;21(12):818-32. [CrossRef]
4. Vazquez JA. Optimal management of oropharyngeal and esophageal candidiasis in patients living with HIV infection. HIV AIDS (Auckl). 2010;2:89-101. [CrossRef]
5. Sudbery PE. Growth of Candida albicans hyphae. Nat Rev Microbiol. 2011;9(10):737-48. [CrossRef]
6. Lo HJ, Köhler JR, DiDomenico B, Loebenberg D, Cacciapuoti A, Fink GR. Nonfilamentous C. albicans mutants are avirulent. Cell. 1997;90(5):939-49. [CrossRef]
7. Ene IV, Brunke S, Brown AJP, Hube B. Metabolism in fungal pathogenesis. Cold Spring Harb Perspect Med. 2014;4(12):a019695. [CrossRef]
8. Brown AJP, Budge S, Kaloriti D, Tillmann A, Jacobsen MD, Yin Z, et al. Stress adaptation in a pathogenic fungus. J Exp Biol. 2014;217(Pt 1):144-55. [CrossRef]
9. Alvarez FJ, Konopka JB. Identification of an N-acetylglucosamine transporter that mediates hyphal induction in Candida albicans. Mol Biol Cell. 2007;18(3):965-75. [CrossRef]
10. Naseem S, Min K, Spitzer D, Gardin J, Konopka JB. Regulation of hyphal growth and N-acetylglucosamine catabolism by two transcription factors in Candida albicans. Genetics. 2017;206(1):299-314. [CrossRef]
11. Biswas S, Van Dijck P, Datta A. Environmental sensing and signal transduction pathways regulating morphopathogenic determinants of Candida albicans. Microbiol Mol Biol Rev. 2007;71(2):348-76. [CrossRef]
12. Ljungdahl PO, Daignan-Fornier B. Regulation of amino acid, nucleotide, and phosphate metabolism in Saccharomyces cerevisiae. Genetics. 2012;190(3):885-929. [CrossRef]
13. Ramachandra S, Linde J, Brock M, Guthke R, Hube B, Brunke S. Regulatory networks controlling nitrogen sensing and uptake in Candida albicans. PLoS One. 2014;9(3):e92734. [CrossRef]
14. Baile MG, Guiney EL, Sanford, EJ, MacGurn, JA, Smolka, MB, Emr SD. Activity of a ubiquitin ligase adaptor is regulated by disordered insertions in its arrestin domain. Mol Biol Cell. 2019;30(25):3057-3072. [CrossRef]
15. Nikko E, Pelham HR. Arrestin-mediated endocytosis of yeast plasma membrane transporters. Traffic. 2009;10(12):1856-67. [CrossRef]
16. Hanumantha Rao K, Roy K, Konopka JB. N-acetylglucosamine transporter Ngt1 undergoes sugar-responsive endocytosis in Candida albicans. Mol Microbiol. 2022;117(3):429-49. [CrossRef]
17. Nikko E, Pelham HRB. Arrestin-mediated endocytosis of yeast plasma membrane transporters.Traffic. 2009;10(11):1856-67. [CrossRef]
18. Lee IR, Lim J, Steen J, Koh TH, Choi J, Jang J, et al. Amino acid permeases and virulence in Cryptococcus neoformans. PLoS One. 2016;11(9):e0163919. [CrossRef]
19. Vaknin Y, Shadkchan Y, Levdansky E, Morozov M, Romano J, Osherov N. The pH signaling transcription factor PacC governs pathogenicity and drug resistance in Aspergillus fumigatus via conserved arrestin-like proteins. Eukaryot Cell. 2014;13(8):1090-101.
20. Hanumantha Rao K, Roy K, Konopka JB. N-acetylglucosamine transporter Ngt1 undergoes sugar-responsive endocytosis in Candida albicans. Mol Microbiol. 2022;117(3):429-49. [CrossRef]
21. Fonzi WA, Irwin MY. Isogenic strain construction and gene mapping in Candida albicans. Genetics 1993;134(3):717-28. [CrossRef]
22. Naseem S, Gunasekera A, Araya E, Konopka JB. N-acetylglucosamine (GlcNAc) induction of hyphal morphogenesis and transcriptional responses in Candida albicans are not dependent on its metabolism. J Biol Chem. 2011;286(33):28671-80. [CrossRef]
23. Jones T, Federspiel NA, Chibana H, Dungan J, Kalman S, Magee BB, et al. The diploid genome sequence of Candida albicans. Proc Natl Acad Sci USA. 2004;101(19):7329-34. [CrossRef]
24. Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ. Basic local alignment search tool. J Mol Biol. 1990;215(3):403-10. [CrossRef]
25. Thompson JD, Higgins DG, Gibson TJ. CLUSTAL W:Improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 1994;22(22):4673-80. [CrossRef]
26. Marchler-Bauer A, Bo Y, Han L, He J, Lanczycki CJ, Lu S, et al. CDD/SPARCLE:functional classification of proteins via subfamily domain architectures. Nucleic Acids Res. 2017;45(D1):D200-3. [CrossRef]
27. Skrzypek MS, Binkley J, Binkley G, Miyasato SR, Simison M, Sherlock G. The Candida Genome Database (CGD):Incorporation of Assembly 22, systematic identifiers and visualization of high throughput sequencing data. Nucleic Acids Res. 2017;45(D1):D592-6. [CrossRef]
28. Noble SM, Johnson AD. Strains and strategies for large-scale gene deletion studies of the diploid human fungal pathogen Candida albicans. Eukaryot Cell. 2005;4(2):298-309. [CrossRef]
29. Gunasekera A, Alvarez FJ, Douglas LM, Wang HX, Rosebrock AP, Konopka JB. Identification of GIG1, a GlcNAc-induced gene in Candida albicans needed for normal sensitivity to the chitin synthase inhibitor nikkomycin Z. Eukaryot Cell. 2010;9(10):1476-83. [CrossRef]
30. Gerami-Nejad M, Berman J, Gale C. Cassettes for PCR-mediated construction of green, yellow, and cyan fluorescent protein fusions in Candida albicans. Yeast. 2001;18(9):859-64. [CrossRef]
31. Dueñas-Santero E, Santos-Almeida A, Rojo-Dominguez P, del Rey F, Correa-Bordes J, Vázquez de Aldana CR. A new toolkit for gene tagging in Candida albicans containing recyclable markers. PLoS One. 2019;14(7):e0219715. [CrossRef]
32. Biswas S, Roy M, Datta A. N-acetylglucosamine-inducible CaGAP1 encodes a general amino acid permease which co-ordinates external nitrogen source response and morphogenesis in Candida albicans. Microbiology (Reading). 2003;149(9):2597-608. [CrossRef]
33. Kraidlova L, Schrevens S, Van Zeebroeck G, Van Dijck P. The Candida albicans GAP gene family encodes permeases involved in amino acid uptake and sensing. Eukaryot Cell. 2011;10(9):1219-29. [CrossRef]
34. Grela E, Zdybicka-Barabas A, Pawlikowska-Pawlega B, Cytrynska M. Modes of the antibiotic activity of amphotericin B against Candida albicans. Sci Rep. 2019;9:17029. [CrossRef]
35. Niimi K, Niimi M. The mechanisms of resistance to echinocandin class of antifungal drugs. Japanese J Med Mycol. 2009;50(2):57-66. [CrossRef]
36. Kavanagh K, Dowd S. Histatins:Antimicrobial peptides with therapeutic potential. J Pharma Pharmacol. 2004;56(3):285-289. [CrossRef]
37. Sahoo S, Sharma S, Singh MP, Singh SK, Vamanu E, Rao KH. Metabolic and phenotypic changes induced during N-acetylglucosamine signalling in the fungal pathogen Candida albicans. Biomedicines. 2023;11(7):1997. [CrossRef]
38. Inglis DO, Arnaud MB, Binkley J, Shah P, Skrzypek MS, Wymore F, et al. The Candida genome database incorporates multiple Candida species:Multispecies search and analysis tools with curated gene and protein information for Candida albicans and Candida glabrata. Nucleic Acids Res. 2012;40(Database issue):D667-74. [CrossRef]
39. Shihan MH, Novo SG, Le Marchand SJ, Wang Y, Duncan MK. A simple method for quantitating confocal fluorescent images. Biochem Biophys Rep. 2021;25:100916. [CrossRef]









