1. INTRODUCTION
The term machine learning was conceived by Arthur Lee, Samuel, a pioneer in computer gaming and artificial intelligence at IBM and Stanford in 1959 [1]. It provides computational systems the ability to spontaneously learn and improve from the previous experience without being explicitly programmed [1]. Machine learning is now practiced in an expansive range of practical purposes that include email spam filtering [2], forecasting GPS signals [3], 5G network management [4], Indian classical dance action identification [5], self-driving cars [6], sign language recognition [7], speech recognition [8], stock market prediction [9], thermal wave image quantification [10] and traffic video surveillance [5], and web-searches [11]. Machine learning methods are broadly categorized into supervised and unsupervised methods. Supervised methods are trained on examples with labels and are then used to predict these labels on some other examples, whereas unsupervised methods find patterns in data sets without the use of labels. Machine learning algorithms can be broadly categorized as classification (decision tree and naive Bayes’s classifier), clustering (k-mean and hierarchical), regression (least-squares and logistic regression), dimensionality reduction (principal component analysis (PCA) and multidimensional scaling (MDS), and artificial neural networks (perceptron, back-propagation, and self-organizing map (SOM)). These machine learning methods and algorithms now have become an integral tool for analyzing biological data.
Biological information is broadly categorized into the digital code of the genome and the environmental cues that emerge externally to the genome [12]. This information can be deciphered by automated technologies, to name a few, next-generation sequencing, DNA microarrays, mass spectroscopy, X-ray crystallography, time-lapse microscopic imaging technologies, and quantitative tagged magnetic resonance imaging. These technologies provide information ranging from genome to phoneme. The contemporary research focuses on data derived from these technologies in the context of quantitative data-based mathematical models of protein-protein interactions, metabolic networks, cellular signaling cascades, and cell-to-cell communication. Machine learning algorithms provide excellent opportunities to understand intricate biological systems and now it has been already a mainstream tool of life sciences. Machine learning has been employed for the prediction of protein sorting signals and signals [13], cancer prognosis and prediction [14], predicting chronic diseases such as Alzheimer’s ailments, and diabetes [15], classification of malaria disease [16], empirical studies of cancers [17], genetics and genomics [18], brain imaging [19], tumor detection [20], and medicine [21]. Unveiling current trends in the application of machine learning in life science allow biologists to improve their research, understand and implement new algorithms, and evolve as interdisciplinary scientists over time. Bibliometric analysis is an evolving branch of science applied to measure progress in several fields of science, a systematic investigation of publication trends. Bibliometric analysis can be carried out with two approaches [22]. The first approach of bibliographic data analysis includes evaluating the performance of published publications, specifically assessing scientific actors such as countries, institutions, departments, and researchers. Another type of science mapping is the extraction of knowledge from the intellectual, social, or conceptual architecture of a research arena that could be done employing science mapping analysis based on bibliographic networks. In this study, we focused majorly on the second aspect of bibliographic networks, that is, knowledge extraction to quantify the thematic evolution of machine learning concepts in life sciences based on the research information available in the Medline database.
2. MATERIALS AND METHODS
2.1. Datasets
To map the intellectual structure of machine learning in life science research, we have carried out a coword analysis using Medical Subject Headings (MeSH) terminology of the National Library of Medicine controlled vocabulary thesaurus for the title and abstract of PubMed indexed articles. PubMed’s advanced search strategy “machine learning” [Title] was employed to retrieve the citations from the Medline database. We retrieved 9162 articles from Medline, the primary bibliographic database of the U.S. National Library of Medicine (NLM). The majority of journals were selected for MEDLINE based on the recommendation of the literature selection technical review committee led by life scientists. Furthermore, MEDLINE is an open-access database and specific to high-quality literature in biomedicine. Here, we selected the stringent criterion for choosing the dataset which may not cover all themes associated with the field. Further articles were downloaded separately for four different time frames, 1964–2010, 2011–2015, 2016–2018, and 2019–2020. The numbers of articles are 551, 968, 2702, and 5293, respectively, for 1964–2010, 2011–2015, 2016–2018, and 2019–2020. All results were saved in PubMed text format.
2.2. Implementation of Bibliometric Analysis
The science mapping analysis was carried out using the Bibliometrix R package [23] and the VOSviewer tool [24]. PubMed text files were processed using the Bibliometrix tool for extracting pieces of information for publications, authors, journals, institutes, and countries. VOSviewer has been exploited for constructing a bibliometric network of coauthors and knowledge extraction from cooccurring words. Cooccurrences of two keywords are defined as the number of publications in which both keywords occur together in the abstract, title, or keyword list [25].
To delve into the evolution of the intellectual structure of the field, we classified the themes as quiescent, basic, or evolving. We strategized the bibliographic network analysis for four consecutive time periods 1964–2010, 2011–2015, 2016–2018, and 2019–2020. We have chosen the periods based on the number of publications with at least 500 in each period.
3. RESULTS
3.1. Evaluation of Scientific Actors
The global research impact of machine learning usage in life science research during the time 1964 to September 7, 2020 led to 9433 publications. The year-wise research publication trends are shown in Figure 1a. The trends indicate that machine learning usage is rapidly incorporated in bioscience research. The works have been published in 2052 journals and book sources. Top 10 publishing journals include PloS one, Scientific reports, Sensors (Basel, Switzerland), BMC Bioinformatics, Journal of chemical information and modeling, Bioinformatics (Oxford, England), Studies in health technology and informatics, Conference proceedings: Annual International Conference of the IEEE Engineering, The Journal of chemical physics, and Computer methods and programs in biomedicine [Figure 1b]. Twenty-seven thousand three hundred and sixty-four organizations and 42,629 unique authors were involved with machine learning research in life sciences. Six thousand three hundred and forty-four authors published at least two articles, 2005 with three, 920 with four, 495 with five, and 53 authors with at least 10 publications. The most contributing authors are from the USA, followed by China [Figure 1c]. Notable researchers with a minimum of 20 publications with collaborations are Sean Ekins (University of Maryland), Klaus-Robert Muller (Max-Planck-Institute for Informatics), O. Anatole von Lilienfeld (University of Vienna), and Wei Wang (National University of Singapore). For every 495 authors, the total strength of the coauthorship links was calculated using the VOSviewer tool. The authors with the greatest total link strength (183 items) are chosen and shown in Figure 1d. The mean collaboration index for machine learning in the biosciences theme is 3.96, which indicates the intricacy of this research theme and hence the quality of the output publication works.
 | Figure 1: Scientific factors of machine learning in biosciences (a). Global publication trends (b). Most contributing journal sources (c). The number of authors versus author affiliation country (d). Coauthor interaction maps for 183 items. [Click here to view] |
3.2. Evaluation of the Conceptual Structure of Machine Learning in Biosciences
Next, we employed keyword analysis to discern the trends, using preferred statistical approaches, developed, fields of life sciences, and tools and technologies. The cooccurrence of Mesh terms with a full counting strategy suggests that the most occurring keywords are linked to machine learning algorithms and statistical terms, human age-related words, and imaging technologies [Figure 2a]. Five thousand one hundred and seventy-four keywords were obtained while five is set as the minimum number of occurrences of keywords. However, it reduced to 277 when 20 occurrences were considered. The most observed keywords include human, machine learning, female, male, and middle-aged, adult, aged, AI, and SVM [Figure 2a]. Both supervised and unsupervised learning are extensively applied in life science research. For instance, we computed the number of publications machine learning relevant keywords in their title or abstracts [Figure 2b]. The most widely used supervised techniques are support vector machine (1543), neural network (1018), decision tree (437), random forest (376), convolutional neural networks (118), linear discriminant analysis (117), AdaBoost (85), and recurrent neural network (36). Here, the number in parentheses indicates the number of publications with at least one keyword. Unsupervised methods k-means clustering (54) and principal component analysis (205) have also been utilized but to a lesser extent when compared to supervised methods.
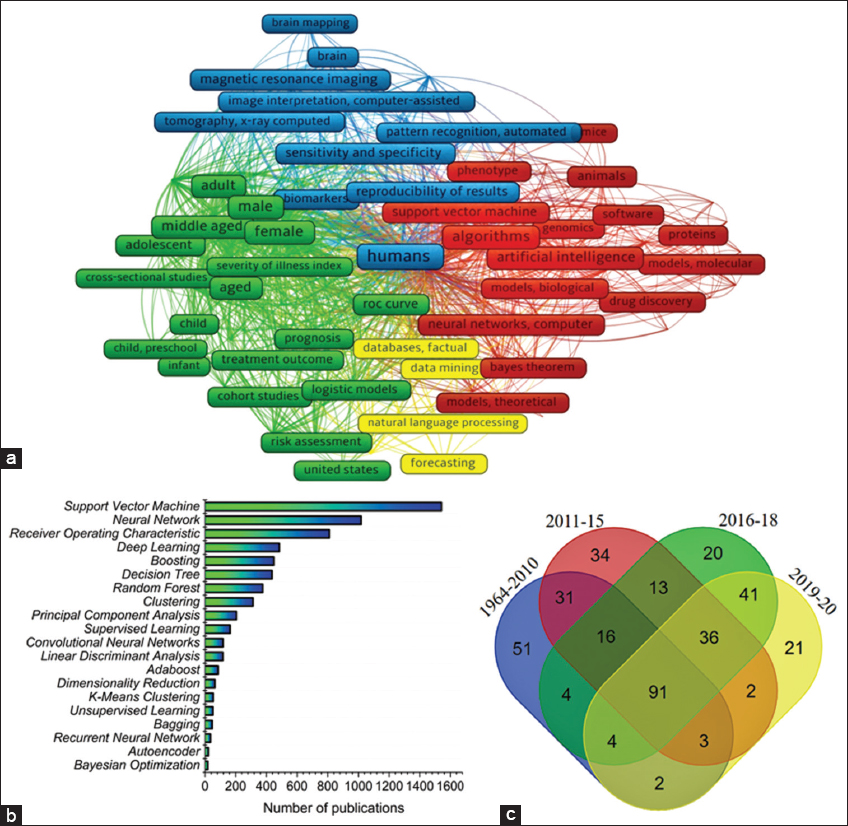 | Figure 2: Machine learning concepts in biosciences. (a) Coword network for the most enriched keywords. Colors indicate clustered items and the curved line indicates the connections among the words. (b) Most occurring machine learning concepts in publications of time frame 1964–2020. (c) Venn diagram indicating the top 200 keywords in four different periods, 1964–2010, 2011–2015, 2016–2018, and 2018–2020. Ninety-one keywords indicate core research theme topics and 51 words may be part of quiescent concepts while 21 may reflect emerging topics. [Click here to view] |
Next, we investigated the evolution of the intellectual structure of the field, for four consecutive time periods 1964–2010, 2011–2015, 2016–2018, and 2019–2020. Furthermore, we loosely categorized the conceptual themes into four categories: 1. Machine learning concepts, 2. molecular biology and biomedicine concepts, 3. tools, methods, and technology concepts, and 4. disease concepts. It should be noted that some of the keywords may overlap with other concepts. The top 200 occurring keywords were chosen for establishing the thematic evolution of the concepts. The concepts may be regarded as quiescent, basic, or evolving. In general, basic conceptual themes will be common for all time periods, while some display emerging trends with a recent period of publication. In general, the keywords conserved in all four timelines are considered as core concepts, and keywords occurring in recent years are regarded as emerging topics. Quiescent themes only exist in older time periods. The top 200 MeSH words identified using VOSviewer were indicated in the Venn diagram [Figure 2c].
3.2.1. Conserved and Basic Concepts
The 91 keywords are enriched in all 4-time frames 1964–2010, 2011–2015, 2016–2018, and 2019–2020 that may be considered as core basic concepts [Figure 2c]. The common research terminology associated with machine learning research in life sciences is as follows.
1. Machine learning keywords: Area under a curve, artificial intelligence, Baye’s theorem, linear models, logistic models, principal component analysis, sensitivity and specificity, regression analysis, natural language processing, neural networks, cluster analysis, computer simulation, data mining, databases, decision trees, forecasting, multivariate analysis, pattern recognition, and fuzzy logic.
2. Molecular biology and biomedicine keywords: Adolescent, adult, aged, aged 80 and over, infant, middle-aged, child, young adult, pregnancy, amino acid sequence, animals, female, humans, mutation, protein binding, protein conformation, structure-activity relationship, image interpretation, computer-assisted image processing, the severity of illness index, biomarkers, tumor, and biomechanical phenomena.
3. Tools, methods, and technology keywords: Magnetic resonance imaging, pattern recognition, single nucleotide polymorphism, electroencephalography, gene expression profiling, genomics, protein diagnosis, computer-assisted, drug design, drug discovery, and feasibility studies.
4. Disease Alzheimer’s disease, disease progression, prostatic neoplasms, neoplasms, breast neoplasms, prognosis, HIV infection, optic nerve diseases, toxicity tests, skin neoplasms, vision disorders, dementia, and glaucoma.
Based on the coword analysis of the keywords associated with the 4-time frames, it is revealed that gene expression profiling, medical imaging, drug design related, and disease-associated concepts form the core concepts of the theme machine learning research in biosciences.
3.2.2. Declining or Quiescent Topics
As the research progress, some concepts may become less used or become obsolete. Here, we loosely categorized the topic which is only represented in the older time frame 1964–2010 and 1964–2015 as quiescent or declining topics in the context of machine learning applications in life sciences. The tools or methods, namely, quality control, matrix-assisted laser desorption ionization, chromosome mapping, and Fourier transform infrared spectroscopy, have become obsolete as many other high-throughput technologies have evolved. It is revealed that on the machine learning related concepts, terminology as a topic, databases as the topic, evaluation, studies as a topic, database management systems, computerized medical records systems, unified medical language system, and electronic data processing, significant research has not been carried out. Furthermore, systems biology, which is a hot topic during the 2000s, now becomes underrepresented in machine learning research. However, the research may be stagnated or at a slower pace beyond 2015.
The major focus of machine learning applications up to 2015 has been on basic molecular biology concepts and methods. The key topics identified include protein sequence alignment, secondary and tertiary structure prediction, and proteomic data analysis using pattern recognition methods. However, from 2016, the theme expanded to utilize medical imaging datasets. It should be noted that artificial intelligence is a general terminology used for machine learning tools and algorithms [Figure 3a].
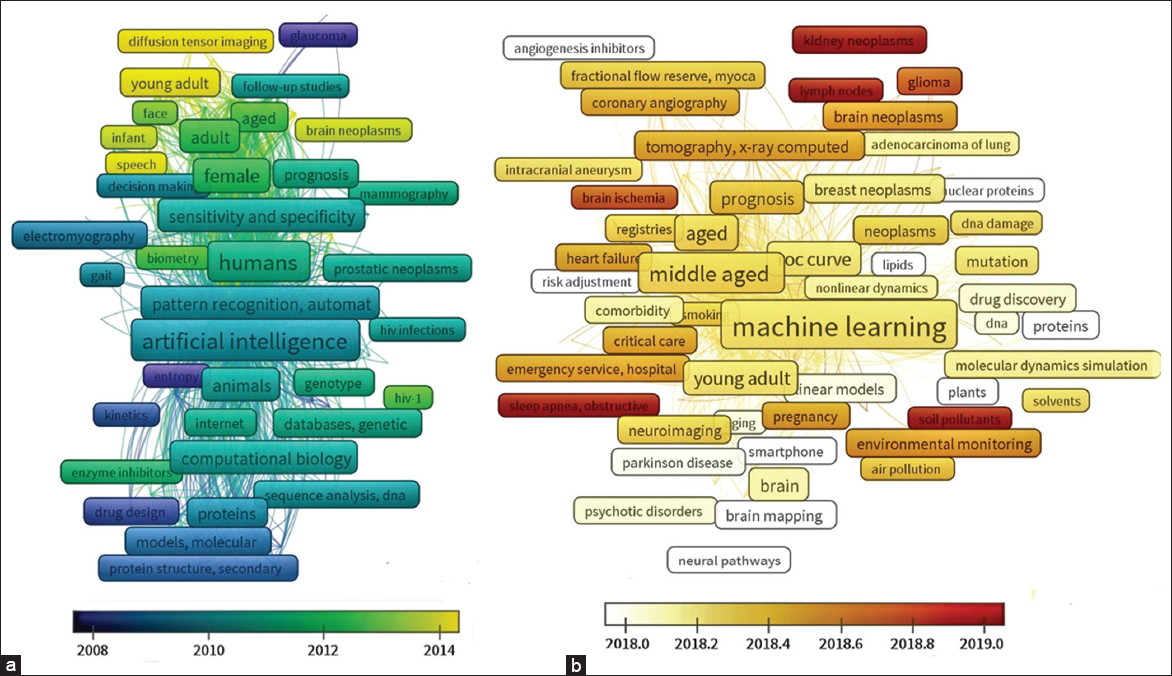 | Figure 3: (a and b) Thematic evolution of machine learning applications in life sciences: Coword analysis for 2-time spans 1964–2015 and 2016–2020 was plotted using VOS viewer. The colors of keywords represent their occurrence as the average publication year. [Click here to view] |
3.2.3. Evolving concepts
During the 2016–20 period, expansive growth of machine learning literature is observed with the evolution of many concepts. The application of machine learning has expanded to new fields in specific medical imaging technologies [Figure 3b]. The commonly enriched concepts for 2016–18 and 2019–20 were listed as follows.
1. Machine learning keywords: Supervised machine learning, unsupervised machine learning, deep learning, and big data.
2. Molecular biology and biomedicine keywords: Precision medicine, radiology, registries, intensive care units, delivery of health care, early diagnosis, exercise, hospital mortality, hospitalization, survival rate, and social media.
3. Tools, methods, and technology keywords: Accelerometry, clinical decision-making, cognition, computed tomography angiography, coronary angiography, coronary stenosis, cross-sectional studies, fractional flow reserve, high-throughput nucleotide sequencing, and wearable electronic devices.
4. Disease Keywords: Antineoplastic agents, cardiovascular diseases, myocardial, glioma, heart failure, mental disorders, neoplasm staging, comorbidity, and sepsis.
The research is more evolved to apply machine learning in the clinical field with an avalanche of publications related to precision medicine, clinical decision-making, heart imaging technology, a multitude of diseases, and physical activity measurements. It indicates that machine learning applications extend from analyzing genomic-related information to clinical and physiological measurement datasets. Machine learning concepts, big data analytics, and deep learning (DL) are also adopted by bioscientists and have become promising tools. It is observed that these tools are majorly applied for image-based diagnostic systems. An interesting observation in evolving concepts includes machine learning research relevant to coronavirus disease 2019 (COVID-19). COVID-19 is an emerging, rapidly evolving pandemic crisis. Based on the bibliographic trends of 2019–20, machine learning has been extensively applied to research on the COVID-19 theme. Figure 4 indicates the coword analysis for a time span of 2 years 2019–2020. Emerged and unique keywords relevant to COVID-19 in this period include beta coronavirus, coronavirus infection, viral pneumonia, pandemics, length of stay, air pollutants, atrial fibrillation, brain ischemia, diabetes mellitus, kidney neoplasms, carcinoma, renal cell, clinical, decision rules, data analysis, drug repositioning, incidence, neoplasm grading, patient selection, postoperative complications, socioeconomic factors, swine, and triage. Altogether, an evaluation of the conceptual structure of machine learning in biosciences revealed trends in quiescent, basic, or evolving themes.
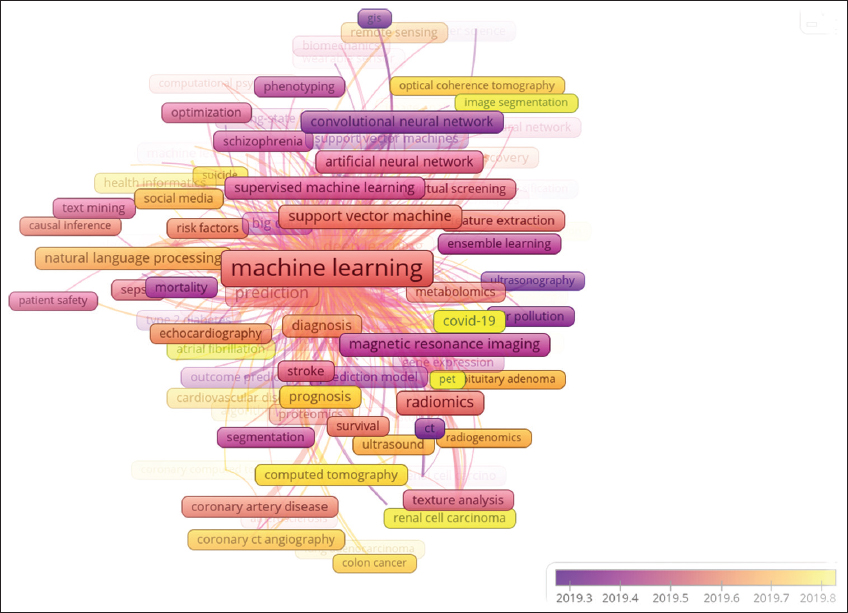 | Figure 4: Machine learning application in the COVID-19 theme. Coword analysis for the time span 2019–2020 indicates that machine learning has been extensively applied to research on the COVID-19 theme. [Click here to view] |
4. DISCUSSION AND CONCLUSIONS
Delving into the avalanche of heterogeneous omics data derived from high-throughput experiments, encompassing “atomic” to “ecosystem” scales are regarded as a great challenge in biology. Bibliometric analysis is an evolving branch of science applied to quantify research progress through systematic investigation of publication trends. Earlier research work of bibliometric analysis for machine learning research has been carried out to provide an overview of the scientific work [23], capture the intellectual structure [24], and scientometrics [26] of the field. Machine learning bibliometric analysis has also been reported in investigating research trends in public health problems [27], mental health in social media [28], COVID-19 [29], and biomedicine [30]. However, the general trend of machine learning bibliometric analysis in life sciences has not been reported. In this survey, we depicted a global picture of the machine learning application in biosciences based on PubMed indexed research publications. We reported the results of the bibliometric analysis with two approaches, namely, scientific actors and conceptual architecture. Assessment of scientific actors, namely, countries, organizations, and researchers, indicated that the output publication work is high quality in nature. Notable publishing journals include PloS one, Scientific reports, Sensors (Basel, Switzerland), BMC Bioinformatics, Journal of chemical information and modeling, and Bioinformatics. The mean collaboration index observed in this study (3.96) often regarded as good quality research scale [31]. After 2012, the rate of publication increased exponentially. A rapid surge in publications may be attributed to breakthroughs such as deep learning, amplified computational power, and storage, improved high-throughput technologies in biosciences [32]. Next, we rigorously reported the second aspect of bibliographic networks, that is, knowledge extraction to quantify the thematic evolution. The two divisions of machine learning techniques supervised learning (support vector machine, neural network, decision tree, random forest, convolutional neural networks, linear discriminant analysis, and recurrent neural network) and unsupervised learning (k-means clustering, and principal component analysis) were extensively used in the life sciences. Our results are in line with a similar bibliometric analysis on artificial intelligence in health care [33]. The authors revealed that artificial neural networks, support vector machines, and convolutional neural networks have more influence on health-care research [33]. Furthermore, a longitudinal cohort bibliographic coupling analysis for 4-time periods, 1964–2010, 2011–2015, 2016–2018, and 2019–2020 captured quiescent, basic, or evolving topics in the conceptual structure of this field. Notable quiescent topics (for which research may be stagnated or at a slower pace) observed are matrix-assisted laser desorption ionization, chromosome mapping, Fourier transform infrared spectroscopy, unified medical language system, electronic data processing, and systems biology. Expectedly, gene expression profiling, medical imaging, drug design related, and disease-associated concepts are conserved in 4-time frames and form the core concepts. Evolved research happened in the clinical field with several research publications in concepts, precision medicine, clinical decision-making, heart imaging technology, a multitude of diseases, and physical activity measurements. A significant observation in our study is that the application of machine learning on COVID-19 datasets. Bibliographic trends of 2019–20 revealed that machine learning has been extensively applied to research on the COVID-19 theme. Machine learning has been applied for analysis on novel coronavirus (COVID-19) [34], differentiating novel coronavirus pneumonia from general pneumonia [35], severity detection for the coronavirus disease 2019 (COVID-19) [36], vaccine design using reverse vaccinology [37], modeling the trend of coronavirus disease spread [38], and proteins structures [39], etc. In summary, our results indicated that during the initial period, machine learning was employed for pattern recognition in biosequences, followed by structural biology and drug design. Recently, the trend moved to personalized medicine and high-throughput medical imaging and big data analytics. The knowledge of machine learning evolving trends provides opportunities to conduct good research on advancing big data sources with velocity, variety, and veracity characteristics. Unveiling current trends in the application of machine learning in life science allow biologists to improve their research, understand and implement new algorithms, and evolve as interdisciplinary scientists over time.
5. AUTHORS’ CONTRIBUTIONS
All authors made substantial contributions to conception and design, acquisition of data, or analysis and interpretation of data; took part in drafting the article or revising it critically for important intellectual content; agreed to submit to the current journal; gave final approval of the version to be published; and agreed to be accountable for all aspects of the work. All the authors are eligible to be an author as per the International Committee of Medical Journal Editors (ICMJE) requirements/guidelines.
6. FUNDING
The authors report no funding source for the work.
7. CONFLICTS OF INTEREST
The authors report no financial or any other conflicts of interest in this work.
8. ETHICAL APPROVALS
This computational study does not requires any kind of ethical approvals.
9. DATA AVAILABILITY
Derived data supporting the findings of this study are available from the corresponding author (V.R.Y) on request.
10. PUBLISHER’S NOTE
This journal remains neutral with regard to jurisdictional claims in published institutional affiliation.
REFERENCES
1. Samuel AL. Some studies in machine learning using the game of checkers. IBM J Res Dev 1959;3:210-29. [CrossRef]
2. Guzella TS, Caminhas WM. A review of machine learning approaches to spam filtering. Expert Syst Appl 2009;36:10206-22. [CrossRef]
3. Srivani I, Prasad GS, Ratnam DV. A deep learning-based approach to forecast ionospheric delays for GPS signals. IEEE Geosci Remote Sens Lett 2019;16:1180-4. [CrossRef]
4. Bashir AK, Arul R, Basheer S, Raja G, Jayaraman R, Qureshi NM. An optimal multitier resource allocation of cloud RAN in 5G using machine learning. Trans Emerg Telecommun Technol 2019;30:e3627. [CrossRef]
5. Appathurai A, Sundarasekar R, Raja C, Alex EJ, Palagan CA. Nithya A. An efficient optimal neural network-based moving vehicle detection in traffic video surveillance system. Circuits Syst Signal Proc 2020;39:734-56. [CrossRef]
6. Bojarski M, Del Testa D, Dworakowski D, Firner B, Flepp B, Goyal P, et al. End to end learning for self-driving cars. arXiv preprint arXiv:1604.07316;2016.
7. Rao GA, Kishore PV, Sastry AS, Anil Kumar D, Kiran Kumar E. Selfie continuous sign language recognition with neural network classifier. In:Proceedings of 2nd International Conference on Micro-Electronics, Electromagnetics and Telecommunications. Berlin, Germany:Springer;2018.
8. Liu SS, Tian YT. Facial expression recognition method based on gabor wavelet features and fractional power polynomial kernel PCA. In:International Symposium on Neural Networks. Berlin, Germany:Springer;2010. [CrossRef]
9. Nadh VL, Prasad GS. Stock market prediction based on machine learning approaches. In:Computational Intelligence and Big Data Analytics. Berlin, Germany:Springer;2019. 75-9. [CrossRef]
10. Lakshmi AV, Muzammil VG, Parvez M, Subhani SK, Ghali VS. Artificial neural networks based quantitative evaluation of subsurface anomalies in quadratic frequency modulated thermal wave imaging. Infrared Phys Technol 2019;97:108-15. [CrossRef]
11. Pazzani M, Billsus D. Learning and revising user profiles:The identification of interesting web sites. Mach Learn 1997;27:313-31. [CrossRef]
12. Hood L, Heath JR, Phelps ME, Lin B. Systems biology and new technologies enable predictive and preventative medicine. Science 2004;306:640-3. [CrossRef]
13. Nielsen H, Brunak S, von Heijne G. Machine learning approaches for the prediction of signal peptides and other protein sorting signals. Protein Eng 1999;12:3-9. [CrossRef]
14. Kourou K, Exarchos TP, Exarchos KP, Karamouzis MV, Fotiadis DI. Machine learning applications in cancer prognosis and prediction. Comput Struct Biotechnol J 2015;13:8-17. [CrossRef]
15. Changala R, Rao DR. Development of predictive model for medical domains to predict chronic diseases (diabetes) using machine learning algorithms and classification techniques. ARPN J Eng Appl Sci 2019;14:1202-12.
16. Sajana T, Narasingarao M. An ensemble framework for classification of malaria disease. ARPN J Eng Appl Sci 2018;13:3299-307.
17. Sivakumar S, Nayak SR, Vidyanandini S, Jayaraman AK, Palai P. An empirical study of supervised learning methods for breast cancer diseases. Optik 2018;175:105-14. [CrossRef]
18. Libbrecht MW, Noble WS. Machine learning applications in genetics and genomics. Nat Rev Genetics 2015;16:321-32. [CrossRef]
19. Lemm S, Blankertz B, Dickhaus T, Müller KR. Introduction to machine learning for brain imaging. Neuroimage 2011;56:387-99. [CrossRef]
20. Patil JS, Pradeepini G. Brain tumor levels detection in three dimensional MRI using machine learning and MapReduce. Indian J Public Health Res Dev 2019;10:1465-71. [CrossRef]
21. Rajkomar A, Dean J, Kohane I. Machine learning in medicine. N Engl J Med 2019;380:1347-58. [CrossRef]
22. Gutiérrez-Salcedo M, Ángeles Martínez M, Moral-Munoz JA, Herrera-Viedma E, Cobo MJ. Some bibliometric procedures for analyzing and evaluating research fields. Appl Intell 2018;48:1275-87. [CrossRef]
23. Rincon-Patino J, Ramirez-Gonzalez G, Corrales JC. Exploring machine learning:A bibliometric general approach using SciMAT. F1000Research 2018;7:1210. [CrossRef]
24. Bhattacharya S. Some salient aspects of machine learning research:A bibliometric analysis. J Sci Res 2019;8:s85-92. [CrossRef]
25. Van Eck NJ, Waltman L. Visualizing bibliometric networks. In:Measuring Scholarly Impact. Berlin, Germany:Springer;2014. 285-320. [CrossRef]
26. Ho YS, Wang MH. A bibliometric analysis of artificial intelligence publications from 1991 to 2018. COLLNET J Scientometrics Inf Manag 2020;14:369-92. [CrossRef]
27. dos Santos BS, Steinera MT, Fenericha MT, Lima RH. Data mining and machine learning techniques applied to public health problems:A bibliometric analysis from 2009 to 2018. Comput Ind Eng 2019;138:106120. [CrossRef]
28. Kim J, Lee D, Park E. Machine learning for mental health in social media:Bibliometric study. J Med Internet Res 2021;23:e24870. [CrossRef]
29. De Felice F, Polimeni A. Coronavirus disease (COVID-19):A machine learning bibliometric analysis. In Vivo 2020;34 3 suppl:1613-7. [CrossRef]
30. Shukla N, MerigóJM, Lammers T, Miranda L. Half a century of computer methods and programs in biomedicine:A bibliometric analysis from 1970 to 2017. Comput Methods Programs Biomed 2020;183:105075. [CrossRef]
31. Stallings J, Vance E, Yang J, Vannier MW, Liang J, Pang L, Dai L, et al. Determining scientific impact using a collaboration index. Proc Natl Acad Sci 2013;110:9680-5. [CrossRef]
32. Topol EJ. High-performance medicine:The convergence of human and artificial intelligence. Nat Med 2019;25:44-56. [CrossRef]
33. Guo Y, Hao Z, Zhao S, Gong J, Yang F. Artificial intelligence in health care:Bibliometric analysis. J Med Internet Res 2020;22:e18228. [CrossRef]
34. Yadav M, Perumal M, Srinivas M. Analysis on novel coronavirus (COVID-19) using machine learning methods. Chaos Solitons Fractals 2020;139:110050. [CrossRef]
35. Liu C, Wang X, Liu C, Sun Q, Peng W. Differentiating novel coronavirus pneumonia from general pneumonia based on machine learning. Biomed Eng Online 2020;19:66. [CrossRef]
36. Yao H, Zhang N, Zhang R, Duan M, Xie T, Pan J, et al. Severity detection for the coronavirus disease 2019 (COVID-19) patients using a machine learning model based on the blood and urine tests. Front Cell Dev Biol 2020;8:683. [CrossRef]
37. Ong E, Wong MU, Huffman A, He Y. COVID-19 coronavirus vaccine design using reverse vaccinology and machine learning. Front Immunol 2020;11:1581. [CrossRef]
38. Liu Z, Huang S, Lu W, Su Z, Yin X, Liang H, et al. Modeling the trend of coronavirus disease 2019 and restoration of operational capability of metropolitan medical service in China:A machine learning and mathematical model-based analysis. Glob Health Res Policy 2020;5:1-11. [CrossRef]
39. Heo L, Feig M. Modeling of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) proteins by machine learning and physics-based refinement. bioRxiv 2020;2020:8904. [CrossRef]