1. INTRODUCTION
Lignocellulosic biomass (LCB) is the most ubiquitous polymer on the earth, and its plentiful source makes it very convenient for usage [1]. Agricultural residues are one of the primary sources of LCB. Burning of lignocellulosic crop residues causes remarkable pollution in the atmosphere and huge nutritional losses in the soil. To overcome these problems, these crop residues can be used to produce second-generation biofuels [2]. LCB consists solely of three structural polymers: Cellulose, hemicellulose, and lignin, organized within a highly resistant matrix. Cellulose, the predominant carbohydrate component, is a linear homopolysaccharide composed of repeated D-glucose monomers linked by β-1,4-glycosidic bonds. Enzymatic and chemical hydrolysis methods are used for the breaking down of cellulose into fermentable sugar monomers, which are essential substrates for further bioethanol synthesis. This conversion mostly relies on the class of enzyme known as cellulases, which is a group of synergistic enzymes, namely, endoglucanases (E.C. 3.2.1.4), cellobiohydrolase (E.C. 3.2.1.91), and β-glucosidase (E.C. 3.2.1.21) [3-5]. Endoglucanases break cellulose chains at random internal sites, producing shorter fragments known as cello-oligosaccharides. Cellobiohydrolases then act on the ends of these fragments and release cellobiose units. Finally, β-glucosidase hydrolyses cellobiose into glucose.
Till now, a lot of research has been done on the enzymatic depolymerization of LCB by various organisms, including bacteria and fungi. Cellulase enzymes produced by fungi are widely used in commercial sectors due to their high enzymatic activity, but recently, researchers have been focusing on bacterial cellulase enzymes due to their rapid growth, multi-enzyme complex expression, stability in extreme environments, and resistance to various environmental stresses, which sets bacteria apart from their fungal counterparts [6,7]. Depolymerization of LCB has been reported for various cellulolytic bacterial species isolated from a variety of sources, such as Bacillus cereus A49 [8], Klebsiella oxytoca M21WG [7], and Klebsiella spp. Z6WG [7], Klebsiella pneumoniae [9], and Enterococcus faecium BS5 [10]. Some other recently isolated cellulolytic bacterial strains with high CMCase activity include Stutzerimonas stutzeri CS7, isolated from the Solanum melongena soil [10]; Bacillus pumilus XM, which is isolated from Père David’s deer feces [11]; and Enterobacter xiangfangensis isolated from the gut of the grasshopper Eyprepocnemis alacris alacris [9].
Many cellulolytic bacterial strains are isolated from diverse sources, such as municipal waste, industrial waste, agricultural leftovers, animal excreta, crop-processing trash, logging slash, and sawdust from timber extraction [12]. However, there is still research ongoing where potential strains of bacteria possessing multiple activities (CMCase, glucosidases, etc.) in the mesophilic range are studied, as there is a growing demand of mesophilic bacteria in simultaneous saccharification and fermentation, which helps to reduce the cost of bioethanol production [13]. Bacterial cellulases play a major role in several industries, especially in food and beverage production, paper and pulp processing, and textile treatments, including biostoning and biopolishing. These enzymes are extensively commercialized through submerged fermentation by many biotechnology companies.
Therefore, in the present research, a cellulase producing bacterial strain was isolated from cow (Bos indicus) fecal matter collected in Godhra, Gujarat, India. Cow dung is undigested food residue that is excreted by herbivorous bovine animals after the consumption of food. Cow dung is a mixture of feces and urine in the ratio of 3:1, and it mainly contains cellulose, hemicellulose, and lignin [14]. Cow dung is the source for various microorganisms, which contain different species of bacteria (Bacillus spp., Corynebacterium spp., etc.), protozoa, and yeast [15,16].
The potential cellulase-producing bacteria were screened out based on the hydrolytic zone on Bushnell Haas (BH) agar plates having carboxymethyl cellulase (CMC) and initial CMCase activity of crude enzyme. B. pumilus A5 was determined to be the bacterial isolate that was chosen after it was subjected to morphological analysis, biochemical profiling, and molecular analyses. Cellulase production of this strain was optimized using the one factor at a time (OFAT) method, after which the activity of the resulting crude enzyme extract was further evaluated and optimized.
2. MATERIALS AND METHODS
2.1. Substrates and Chemicals used in Experiments
CMC; medium viscosity and BH broth were obtained from Sigma-Aldrich, USA. Media and reagents, including Luria-Bertani (LB) broth, yeast extract, agar powder, peptone, tryptone broth, glucose phosphate broth (GPB), Congo red, Lugol’s iodine, and apple pectin, along with Simmon’s citrate agar and triple sugar iron (TSI) agar, were obtained from HiMedia Pvt. Ltd., India. 4-nitrophenyl β-D-glucopyranoside (pNPG) was purchased from Sisco Research Laboratories Pvt. Ltd (SRL), India. All chemicals and reagents used in this study were of analytical grade.
2.2. Sample Collection and Screening of Bacterial Strain
A fecal sample from B. indicus (Malvi breed) was collected from Godhra, Panchmahal, Gujarat, India (22.7788° N, 73.6143° E) using sterile forceps and was kept at 4°C until further use. 1 g of fecal sample was mixed with 10 mL of sterile 0.85 % (w/v) saline solution, and was placed on an orbital shaker (120 rpm) for 1 h until the mixture was homogenized. The homogenized mixture was serially diluted up to 10−8 using saline solution and from each dilution, 100 μL of sample was spread onto CMC-BH agar plates composed of g/L: (CMC (10.0); K2HPO4 (1.0); MgSO4.7H2O (0.2); KH2PO4 (1.0); FeCl3.6H2O (0.05); NH4NO3 (1.0); CaCl2 (0.02); and agar powder (15.0) and incubated at 37°C for 96 h [5]. Morphologically distinct bacterial colonies were selected and further subcultured by streaking on BH-CMC agar plates, followed by incubation at 37°C for 2 days. Each purified colony was grown into fresh LB broth and incubated at 37°C for 16 h. For the qualitative analysis, 3 μL of each overnight-grown culture was placed on CMC-BH agar plates and incubated at 37°C for 48 h. After incubation, agar plates were flooded with 0.3% Congo-red solution for 20 min and counterstained with 1M NaCl for 10 min [17]. For quantitative analysis, the selected isolates were cultured in 50 mL of cellulase production medium (pH 7.0) composed of g/L: (CMC (10.0); K2HPO4 (1.0); MgSO4.7H2O (0.2); KH2PO4 (1.0); FeCl3.6H2O (0.05); NH4NO3 (1.0); CaCl2 (0.02) and yeast extract (5.0) at 37°C, 180 rpm for 72 h. The freshly grown culture was subjected to centrifugation at 10,000 × g for 20 min at 4°C, and the resultant cell-free crude supernatant was utilized to measure CMCase activity. CMCase activity of each crude enzyme was determined using the 3,5-dinitrosalicylic acid (DNS) method [18].
2.3. Identification of Potent Cellulolytic Bacteria
Selected bacterial isolates were subjected to morpho-biochemical characterization and compared with the Bergey’s manual of systematic bacteriology [19]. The Gram stain method was performed to differentiate between Gram-negative and Gram-positive bacteria [20]. The starch hydrolysis property of the selected strain was done by streaking on a starch agar plate, followed by incubation at 37°C for 24 h. Starch agar plates were soaked with Lugol’s iodine solution and examined for a distinct, colorless zone [21]. An isolated bacterial colony was placed onto a sterile, dry glass slide for the catalase test using a sterile nichrome wire loop. After adding a few drops of 3% hydrogen peroxide (H2O2) to the colony, the reaction was examined for the immediate release of gas bubbles (effervescence), which indicated a positive catalase activity [22]. The oxidase test was performed by impregnating a filter paper disc with a 1% (w/v) tetramethyl-p-phenylenediamine dihydrochloride solution, a synthetic electron donor referred to as Wurster’s reagent, and then air drying it. A small portion of the bacterial colony was subsequently put onto the treated disc, and color development was detected within 10 s [23]. Bacterial culture was grown into GPB with incubation of 24–48 h at 37°C. In the methyl red test, a few drops of methyl red were added into GPB and observed for the appearance of a red color. For the Voges-Proskauer (VP) test, 0.6 mL of α-naphthol followed by 0.4 mL of KOH solution were added and observed for the red color development [22]. The bacterial culture was inoculated into peptone nitrate broth and incubated at 37°C for 24 h, for the nitrate reduction test. Following incubation, 0.5 mL of sulfanilic acid and α-naphthylamine were added, and development of a red color was observed, which signifies the nitrate reduction [21]. The indole production was evaluated by inoculating a loopful of the test culture into tryptone broth and incubated at 37°C for 24 h. After incubation, 1 mL of Kovac’s reagent was added, and the appearance of a pink ring at the surface was examined [22]. The TSI test involved streaking a loopful of test culture onto the TSI agar and incubated at 37°C for 24 h. Acid production, gas formation, and H2S production were observed in both the butt and the slant [22]. A loopful of test culture was streaked onto Simmon’s citrate agar slants and incubated at 37°C for 24 h, and the color change from green to blue was observed [23].
2.4. Analysis of 16s Ribosomal RNA (rRNA) of Cellulolytic Bacteria
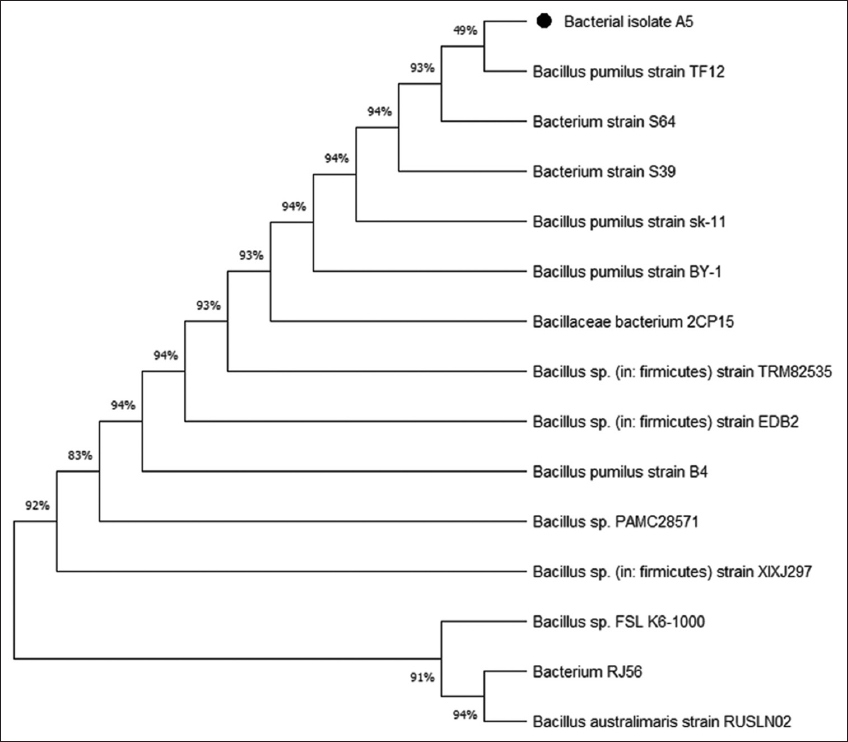
16S rRNA amplification, sequencing, and molecular identification of the bacterial isolate A5 was conducted at Yaazh Xenomics, Coimbatore, Tamil Nadu, India. The Qiagen DNA extraction kit was used to isolate DNA from bacterial samples. The bacterial strain was identified using polymerase chain reaction using universal primers 27F (5’-CCAGAGTTTGATCMTGGCTCAG-3’) and 1492R (5’-TACGGYTACCTTGTTACGACTT-3’) [24]. The 16S gene sequence was then subjected to basic local alignment search tool (BLAST) analysis at the National Center for Biotechnology Information (NCBI) GenBank database, followed by multiple sequence alignment using Thompson et al. [25]. A phylogenetic tree was constructed using the maximum likelihood method, utilizing molecular evolutionary genetics analysis (MEGA) 11.0 software [26].
2.5. Crude Enzyme Production and Endo-β-1, 4-Glucanase Enzyme Assay
Bacterial isolate (A5) with the highest initial CMCase activity was selected and cultured into LB broth and incubated at 37°C, 180 rpm for 16 h. 1 mL of freshly grown bacterial culture was inoculated into 100 mL of enzyme production media (g/L: CMC (10.0); BH (3.27); yeast extract (5.0) at pH 7.0) and incubated at 37°C at 180 rpm for 72 h. The enzyme production medium was subject to centrifugation at 10,000 × g for 10 min after the incubation and the cell-free extract was collected for further analysis [27,28]. CMCase activity was determined by adding 300 μL of crude enzyme along with 600 μL of substrate (1% CMC in 50 mM phosphate buffer, pH 7.0) and incubated for 20 min at 37°C. After incubation, 900 μL of DNS solution was added into the reaction mixture, followed by 10 min incubation in a boiling water bath. The reaction mixture was cooled down after incubation, and 900 μL of distilled water was added before measuring the absorbance using a spectrophotometer at 540 nm and compared to a blank containing all the reagents except the crude enzyme [29]. One unit of CMCase activity was defined as 1 mL of enzyme solution that catalyzes CMC hydrolysis and produces 1 μmoL of glucose/min under given assay conditions [30].
2.6. CMCase Production Optimization
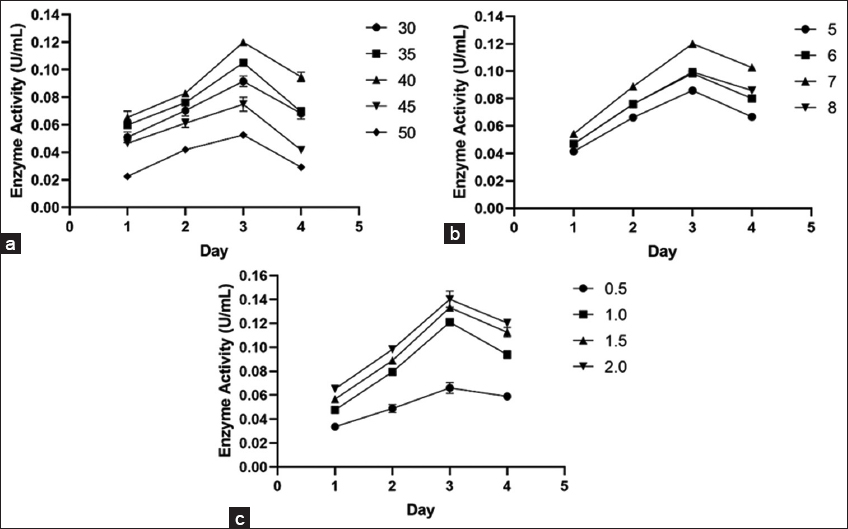
The influence of temperature, incubation time, pH, and CMC concentration was determined by using OFAT. The effect of incubation time on the CMCase production was analyzed by incubating freshly grown bacterial culture in enzyme production media at 37°C, pH 7.0 for 96 h. Production medium was withdrawn at regular intervals and centrifuged at 10,000 × g for 10 min to obtain crude extract. CMCase activity of crude extract was determined through the DNS method as described in section 2.5 [11]. The effect of incubation temperature on the production of CMCase was analyzed by incubating 1 mL of freshly grown bacterial culture into enzyme production media (100 mL), followed by incubation at temperatures ranging from 35 to 50°C at 180 rpm for 96 h. The impact of varying pH levels on CMCase production was assessed by inoculating 1 mL of freshly grown culture into enzyme production medium of different pH levels (5.0, 6.0, 7.0, and 8.0), followed by incubation at 180 rpm for 96 h at the optimal temperature. The enzyme production medium (100 mL) containing different volumes of CMC concentration (0.5, 1.0, 1.5, and 2.0%) was incubated at the optimal incubation temperature and pH to examine the influence of substrate concentration on CMCase production [31].
2.7. Crude Enzyme Activity Characterization
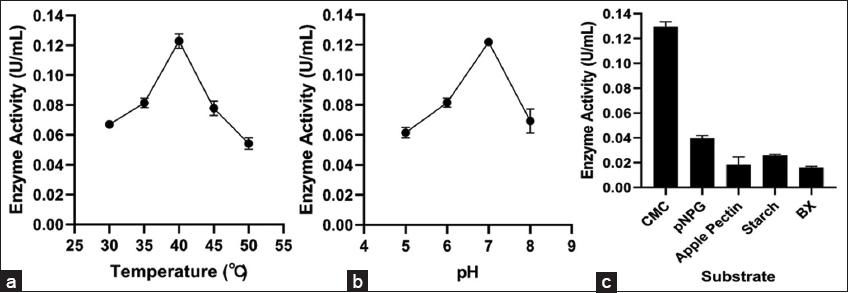
The optimal temperature for endo-β-1,4-glucanase activity from the bacterial isolate was determined by incubating 300 μL of the crude enzyme with 600 μL of CMC (1% w/v in 50 mM phosphate buffer, pH 7.0) at temperatures ranging from 30°C to 50°C for 20 min. The enzymatic process was halted by the addition of 900 μL of DNSA reagent, and the mixture was then incubated in a boiling water bath for 10 min to develop the color. The pH optimum was determined by incubating 300 μL of the crude enzyme with 600 μL of CMC (1% w/v in buffer of pH from 5.0 to 8.0 using 50 mM citrate-phosphate or Tris-HCl buffer) at 40°C for 20 min, followed by the addition of DNSA and a subsequent boiling period of 10 min. Upon cooling, each reaction mixture was diluted by adding 900 μL of distilled water, and absorbance was recorded at 540 nm to evaluate enzyme activity across varying temperature and pH parameters [32].
2.8. Substrate Specificity
The crude B. pumilus A5 extract was tested against substrates: CMC, BX, pNPG, starch, and apple pectin. 900 μL reaction mixture containing 600 μL substrate (1% in 50 mM phosphate buffer, pH 7.0) and 300 μL crude enzyme was incubated for 20 min at 40°C. After incubation, 900 μL of DNS reagent was added to the reaction mixture and further incubated in a boiling water bath for 10 min, followed by the addition of 900 μL of distilled water, and the absorbance was measured at 540 nm using a spectrophotometer [33,34]. All assays were carried out in triplicate. One unit of CMCase, xylanase, amylase, and pectinase were defined as the amount of enzyme used to produce 1 μmoL of glucose, xylose, maltose, and galacturonic acid, respectively, per minute under defined assay conditions [35]. The β-glucosidase activity was assessed by adding crude enzyme (500 μL) with 1,000 μL of 5 mM pNPG, prepared in 50 mM phosphate buffer (pH 7.0), and incubated for 20 min at its optimum temperature. After incubation, 1,500 μL of Na2CO3 (1 M) was added to the reaction mixture to terminate the process. Absorbance was measured at 405 nm using a spectrophotometer using para-nitrophenol (pNP) as a standard [30,36]. One unit of β-glucosidase activity is defined as the amount of enzyme used to produce 1 μmol of pNP/min under specific assay conditions [35].
2.9. Data Analysis
All graphs were generated using GraphPad Prism version 8.0. Cellulase production under varying conditions was evaluated by two-way analysis of variance at p < 0.0001. A phylogenetic tree was created using MEGA 11.0 software.
3. RESULTS AND DISCUSSION
3.1. Isolation and Screening of Cellulolytic Bacteria
Five bacterial isolates were selected on the basis of morphological differences after spreading the B. indicus feces sample on CMC-BH agar plates. Out of these five isolates, only isolate A5 demonstrated a visible hydrolytic zone after staining with 0.3% Congo red solution [Figure 1a]. All isolates were also investigated quantitatively, and isolate A5 demonstrated notable initial CMCase activity (0.064 ± 0.0045 U/mL) after 72 h of incubation at 37°C [Figure 1b]. The initial CMCase activity of isolate A5 was comparable with some other known cellulolytic producing strains isolated from various environments, which include Cellulomonas spp. (0.033 U/mL) [36], Micrococcus spp. (0.015 U/mL) [36], Brevibacillus spp. JXL (0.02 U/mL) [37], Geobacillus spp. (0.012 U/mL) [38], Pseudomonas spp. NE (0.061 U/mL) [32] and Bacillus subtilis AS3 (0.07 U/mL) [39].
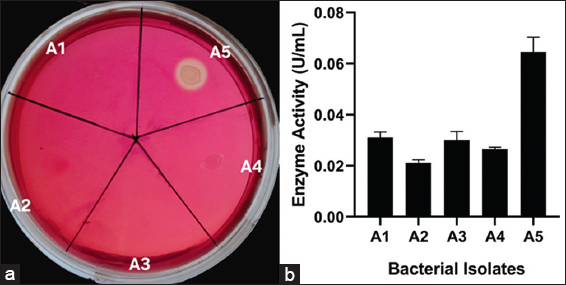 | Figure 1: Primary screening of cellulase producing bacterial isolates on Bushnell Haas Carboxymethyl cellulose agar plates (a) Plates showing clear hydrolytic zone surrounding bacterial isolates after staining with 0.3% Congo red solution (b) Initial CMCase activity of bacterial isolates isolated from the fecal sample of cow. [Click here to view] |
3.2. Identification of Potent Cellulolytic Bacterial Strain
Selected bacterial strain A5 was subjected to cultural characterization, such as size, shape, and pigmentation. Microscopic examination under an oil immersion lens (×100) revealed that bacterial isolate A5 was Gram-positive rod-shaped in nature [Figure 2]. According to biochemical characterization, isolate A5 indicated negative results for the indole, starch hydrolysis, and nitrate reduction tests, while yielding positive results for the VP, methyl red, oxidase, catalase, and citrate utilization tests. In the case of the TSI test, the color of the butt and slant remained orange with no H2S production [Table 1]. Similar biochemical profiles were reported by Parvathi et al. [40] and Kapilan and Arasaratnam [41] for B. pumilus. These morpho-biochemical characteristics, together with a comparison with the Bergey’s manual of systematic bacteriology, imply that isolate A5 is most likely B. pumilus.
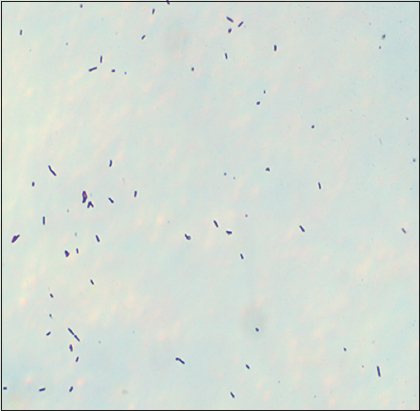 | Figure 2: Microscopic observation of bacterial cells of B. pumilus A5 after Gram staining method [Click here to view] |
Table 1: Morphological and biochemical characterization of B. pumilus A5 isolated from the feces of a cow.
| Characterization | Reference reactions | A5 |
|---|---|---|
| Shape | Short rod | |
| Gram staining | + | |
| Pigmentation | Off white | |
| Colony texture | Opaque | |
| Colony margin | Irregular | |
| Methyl Red | Positive (pink color) | + |
| Voges Proskauer | Positive (pink color) | + |
| Indole production | Negative (no color change) | - |
| Catalase | Positive (effervesces observed) | + |
| Oxidase | Positive (purple color) | + |
| Nitrate reduction | Negative (no color observed after A and B, but after the addition of zinc) | - |
| Starch hydrolysis | Negative (no clear zone) | - |
| Citrate utilization | Positive (blue color) | + |
| TSI (H2S) | Negative | - |
| Gas production | No gas production | - |
| TSI (Butt color) | Orange | |
| TSI (Slant color) | Orange |
TSI: Triple sugar iron, +: Positive, −: Negative.