1. INTRODUCTION
Woodfordia fruticosa (Linn.) Kurz (family: Lythraceae) is commonly called “Dhainyaro” and is housed from 200 to about 1,800 m above the ocean surface. Therapeutic values for its various parts are reported [1] with commercial formulations incorporating them. It has, thus, evoked significant curiosity in the scientific community to appraise the therapeutic properties with scientific rationality for its use in medical therapy. There are a few reports on the use of flowers and leaves exhibiting experimentally proved anti-inflammatory, antitumor, anti-leucorrhoeic, and immunomodulatory activities. The biological compounds reported in parts like flowers, leaves, and stem of W. fruticosa have been credited to provide therapeutic benefits [2–4].
Endophytic microbes of medicinal flora have gained awareness as they are accepted to be magnificent assets of secondary metabolites endowed with biological capacity [5–10]. As the secondary metabolites produced by fungal endophytes have significance in both academic and industrial scenarios, the study of W. fruticosa-associated fungi yields wider appreciation toward its diversity. As a first report about the biological activities of the extract from endophytic fungi residing in W. fruticosa leaves, this study reports the fungus Mucor souzae as a new source with the ability to yield molecules with application in pharmacy.
2. MATERIALS AND METHODS
2.1. Plant Samples
Leaf samples of W. fruticosa (Linn.) Kurz were gathered from Gokarna, which is along the west coast of India, in the taluk of Kumta, and is in the district of Uttara Kannada of Karnataka state. The herbarium specimen has been deposited (W. fruticosa # IOE LP0006). The specimens were placed in plastic containers, sorted during a stipulated time period (24–48 hours), and were processed for growth and analysis of fungal endophytes.
2.2. Culturing of Leaf Endophytes
The leaves collected were cleaned under tap water (five times), resulting in the clearing of dust and grime particles. The sterilization of cleaned leaf surface was carried out under sterile conditions. The steps involved several rinses with ethanol (70%, v/v) for a minute. The next step was cleaning with sodium hypochlorite (3 minutes, containing 3% accessible chlorine; 4.5% v/v diluted; HiMedia, India). Lastly, a rinse was given using distilled water (sterile), resulting in the removal of any remnants of sterilants on the surface [11]. All leaf bits were then tapped, resulting in the removal of adhering water droplets. Leaf bits (10–15 bits; 0.5–1 cm) were placed in media (sterile water agar; 15 g.l−1) containing chloramphenicol (240 ppm; Sigma Aldrich, St. Louis, MO) in a sterile petri dish. Incubation was for 15 days for 16 hours (28°C; dark alternate with light cycles). The fungal endophytes sprouting from the bits of leaves were photographed. Furthermore, they were then placed on potato dextrose agar (PDA) media (HiMedia, India) to induce maturation, including sporulation. Then the grown fungal endophytes were stored (−80°C) on PDA and 15% glycerol (v/v) in cryovials.
2.3. Isolation of Genomic DNA from Fungal Endophytes, its PCR Amplification, and Molecular Typing
Initially, the fungal endophytes were identified depending on their colony characteristics. Their spores and fruiting bodies were photographed under the microscope (Research Stereo Zoom Microscope; Stereo Discovery V20; Carl Zeiss, Germany). Furthermore, typing was carried out as reported in standard manuals specifically for fungal identification [11]. Endophytic fungi were cultivated on PDA media and the fresh culture was harvested for further work. The DNA from this culture was extracted by cetyltrimethylammonium bromide protocol [12,13]. DNA was kept in a buffer (Tris EDTA; 100 μ l). Quantification of the genomic DNA and its purity were checked in a nano-spectrophotometer (Thermo 2000C; Thermo Fisher Scientific, USA). Corresponding to pure desirability (1.6/1.8) indicated no contamination with ribose nucleic acid, protein or contaminant, phenol. The DNA in the resultant sample was confirmed in 1% agarose gel electrophoresis and the results were documented (Geldoc XRT; BioRad, USA). Universal internal transcribed spacer (ITS) primers for fungi were used to amplify the extracted DNA (ITS1: 5?-TCC GTAGGTGAACCTGCG G-3?; ITS4: 5?-TCCTCCGCTTATTGATATGC-3?) [14]. The reaction was carried out in polymerase chain reaction (PCR) tubes (0.2 ml) containing the reaction mixture (25 μ l) and genomic DNA (1 μ l; 50 ng/μ l) in a thermal cycler (Mastercycler gradient; Eppendorf, Germany). The program was as follows: initial denaturation (5 minutes, 95°C followed by 3 minutes, 94°C) and primer annealing (1 minute, 55°C), followed by extension (2 minutes, 72°C and 10 minutes extension), and it was repeated for 35 cycles. The product of this PCR amplification (5μ l) was assessed on 1% agarose gel electrophoresis and the results were reported in comparison to a commercially available standard molecular ladder. The amplified PCR samples were sequenced at Bangalore (Chromous Biotech). The resulting DNA sequences were analyzed by BLAST with NCBI.
2.4. Preparation of Endophyte Extracts
Agar pieces (0.5 cm2) with actively growing fungal endophytic colonies were inoculated onto sterile potato dextrose broth (1 l). The flasks were incubated with alternate light (8 hours) and dark cycles (16 hours) in a stationary phase (25°C ± 2°C) for 15 days. The mycelial mat in the broth was sonicated and the whole mass was filtered. For extraction of bioactive ingredients, the filtrate was transferred to a separating funnel containing an equal volume of ethyl acetate. The mixture was strongly agitated. The solvent layer on the top was separated and condensed to 10 ml in a Heidolph system, an evaporator with rotations at 42°C [15]. The fraction that was dry was solubilized in methanol (5 ml). It was stored (4°C) in a colored glass container for further experimentation.
2.5. Estimation of Total Phenol Concentration in the Endophyte Extract
The phenol content in endophyte extracts was determined according to the calorimetric method [16]. The extract (100 μl) was mixed with a Folin–Ciocalteu reagent (0.75 ml; 1:10 diluted, HiMedia, India) and incubated (22°C, 5 minutes). Saturated sodium carbonate (60 g/l) was added to neutralize it. Further incubation was carried out in the dark (1.5 hours, 22°C). The blue color was measured at 725 nm (Hitachi U-3900 UV/visible spectrophotometer). Phenol content was estimated using the standard curve of gallic acid (25–250 μg/ml; Sigma Aldrich, St. Louis, MO). The concentration of phenol in the extract was reported as 153.39 gallic acid equivalents (GAE mg/mg extract). All the estimations were carried out in triplicate.
2.6. Estimation of Total Flavonoid Content
The sample extract (100 μl) was combined with the methanolic solution of aluminum chloride (2%; 100 μl). After incubation (30 minutes), the absorbance of the reaction was read at 430 nm. The flavonoid content was reported using the Rutin standard analytical curve (10–100 μg/ml; Sigma Aldrich, St. Louis, MO). The flavonoid concentration in the extract was expressed in mg Rutin (mg/g of extract). All estimations were carried out in triplicate.
2.7. 1,1-Diphenyl-2-Picrylhydrazyl (DPPH) Radical Scavenging Assay
DPPH was obtained from Sigma Aldrich (St. Louis, MO). The procedure for estimation of the antioxidant potential was followed as described by a previously reported protocol [17]. The results were measured at 517 nm using a Spectra max 340 Colorimeter from Molecular devises, USA. A methanolic DPPH solution with radicals (300 μM) was prepared. The final mixture included an extract (5 μl) mixed with a DPPH solution (95 μl) at 37°C for 30 minutes. The scavenging activity of DPPH radical (percent) was estimated in comparison with methanol (negative control), ascorbic acid, and quercetin (25–250 μg/ml; positive controls; Sigma Aldrich, St. Louis, MO). The values (IC50) representing the concentration of the endophyte extract required to scavenge DPPH (50%) radicals was reported.
2.8. Biological Activity Studies
2.8.1. Agar disk diffusion method
Escherichia coli (MTCC 724) and Pseudomonas aeruginosa (ATCC 27853) were the Gram-negative bacteria used in the study. Staphylococcus aureus (MTCC 96) and Bacillus subtilis (MTCC 441) were the Gram-positive bacteria used in the study. They were obtained from the Institute of Microbial Technology (Chandigarh, India). The endophyte extract (1, 2, 5.0, and 10.0 μg) and positive control (streptomycin 2 μ l of 1 mg/ml; Sigma Aldrich, St. Louis, MO) were used. The samples were introduced on Whatman’s paper disks (10 mm) placed on Mueller Hinton agar as per a previously reported protocol [18]. They were incubated for 15–18 hours at 37°C. The inhibition zone diameter was reported in comparison with streptomycin.
2.8.2. Assay of the loss of biofilm by confocal laser scanning microscopy
The biofilm structure was observed in confocal laser scanning microscopy (CLSM) (LSM 710; Carl Zeiss, Germany) and reported. The experiment was carried out as per a previously reported procedure [19]. The overnight bacterial cultures, Gram-negative bacteria, and P. aeruginosa (ATCC 27853), along with E. coli (MTCC 724), were prepared. A 6-well plate was used for the assay. Cover slips were introduced in all the wells. The media (2 ml) was poured over them. The controls consisted of a cover slip with media. To the experimental wells, overnight-grown bacterial cells (750 μl) were added. The endophyte extracts (200 μl, 750 μg) were then introduced into the wells. The 6-well plates were incubated at 37°C for 24 hours. The cover slips in wells with biofilm formed on them were carefully removed. The biofilm was washed with phosphate buffer saline. The film was stained with ethidium bromide and acridine orange (24 μl; concentration 1 μg in sterile distilled water, 400 μl; HiMedia, India). Furthermore, they were carefully washed (thrice). The cover slips containing stained biofilms were placed on a glass slide in an inverted position. The samples were imaged and CLSM images were presented.
2.8.3. Evaluation of M. souzae extract-induced cell cytotoxicity
2.8.3.1. Animal cell lines and their culturing with 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide assay (MTT) assay. Human cell lines, HeLa (cervical carcinoma), obtained from the National Center for Cell Science (NCCS, Pune, India) were used. They were cultivated in Dulbecco's Modified Eagle Medium media (Sigma Aldrich; St. Louis, MO). Fetal bovine serum was added to it (10%; Gibco BRL, USA). MTT assay of the fungal extract was reported by MTT (Sigma Aldrich, St. Louis, MO) [20]. Three wells were used for each setup in triplicate.
2.8.3.2. DNA fragmentation assay. HeLa cells (5 × 104 cells/ml) were treated with M. souzae extract (60 mg/ml) and camptothecin (2 mg/ml; Sigma Aldrich, St. Louis, MO). The cells (treated and controls) were harvested and a DNA fragmentation procedure was carried out as reported earlier [20].
2.8.3.3. Apoptosis assessed by nuclear morphology. HeLa cells were treated with M. souzae (60 μ g/ml) and camptothecin (2 μ g/ml). They were stained with acridine orange and propidium iodide [Acridine orange/propidium iodide (AO/PI); 1 mg/ml] as reported earlier [20]. Changes after treatments in the apoptotic nuclear morphological were observed by CLSM LSM 710 (Carl Zeiss, Germany).
2.9. Chromatographic Analysis by Thin-Layer Chromatography (TLC)
TLC was carried out on precoated (0.25 mM silica gel) glass plates (10 × 5 cm; 60 F254, Merck, Germany). Prior to use, they was activated in an oven (110°C, 1 hour). The ethyl acetate extract (10 μ , 5 mg/ml) was applied by means of a capillary tube. The standards quercetin and kaempferol (10 μ l and 1 mg/ml, respectively; Merck, Germany) solubilized in methanol were also applied along with the extracts by using the capillary tube. The plates were developed in glass chambers by solvent system hexane–ethyl acetate–methanol (10:75:15; v/v). The secondary metabolites of M. souzae ethyl acetate extract were visualized by spraying with anisaldehyde sulfuric acid (HiMedia, India) with subsequent heating at 115°C. The chromatogram images were visualized under UV365 and the images were captured by a digital camera.
2.10. Statistical Analysis
All the experimental results and reported values were in triplicate. The results are reported as mean ± standard deviation (SD) after subjecting to variance scrutiny, followed by Tukey’s test. This lead us to analyze the differences among the M. souzae ethyl acetate extract and control conditions at p-value < 0.001.
3. RESULTS AND DISCUSSION
Fungal endophytes as asymptomatic symbiotic colonizers are present in the internal tissues of host plants, an association sound to the host, wherein they reside as in several scenarios that are neither mutualistic nor pathogenic, and have been reported with the capacity to produce the same biologically active compound as reported in their host plant. Thus, it resulted in several leads and discoveries of unique secondary metabolites that have been used as drugs [21]. In this direction, our work is on the study of endophytic fungi of W. fruticosa (Linn.) Kurz (family: Lythraceae) tree leaves, an endemic tree from the western coastal region, Karnataka, India, that provides a view into the fungal endophytic consortia.
3.1. Fungal Endophytes by Morphotyping and Molecular Typing
In the present study, the fungal endophytes of the leaves were sporulated and reported by morphotyping (Fig. 1) and molecular typing by analysis of ITS in their DNA sequence. A set of 21 endophytes were obtained from 82 bits of leaves.
DNA from the endophytes was isolated. The ITS region of DNA was amplified. A clear band at 600 bp (Fig. 2) obtained on agarose gel was also reported. The band was purified and sent for DNA sequence analysis. The obtained results of DNA sequences were analyzed by the BLAST tool at NCBI. Woodfordia fruticosa endophytes identified by molecular typing include Mucor spp., Colletotrichum spp., Alternaria spp., Penicillium spp., Fusarium spp., and Aspergillus spp.
 | Figure 1: An approach of the reporting fungal endophyte, M. souzae, from leaf segments of W. fruticosa. [Click here to view] |
 | Table 1: Details of the fungal endophytes from leaves of W. fruticosa. [Click here to view] |
Mucor sp. was identified as M. souzae by the colony, spore morphology (Fig. 1ci, ii and d) and ITS analysis (Table 1). This isolate reported was taken up for further studies. Its ethyl acetate extract was screened for biological activities and secondary metabolites were reported.
3.2. Total Secondary Metabolites (Phenol and Flavonoid) Contents
In the current study, crude extracts from fungal endophytes were screened and our work was the first study reporting antioxidant and antibacterial capacities of the extracts of M. souzae, endophytic fungi of W. fruticosa tree leaves. Endophyte extracts exhibited several biological properties, like antioxidant and antibacterial with biofilm inhibition capacities.
Phenol concentration was expressed as GAE (mg/g of extract). Flavonoid concentration was reported as Rutin equivalents (mg/g of extract). Mucor souzae extract contained a total phenol content of 151.39 ± 0.124 (GAE; mg/g of extract). The total flavonoid content of 72.05 ± 0.32 (Rutin equivalents; mg/g of extract) was reported in the extract (Table 2).
 | Table 2: Total secondary metabolites content (phenol and flavonoid) in the extract of fungal endophytes reported from W. fruticosa. [Click here to view] |
 | Table 3: Scavenging DPPH radical by M. souzae extract of W. fruticosa leaves. [Click here to view] |
3.3. DPPH Radical Scavenging Property of the Extract
The test compounds with hydrogen-donating capacity can decolorize DPPH. The activities were recorded and compared to the reference standards, ascorbic acid and quercetin. Endophyte, M. souzae ethyl acetate extract exhibited DPPH scavenging activity with the IC50 value of 58.64 ± 4.38 μg/ml. The reference compounds, ascorbic acid and quercetin, exhibited IC50 values of 29.31 ± 4.21 and 23.58 ± 3.23 μg/ml, respectively, as reported in Table 3.
3.4. Antibacterial Activity
3.4.1. Agar disk diffusion method
The extracts inhibited bacterial growth resulting in clear ZOI indicating an antibacterial effect. Inhibitions of both Gram-negative and Gram-positive bacteria were recorded (Fig. 3). This antibacterial property of the M. souzae ethyl acetate extract was screened using four microbes (Fig. 3), S. aureus (MTCC 96) and B. subtilis (MTCC 441) (the Gram-positive bacteria) and E. coli (MTCC 724) and P. aeruginosa (ATCC 27853) (the Gram-negative bacteria). The resulting inhibition zones (ZOI) were also recorded. Endophyte extract (10 μ g) significantly arrested the growth of P. aeruginosa (5 mm, ZOI) and S. aureus (2 mm, ZOI), followed by E. coli and B. subtilis (4 and 3 mm, ZOI, respectively).
 | Figure 2: Amplifications of ITS in the DNA of endophytes and its analysis on agarose gel (1% w/v).Lane M: DNA ladder (100–1 kbp); lane 1: Mucor spp; lane 2: Colletotrichum spp; lane 3: Alternaria spp; lane 4: Penicillium spp; lane 5: Fusarium spp; and lane 6: Aspergillus spp. [Click here to view] |
3.4.2. CLSM investigation to record biofilm loss
Gram-negative strains of P. aeruginosa (ATCC 27853) and E. coli (MTCC 724) and the biofilm forming microbes were incorporated for this study and the loss of biofilm was visualized by CLSM. From sample observations of controls, P. aeruginosa (Fig. 4a) and E. coli (Fig. 4c) formed a biofilm on the support, and live cells were visualized and stained green by acridine orange. The addition of the extract (200 μl, 750 μg) resulted in a significant loss of biofilm that was recorded by CLSM. Patches and broken-down biofilm architecture along with diminished microcolonies containing dead cells of P. aeruginosa (Fig. 4b) and E. coli (Fig. 4d) were observed to be red as they were stained with ethidium bromide.
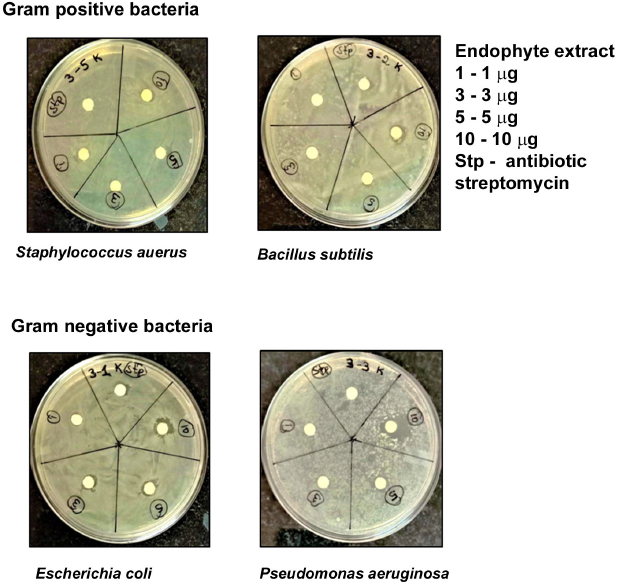 | Figure 3: Antibacterial assay of M. souzae ethyl acetate extract by the disk diffusion method. [Click here to view] |
3.5. Kaempferol and Quercetin Presence in Ethyl Acetate Extract by TLC
Woodfordia fruticosa was reported to be rich in secondary metabolites (kaempferol and quercetin). Previously, Mucor fragilis isolated from Sinopodophyllum hexandrum (Royle) Ying rhizomes harvested at high ranges in the Taibai Mountains (China) had the ability to produce podophyllotoxin and kaempferol [22]. In the present study, secondary metabolite from M. souzae extract needed to be characterized. In this direction, we carried out a preliminary study by TLC. Kaempferol (3,4′,5,7-tetrahydroxyflavone) and quercetin (3,3’,4’,5,7-pentahydroxyl-flavone) flavonoids were detected via TLC analysis compared with the standards (Fig. 5). The developed bands were observed under the UV light.
Flavonoids have been reported to show up to a sixfold stronger antibacterial activity in comparison to the standard antibacterial drugs in the market. They are low molecular weight polyphenolics with the core structure being the C6–C3–C6 skeleton. The amphipathic features (hydrophilic and hydrophobic moieties) with substituents (alkyl chains, heterocyclic moieties containing oxygen or nitrogen, and prenyl groups) contribute to antibacterial properties. The mode of action includes cytoplasmic membrane function alterations, inhibition of energy metabolism, and decreased cell attachment resulting in disrupted biofilm formation, changes in permeability of cell membrane, and attenuation of pathogenicity to cite a few [23].
Pure kaempferol exhibited disruption of bacteria. It was carried out by interaction with the head group with a polar charge in the model membrane used in the study under in-vitro conditions [24]. Pure quercetin exhibited its antibacterial capacity by blocking the charges on amino acids in the porins [25].
3.6. Evaluation of M. souzae Extract-Induced Cell Cytotoxicity via Apoptosis
Maximum cell death was observed and IC50 values were reported: M. souzae extract (IC50 61 ± 3.5 μg/ml) and camptothecin (IC50 2.3 ± 1.6 μg/ml). DNA fragmentation experiment results exhibited fragmentation of the same to smaller entities (Fig. 6i). A smear was observed indicating nonspecific DNA degradation. These results corroborate apoptotic induction capacity of M. souzae extract [26]. To confirm apoptosis, HeLa cells were stained with AO and PI. After this dual-stain, an emission of green fluorescent wavelengths by AO and orange fluorescent wavelengths by PI were captured by CLSM. Apoptosis with chromatin condensation was visualized as an orange fluorescence in the nuclear region (Fig. 6ii, b and c) of HeLa cells when they were treated with M. souzae and camptothecin. The control cells were visualized as green, indicating live cells (Fig. 6ia). Chromatin condensation affirmed HeLa cell cytotoxicity by M. souzae extract as an apoptotic affair.
 | Figure 4: Biofilm inhibition assay of M. souzae ethyl acetate extract. Controls, P. aeruginosa (a); E. coli (c); M. souzae extract + P. aeruginosa (b); and M. souzae extract + E. coli (d). [Click here to view] |
 | Figure 5: Thin-layer chromatography detecting the presence of quercetin and kaempferol in M. souzae ethyl acetate extract as visualized in UV. [Click here to view] |
 | Figure 6: Evaluation of M. souzae extract-induced cell cytotoxicity via apoptosis. i) DNA was subjected to fragmentation. The resultant samples were analyzed by agarose gel electrophoresis. Lane M: DNA ladder (1 Kbp); lane 1: DNA of control (only HeLa cells); lane 2: DNA of HeLa cells + M. souzae extract (60 mg/ml); lane 3: DNA of HeLa cells + camptothecin (2 mg/ml). ii) Induction of apoptotic nuclear morphology in HeLa cells by M. souzae extract (iib) and camptothecin (iic) in comparison to control (iia) cells. They were dual-stained with AO and PI and visualized under a confocal laser scanning microscope. V, viable cells; CC, chromatin condensation in apoptotic cells. [Click here to view] |
Cancer cell lines facilitate research on new cytotoxic entities by being useful tools in the anticancer studies and the structure–activity relationship that aids in revealing the potencies of new molecules with therapeutic properties. In this context, studies have revealed the hydroxylation at the C5 position as in the case of quercetin, which can be considered important for cytotoxic activity. Similarly, hydroxylation at 3, 5 7, 4` and its derivative, kaempferol-3-O-p-D-glucopiranoside, have reported cytotoxic capacities [27].
4. CONCLUSION
The current report on the fungal endophyte M. souzae screened from a consortium of endophytes from W. fruticosa tree leaves by its characterization via morphotyping and molecular typing by ITS–DNA sequence analysis adds to the current trend of knowledge with respect to several studies indicating Mucor spp. as a rich source of secondary metabolites [28–30]. However, this study identifies Mucor spp. investigated only at the crude extract level. Although kaempferol and quercetin from ethyl acetate extract were reported by TLC analysis in comparison to the use of standards, further investigations are required to determine other secondary metabolites from M. souzae that could lead to identification of many more new molecules toward therapeutic applications. Chemical investigation to identify potential active ingredients is still ongoing. Data collection is a treasure toward unraveling the capacity of Mucor endophytic species as it is reported to be a true fungal factory required for exploitation of a broad range of applications.
5. ACKNOWLEDGMENTS
KD, PS, SB, KRK, and SS acknowledge the financial support and the experimentation facilities given by the University Grants Commission, New Delhi, Government of India (Grant number UOM/IOE/RESEARCH/I/2010-11, dt 22.04.2010). The authors thank the University Grants Commission, New Delhi, India, for recognizing University of Mysore as an Institution of Excellence. SS planned and supervised the work and finalized the manuscript. KD carried out the extraction, antioxidant study, antibacterial activity, and analyzed the data. KRK, PS, and SB identified fungal strains, conducted ITS experiments, and analysis. Animal cell culture experiments were carried out by SS. All the authors edited the manuscript approved the final version of the manuscript.
6. CONFLICT IN INTEREST
The authors report no financial or any other conflicts of interest in this work.
7. AUTHOR CONTRIBUTIONS
All authors made substantial contributions to conception and design, acquisition of data, or analysis and interpretation of data; took part in drafting the article or revising it critically for important intellectual content; agreed to submit to the current journal; gave final approval of the version to be published; and agree to be accountable for all aspects of the work. All the authors are eligible to be an author as per the international committee of medical journal editors (ICMJE) requirements/guidelines.
8. ETHICAL APPROVALS
This study does not involve experiments on animals or human subjects.
9. DATA AVAILABILITY
All data generated and analyzed are included within this research article.
10. PUBLISHER’S NOTE
This journal remains neutral with regard to jurisdictional claims in published institutional affiliation.
REFERENCES
1. Bhattarai S, Bhuju DR. Medicinal usefulness of Woodfordia fruticosa (Linn.) Kurz of Nepal. In: Rai M, Acharya D, Jose LR (eds.). Ethnomedicinal plants: revitalization of traditional knowledge of herbs, Science Publishers (Edenbridge Ltd, British Channel Islands) Enfield, NH, pp 253–68, 2011.
2. Yoshida T, Takuo O, Aya N. Woodfordins A, B and C, dimeric hydrolyzable tannin from Woodfordia fruticosa flowers. Heterocycles 1989;29:2267–71. CrossRef
3. Yoshida T, Cho T, Aya N, Miyamoto K, Koshiura R, Okuda T. Woodfordin C, a macro-ring hydrolysable tannin dimmer with antitumor activity, and accompanying dimmers from Woodfordia fruticosa flowers. Chem Pharm Bull 1990;38:1211–7. CrossRef
4. Motegi AK, Kuramochi H, Kobayashi F, Ekimoto H, Takahashi K, Kadota S, et al. Woodfruticosin (Woodfordin C) a replacement inhibitor of DNA topoisomerase II: experimental antitumor activity. Biochem Pharmacol 1992;44:1961–5. CrossRef
5. Bernstein ME, Carroll GC. Internal fungi in old-growth Douglas Fir foliage. Can J Bot 1977;55:644–53. CrossRef
6. Zhao J, Shan T, Mou Y, Zhou L. Bioactive compounds from plant-derived endophytic fungi. Mini Rev Med Chem 2011;11:159–68. CrossRef
7. Stierle A, Strobel G, Stierle D. Taxol and taxane production by an endophytic fungus yew of Pacific, Taxomyces andreanae. Science 1993;260:214–6. CrossRef
8. Strobel G, Daisy B. Microbial endophytes bioprospecting and their natural products. Microbiol Mol Biol Rev 2003;67:491–502. CrossRef
9. Stierle AA, Stierle DB. Bioactive compounds from four endophytic Penicillium sp. isolated from the yew tree of Northwest Pacific. In: Rahman A (ed.). Bioactive natural products, Elsevier Science Publishers, Amsterdam, Netherlands, vol. 24, pp 933–78, 2000. CrossRef
10. Schulz B, Wanke U, Draeger S, Aust HJ. Effectiveness of surface sterilization of endophytes from herbs and shrubs. Mycol Res 1993;97:1447–50. CrossRef
11. Barnett HL, Hunter BB. Imperfect fungi illustrated genera. III edition, Burgess Publishing Co., Minneapolis, MN, vol. 241, pp 1–32, 1972.
12. Rigerte L, Blumenstein K, Terhonen E. Identification of root endophytes by new R-based methodology to optimize the against Heterobasidion parviporum. Microorganisms 2019;7:1–19. CrossRef
13. Doyle JJ, Doyle JL. DNA isolation by rapid method in little quantities from fresh leaf tissue. Phytochem Bull 1987;19:11–5.
14. White TJ, Bruns T, Lee S, Taylor JW. Direct sequencing of fungal ribosomal RNA genes for phylogenetics and its amplification. In: Gelfand DH, Sninsky JJ, Innis MA, White TJ (eds.). A guide to methods and applications of PCR protocols, Academic Press, New York, NY, pp 315–22, 1990. CrossRef
15. Li G, Kusari S, Lamshöft M, Schüer A, Laatsch H, Spiteller M. Antibacterial secondary metabolites from an endophytic fungus, Eupenicillium sp. LG41. J Nat Prod 2014;77:2335–41. CrossRef
16. Sekhar S, Karmakar R, Prakash HS. Curative properties of Buchanania lanzan: as evaluated by its anti-oxidant, anti-inflammatory and DNA protective properties. J Nat Pharm 2012;3:71–7. CrossRef
17. Brand-Williams W, Cuvelier ME, Berset C. Use of a free radical method to gauge antioxidant activity. LWT 1995;28:25–30. CrossRef
18. Klan?cnik A, Piskernik S, Jeršek B, Možina SS. Diffusion and dilution evaluation for antibacterial activity of plant extracts. J Microbiol Methods 2010;81:121–6. CrossRef
19. Zhang W, Wang Y, Tian RM, Bougouffa S, Yang B, Cao HL, et al. Species sorting during biofilm assembly by artificial substrates during a cold seep system. Sci Rep 2015;4:6647–54. CrossRef
20. Kumari M, Taritla S, Sharma A, Jayabaskaran C. Antiproliferative and antioxidative bioactive compounds in extracts of marine-derived endophytic fungus Talaromyces purpureogenus. Front Microbiol 2018;9:1–12. CrossRef
21. Chandra S. Novel sources of anticancer lead molecules from endophytic fungi. Appl Microbiol Biotechnol 2012;95:47–59. CrossRef
22. Huang J, Zhang J, Zhang XR, Zhang K, Zhang X, He XR. A unique source (Mucor fragilis) of the key pharmaceutical agents podophyllotoxin and kaempferol, Pharma Biol 2014;52:1237–43. CrossRef
23. Farhadi F, Khameneh B, Iranshahi M, Iranshahy. Antibacterial activity of flavonoids and their structure–activity relationship: an update review. Phytother Res 2018;33:13–40. CrossRef
24. He M, Wu T, Pan S, Xu X. Antimicrobial mechanism of flavonoids against ATCC 25922, E. coli by model membrane study. App Surface Sci 2014;305:515–21. CrossRef
25. Alvarez M, Debattista N, Pappano N. Antimicrobial activity and synergism of some substituted flavonoids. Folia Microbiol 2008;53:23–8. CrossRef
26. Kuriakose GC, Lakshmanan DM, Arathi BP, Kumar HRS, Krishna ATH, Ananthaswamy K, et al. Extract of Penicillium sclerotiorum an endophytic fungus isolated from C. fistula L. causes cell cycle arrest and apoptosis in human cervical cancer cells by mitochondrial membrane depolarization. Biomed Pharmacother 2018;105:1062–71. CrossRef
27. Lopez-lazaro M, Galvez M, Martin-Cordero C, Ayuso MJ. Cytotoxicity of flavonoids on neoplastic cell lines. Structure-activity relationship. Stud Nat Prod Chem 2002;27:891–1291. CrossRef
28. Ferreira JA, Lennartsson PR, Edebo L, Taherzadeh MJ. Present status and future prospects of zygomycetes-based biorefinery. Bioresour Technol 2013 135:523–32. CrossRef
29. Detha AIR, Datta FU. Antimicrobial activity of traditional wines (Sopi and Moke) against Salmonella sp. and Escherichia coli. J Adv Vet Anim Res 2016;3:282–5. CrossRef
30. Detha A, Sudarwanto M, Latif H, Datta FU, Rahayu P. Fractionation and identification antimicrobial activity of Sumba mare milk protein against causative agent of subclinical mastitis. Glob Vet 2013;11:674–80.