1. INTRODUCTION
Human activities, such as improper industrial waste disposal, rapid urbanization, and the excessive use of fertilizers and pesticides are significant sources of heavy metal pollution in the soil ecosystem. González Henao and Ghneim-Herrera [1] reported that approximately 5 million soil sites worldwide are contaminated with heavy metals above regulatory limits. Among various anthropogenic sources of metal contamination, foundry waste plays a major role. In India, about 1.71 million tons of foundry waste are generated annually, which contaminates the soil ecosystem and causes a significant threat to the environment, as stated by Faisal et al. [2]. Foundry contaminated soil is of particular concern because it contains toxic metals, such as nickel. Accordingly, considerable concentrations of nickel have been detected in spent foundry soil, as reported by Cioli et al. [3] and Bozym [4]. The continuous release of nickel into the environment is dangerous because it accumulates in flora and fauna and ultimately threatens human health and organisms at all trophic levels [5].
In addition to environmental impacts, nickel exposure in humans can damage vital organs, such as the lungs, kidneys, heart, and skin, leading to pulmonary fibrosis, cardiovascular diseases, renal edema, and contact dermatitis. Furthermore, chronic exposure to nickel has been associated with increased risk of respiratory cancer [6]. Considering the health and environmental risks associated with nickel, its removal from the contaminated environment becomes essential. Conventional remediation methods, including soil replacement, soil spading, soil washing, electrokinetic remediation, chemical reduction, chemical stabilization, and photocatalysis, have been employed for nickel remediation. However, these techniques are often costly and damage the environment, which procures a desperate alternative. Bioremediation was chosen as the best replacement for the chemical method, as reported by El-Naggar et al. [7] and Gabr et al. [8].
The present study focused on the use of indigenous nickel-resistant bacteria for the biosorption of nickel from contaminated foundry soil. These native bacterial strains are well-adapted to the environmental conditions where they were isolated and can effectively remove nickel. To enhance the bacterial remediation efficiency under in situ conditions, the native strains were immobilized in non-toxic, supportive matrices, such as alginate, agarose, and chitosan. This approach improves the stability, reusability, and efficiency of microbes in nickel removal [9]. Besides evaluating the biosorption capacity, this study focuses on identifying critical genes responsible for nickel resistance and remediation, with emphasis on their roles in metal binding and cellular transport mechanisms. Overall, this study aimed to develop an environmentally sustainable and practically applicable bioremediation approach using immobilized bacteria for the remediation of nickel-contaminated foundry soils.
2. MATERIALS AND METHODS
2.1. Isolation and Determination of Minimal Inhibitory Concentration (MIC) of Nickel-Resistant Bacteria
The metal-resistant bacterial strain was previously isolated from metal-contaminated foundry soil and identified as Enterobacter cloacae SK1 through 16S rRNA gene sequencing [10]. The MIC of E. cloacae SK1 for nickel was determined using the plate dilution technique [11]. Sterile nickel solution ranging from 100 to 1500 ppm was aseptically mixed with nutrient agar and poured into sterile petri plates. The E. cloacae SK1 (0.5 McFarland standard) culture was then inoculated onto the nickel-amended plates and incubated at 37°C for 48 h. The nickel concentration that completely inhibited the bacterial growth was determined as the MIC [12].
2.2. Effect of Different Growth Parameters on Nickel Biosorption by E. cloacae SK1
2.2.1. Effect of contact time and initial metal ion concentration
To optimize the nickel remediation potential of the active strain, the effect of initial nickel ion concentrations ranging from 50 to 250 ppm and contact time was investigated. Each experimental flask containing nutrient broth was inoculated with 1% of E. cloacae SK1 (0.5 McFarland standard) and incubated at 37°C for various contact times (6, 12, 18, 24, 30, 36, 42, 48, 54, 60, 66, and 72 h) [13,14]. After incubation, samples were withdrawn at each of the above specified contact times and centrifuged at 5000 rpm for 15 min. The nickel concentration in the supernatant was analyzed by adding 1 mL of dimethylglyoxime solution, 2 mL of saturated bromine water, and 5 mL of ammonia solution (dimethylglyoxime method). Distilled water was also subjected to the above-said method served as the blank. Absorbance was then measured using a UV-Vis spectrophotometer (Visible Spectrophotometer, LABMAN, LMSP-V320, India) at a wavelength of 470 nm [15-17]. The nickel removal percentage was calculated using the following formula.
(Ci is the initial nickel concentration in the medium and Cf is the final nickel concentration in the medium).
2.2.2. Effect of pH
The effect of pH on nickel remediation by E. cloacae SK1 was examined at various pH values ranging from 1 to 14. The experiments were carried out using sterile nutrient broth amended with 50 ppm of nickel. Further, the broth was inoculated with 1% of E. cloacae SK1 (0.5 McFarland standard) and incubated at 37°C for 48 h. After incubation, the samples were centrifuged at 5000 rpm for 15 min and analyzed by the dimethylglyoxime method [15-17].
2.2.3. Effect of temperature
To determine the optimal temperature for nickel biosorption, 1% of E. cloacae SK1 (0.5 McFarland standard) was inoculated into the sterile nutrient broth adjusted to pH 7, supplemented with 50 ppm of nickel, and incubated at various temperatures (18°C, 28°C, 37°C, 45°C and 55°C) for 48 h. After incubation, samples were centrifuged at 5000 rpm for 15 min, and analyzed by the dimethylglyoxime method [15-17].
2.2.4. Effect of inoculum concentration
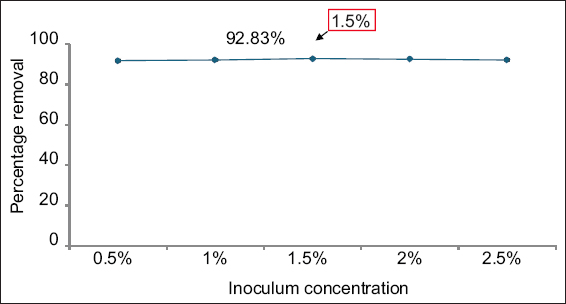
The effect of inoculum concentration on nickel biosorption was investigated using sterile nutrient broth, adjusted to pH 7 and supplemented with 50 ppm of nickel. Inoculum concentrations ranging from 0.5% to 2.5% of E. cloacae SK1 (0.5 McFarland standard) were added individually and incubated at 28°C for 48 h. After incubation, samples were centrifuged at 5000 rpm for 15 min, and analyzed by the dimethylglyoxime method.
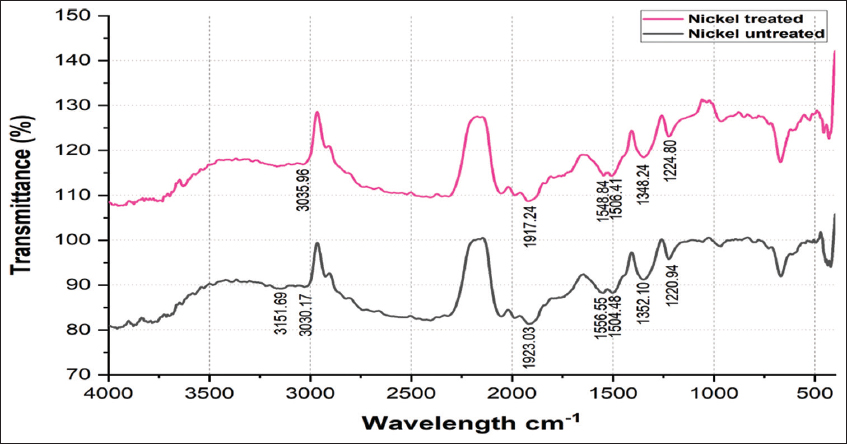
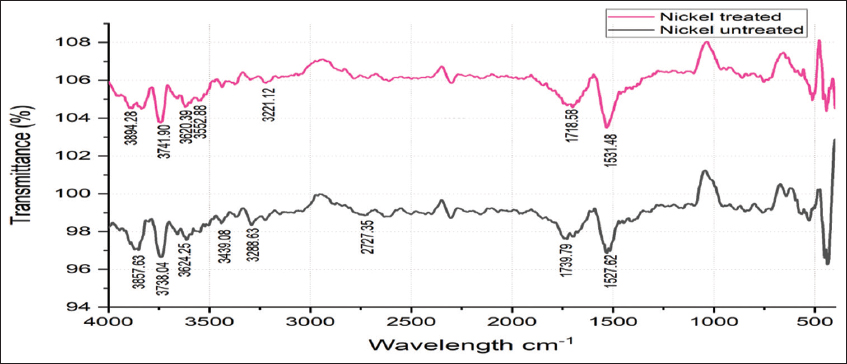
2.3. Fourier Transform Infrared Spectroscopy (FTIR) Analysis of Nickel-Untreated and Nickel-Treated Native E. cloacae SK1
E. cloacae SK1 was cultured in nutrient broth amended with 50 ppm nickel and incubated for 48 h at 28°C (test). Further, control was prepared without the addition of nickel. After incubation, the bacterial pellets were collected by centrifugation at 10,000 rpm for 10 min and resuspended in distilled water. The cell suspensions were then placed on a ZnSe crystal for drying and analyzed using FTIR spectroscopy (Shimadzu, Japan) in the wavelength range of 4000–400 cm−1 [18].
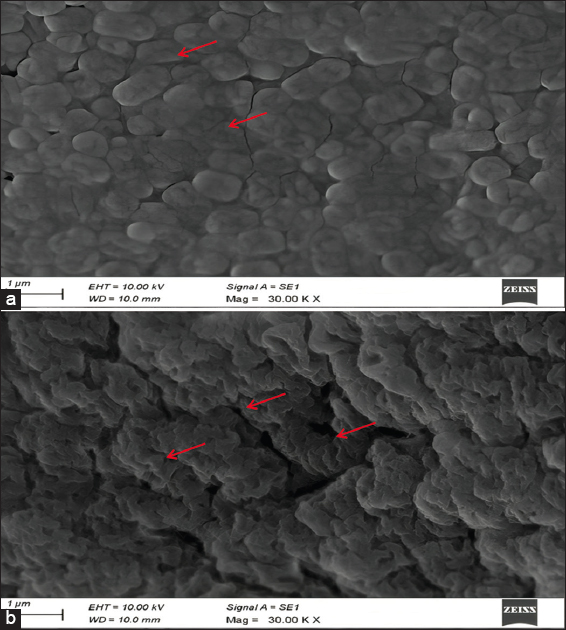
2.4. SEM Analysis of Nickel-Untreated and Nickel-Treated Native E. cloacae SK1
Morphological alterations in E. cloacae SK1 due to nickel exposure were examined using SEM. The strain was inoculated into the nutrient broth amended with 50 ppm nickel (test) and incubated for 48 h at 28°C. Further, control was prepared without the addition of nickel. After incubation, the cell cultures were centrifuged at 10,000 rpm for 10 min, washed three times with Phyto-bio system (PBS), and fixed onto a glass slide for 3 h in 3 mL of 2.5% glutaraldehyde solution. After fixation, the cells were washed 3 times with PBS and dehydrated using an ethanol gradient (30%, 50%, 70%, and 100%). The dehydrated cells were then placed on an aluminium stub for drying, mounted on a carbon-conductive sample holder, coated with platinum, and visualized under SEM (Carl Zeiss EVO 18, Germany) [19].
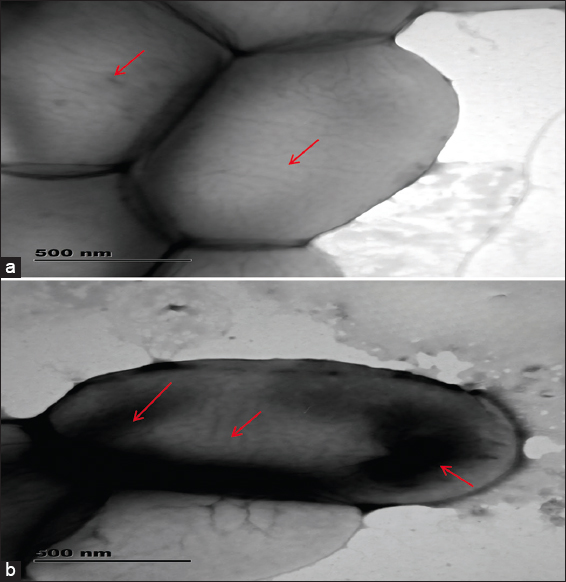
2.5. TEM Analysis of Nickel-Untreated and Nickel-Treated Native E. cloacae SK1
To investigate the intracellular accumulation of nickel in E. cloacae SK1, TEM was performed. E. cloacae SK1 cells were grown in nutrient broth containing 50 ppm nickel (test). Control was prepared without the addition of nickel. After incubation, the cells were harvested using centrifugation (10,000 rpm for 10 min), washed with PBS, and fixed for 3 h in 2.5% glutaraldehyde. After primary fixation, the samples were post-fixed with 1% osmium tetroxide at 4°C for 2 h. The fixed cells were then dehydrated through a graded ethanol series and transferred onto 200-mesh carbon-coated copper grids. After drying, the samples were examined using a TEM (JEOL JEM 2100, Japan) operated at an accelerating voltage of 200 kV [19,20].
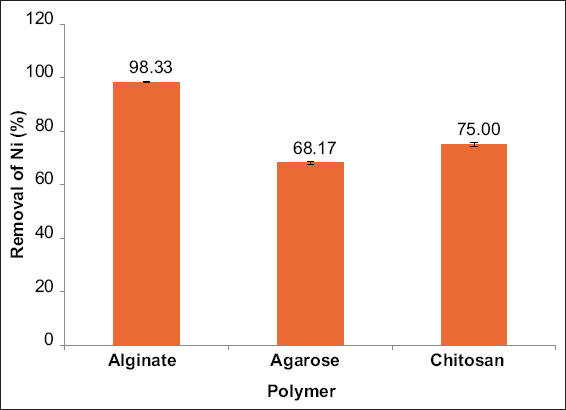
2.6. Detection of Nickel Removal Efficiency by Alginate, Agarose, and Chitosan-Immobilized Cells of E. cloacae SK1
The immobilization of active bacterial strain using polymeric substances, such as alginate, agarose, and chitosan was performed, and the nickel biosorption efficiency was determined. To prepare alginate-immobilized cells of E. cloacae SK1, 2 g of sodium alginate were dissolved in 60 mL of hot, sterile distilled water. Then, 2 g of E. cloacae SK1 biomass was added, and the total volume was adjusted to 100 mL with sterile distilled water. The resulting mixture was extruded through a 2.5 mL syringe into 100 mL of 0.2 M calcium chloride (CaCl2) solution to form uniform-sized beads [21]. Agarose beads were prepared by dissolving 2 g of agarose powder in 60 mL of hot, sterile distilled water, and 2 g of E. cloacae SK1 biomass was added. Then the volume was made up to 100 mL with sterile distilled water. The mixture was added dropwise using a 2.5 mL syringe into paraffin oil to form agarose-immobilized beads [21]. For chitosan immobilization, 2 g of chitosan were mixed with 60 mL of acetic acid and left overnight. Thereafter, 40 mL of E. cloacae SK1 cell culture was added, and the mixture was extruded through a 2.5 mL syringe into 100 mL of 0.25 mol/L sodium hydroxide (NaOH) solution to form uniform-sized chitosan beads [22]. The prepared immobilized cells were then incubated individually in 50 ppm nickel-containing nutrient broth adjusted to pH 7 and incubated for 48 h at 28°C, and their biosorption efficiency was evaluated to determine the most effective polymer for nickel removal.
2.7. FTIR Analysis of Nickel-Treated and Nickel-Untreated Alginate-Immobilized Cells of E. cloacae SK1
Alginate-immobilized E. cloacae SK1 cells were cultured in nickel-amended (50 ppm) nutrient broth and incubated for 48 h at 28°C (test). Control was prepared without the addition of nickel. After incubation, the samples were centrifuged at 10,000 rpm for 10 min to obtain alginate-immobilized cell pellets and resuspended in sterile distilled water. Further, the alginate cell suspensions were placed on a ZnSe crystal for drying and analyzed using FTIR spectroscopy (Shimadzu, Japan) in the wavelength range of 4000–400 cm−1 [18].
2.8. SEM Analysis of Nickel-Treated and Nickel-Untreated Alginate-Immobilized Cells of E. cloacae SK1
Nickel-induced morphological alterations in alginate-immobilized cells of E. cloacae SK1, which was detected by SEM. Alginate-immobilized cells were cultivated in nickel-amended (50 ppm) nutrient broth and incubated for 48 h at 28°C (test). Control was prepared without the addition of nickel. After incubation, the resulting cultures were centrifuged at 10,000 rpm for 10 min to obtain cell pellets. Then the pellets were washed 3 times with PBS solution and fixed for 3 h using 3 mL of 2.5% glutaraldehyde solution. Following fixation, the cells were washed 3 times with PBS and dehydrated through an increased gradient of ethanol solution (30%, 50%, 70%, and 100%). Further, the cells were placed on an aluminium stub for drying and mounted on a carbon-conductive sample holder coated with platinum and visualized under SEM (Carl Zeiss EVO 18, Germany) [19].
2.9. In situ Bioremediation of Nickel-Contaminated Foundry Soil
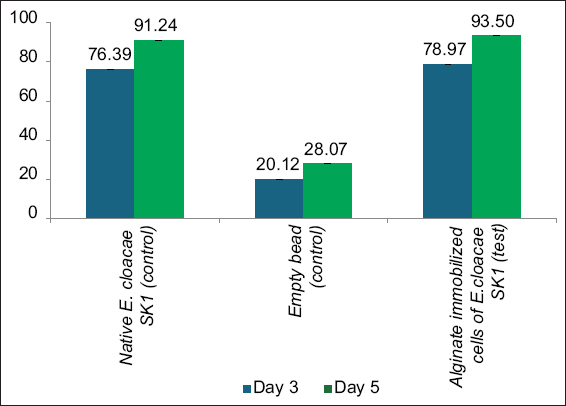
The in situ bioremediation experiment was carried out using the method proposed by Rosariastuti et al. [23] with some modifications. Three experimental plots, each measuring 50 cm × 50 cm, were created in nickel-contaminated foundry soil. The initial nickel concentration in the experimental site was determined using Atomic Absorption Spectrometry (AAS, SHIMADZU, AA-6300, Japan) before the start of the experiment. Further, two plots were served as controls: One receives 10% freshly prepared native E. cloacae SK1 (3 × 109 CFU/g), and the other plot received 10% freshly prepared alginate beads without E. cloacae SK1. The 3rd treatment plot was inoculated with 10% freshly prepared alginate-immobilized E. cloacae SK1 (3 × 109 CFU/g). Further, the soil samples were collected on the 3rd and 5th days of treatment, and the nickel concentrations were measured using AAS (SHIMADZU, AA-6300, Japan) to examine the remediation efficiency of alginate-immobilized cells [24,25].
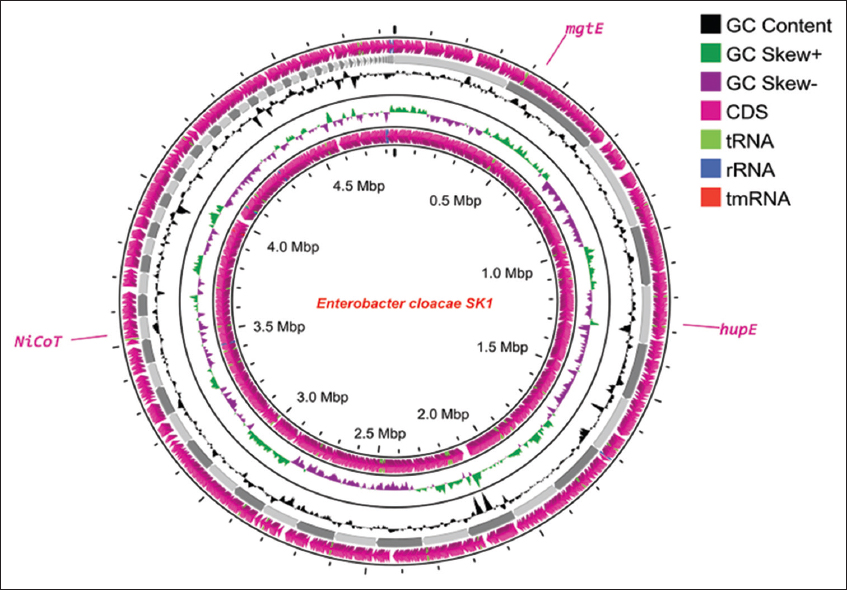
2.10. In silico Analysis: Prediction of the Nickel Resistance Genes in E. cloacae SK1
Genomic DNA from the nickel-resistant strain E. cloacae SK1 was isolated using the AlexGen bacterial DNA extraction kit (Alexius Biosciences, Germany) according to the manufacturer’s protocol. The sequencing library was constructed with the Twist NGS Library Preparation Kit (South San Francisco, CA, USA). The paired-end sequencing was conducted on an Illumina NovaSeq 6000 system (San Diego, CA, USA) [26]. In silico genome analysis of the nickel-resistant strain E. cloacae SK1 was conducted using the rapid annotation using subsystem technology (RAST) server, with SEED viewer. RAST is an online tool that facilitates the identification and functional annotation of genes within the genome, allowing the prediction of key genes involved in nickel resistance and detoxification mechanisms [27].
2.11. Statistical Analysis
One-way analysis of variance (ANOVA) was conducted using the statistical package for the biological and social sciences (SPSS) (version 16.0) to analyze the significance of the data. Further, the results are expressed as mean ± standard deviation [28].
3. RESULT AND DISCUSSION
3.1. Isolation and Determination of MIC of Nickel-Resistant Bacteria
The previously isolated and identified nickel-resistant E. cloacae SK1 (GenBank accession number – OR835983) was used for this study. E. cloacae SK1 showed an MIC of 1100 ppm for nickel, demonstrating a strong ability to tolerate nickel. Pavlic et al. [29] reported that the nickel-resistant strains Staphylococcus aureus strain ATCC 29213, Enterococcus faecalis ATCC 29212, and Escherichia coli ATCC 25922 showed an MIC of 1000 ppm for nickel. In addition, Banerjee et al. [30] reported that the nickel-resistant E. cloacae B1 showed a MIC of 700 ppm for nickel. The variation in MIC was due to the differences in the genomic characteristics of the isolates, as noted by Navarro et al. [31] and Al-Jebouri et al. [32].
3.2. Effect of Different Growth Parameters on Nickel Biosorption by E. cloacae SK1
3.2.1. Effect of contact time and initial metal ion concentration
The influence of contact time and initial metal ion concentration on nickel removal by E. cloacae SK1 was illustrated in Figure 1. At 48 h of contact time, E. cloacae SK1 removed 90.83% of 50 ppm nickel. Beyond this optimum contact time and initial nickel ion concentration, the biosorption efficiency remained constant. Similarly, Paul and Mukherjee [33] and Bisht and Kumar [34] reported that Enterobacter asburiae KUNi5 and Kluyvera cryocrescens M7, respectively, exhibited maximum nickel removal after 48 h. De Padua and Dela Cruz [35] also found that the nickel-resistant strains removed 66–68% of nickel at a concentration of 50 ppm. At lower metal ion concentrations, increased contact time enhanced the availability of active binding sites, thereby improving nickel biosorption. However, after reaching the optimum conditions, the availability of these binding sites decreased due to saturation by metal ions, leading to repulsive forces that hindered further biosorption [36,37].
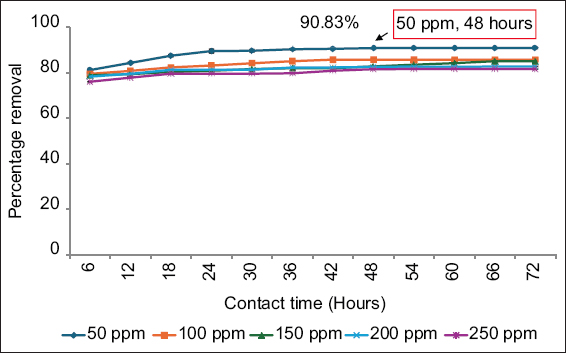 | Figure 1: Effect of contact time and initial nickel ion concentration on the removal of nickel. Data are expressed as mean ± standard deviation (n = 3), (P > 0.05). [Click here to view] |
3.2.2. Effect of pH
E. cloacae SK1 was able to remove a maximal nickel removal efficiency of 90.83% at pH 7 [Figure 2], in accordance with the result reported by Paul and Mukherjee [33]. The nickel removal efficiency was increased up to pH 7, which was due to the deprotonation of functional binding sites of the bacterial cell wall, as stated by Oves et al. [38], Al-Fakih [39], and Gupta and Jain [40]. Similarly, Taran et al. [41], Oves et al. [38] reported that nickel-resistant bacterial strains showed optimum nickel removal at pH 7. Further, under acidic conditions, the metal binding sites were closely associated with hydronium ions (H3O), which create repulsive forces that reduced metal binding efficiency. However, beyond pH 7, the formation of nickel hydroxide (Ni(OH)2) precipitates reduced the biosorption efficiency, as reported by Yu and Jiang [42].
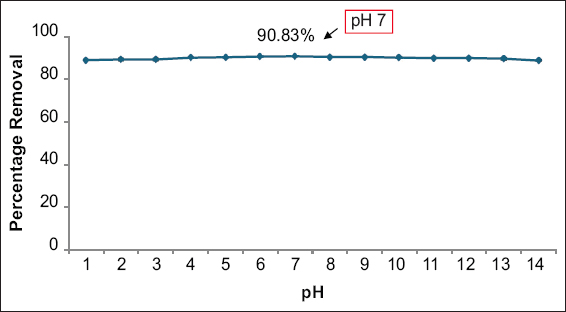 | Figure 2: Effect of pH on the removal of nickel. Data are expressed as mean ± standard deviation (n = 3), (P > 0.05). [Click here to view] |
3.2.3. Effect of temperature
E. cloacae SK1 was able to remove 92.17% of nickel at a temperature of 28°C [Figure 3]. Similar results were observed by Haque et al. [43], where metal-resistant strains, such as E. asburiae ENSD102, Enterobacter ludwigii ENSH201, Vitreoscilla sp. ENSG301, Acinetobacter lwoffii ENSG302, and Bacillus thuringiensis ENSW401 exhibited maximum nickel removal at a temperature of 28°C. Below this optimum temperature, microbial growth and enzyme activity decreased, which consequently lowered the biosorption efficiency. Moreover, above the optimal temperature, the bacterial cell membrane structure was destroyed, leading to reduced nickel removal efficiency [44].
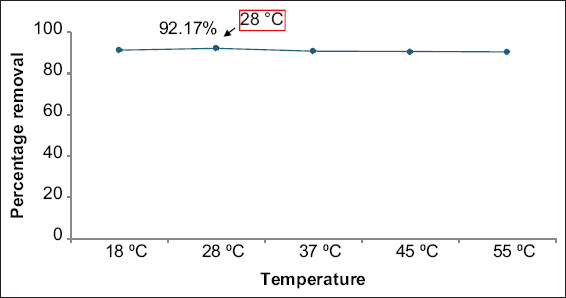 | Figure 3: Effect of temperature on the removal of nickel. Data are expressed as mean ± standard deviation (n = 3), (P > 0.05). [Click here to view] |