1. INTRODUCTION
Phosphorus is an important macronutrient for all living beings. Its assimilation by microorganisms depends on the specific enzymes, as phosphatases (EC 3.1.3), which are able to hydrolyze phosphorus-containing molecules. Organic phosphorus is a common source of phosphorus in soil and for many of these compounds, the phosphorus is linked to a carbon atom through the ester linkage, which can be hydrolyzed by phosphatase action, allowing the mineralization of organic phosphate when levels of free inorganic phosphate in soil are low [1,2]. In general, these enzymes hydrolyze esters and anhydrides of phosphoric acid releasing inorganic phosphate throughout different mechanisms of hydrolysis and substrate specificities [3].
Considering the classic way, phosphatases can be separated into alkaline (EC 3.1.3.1) and acid phosphatases (EC 3.13.2), according to the optimal pH of activity [4]. According to their molecular characteristics, these enzymes are also classified as (1) alkaline phosphatase, (2) high molecular mass acid phosphatase (greater than 90 kDa), (3) low molecular mass acid phosphatase (less than 50 kDa), (4) purple acid phosphatase (metalloenzyme) and (5) V-protein phosphatase [5]. Alkaline phosphatase hydrolyzes different natural and synthetic substrates as adenosine triphosphate (ATP), adenosine diphosphate (ADP), adenosine monophosphate (AMP), p-nitrophenyl phosphate, glucose-6-phosphate, glucose-1-phosphate, glyceraldehyde-3-phosphate and pyrophosphate, due to its capacity to act in alkaline pH [6].
Phosphatases are enzymes with multiple functions in living organisms and widely distributed in both prokaryotic and eukaryotic organisms [7]. In this context, filamentous fungi are interesting microorganisms because they contribute with different ecological process as the decomposition of organic molecules and transformation of N and P, among others [1]. For this purpose, fungi are able to produce a myriad of ectoenzymes, including alkaline phosphatases used in the mineralization of organic phosphorus. The phosphatase activities observed in soils are the result of the extracellular enzymes accumulated in soil particles and intracellular enzymes from microorganisms [8]. The alkaline phosphatase is an important enzyme for mineralization process in different environments, especially in alkaline and phosphorus-limited soils [9]. Some alkaline soils as siderite, found in the environment as clay ironstone, are Fe-rich and can form extensive stratified beds [10,11]. Enzymes with the potential to mineralize these types of soils are an interesting focus of study and, according to our knowledge, no fungal alkaline phosphatase activated by iron has been described until this moment.
The investigation on the ability of fungal strains to produce alkaline phosphatases can be performed using submerged fermentation, as demonstrated by Coban and Demirci [12] for Aspergillus ficuum and by Suresh and Das [13] for Aspergillus fumigatus. As advantages, the composition of the culture medium for SbmF can be precisely defined; the fermentative parameters as temperature and pH are easily controlled, facilitating the scale up of the process; facility of sterilization reducing the contamination; high yield of the product [14,15]. The use of SbmF allows the control of the inorganic phosphate (Pi) concentration in the medium. Pi is an important macronutrient for all organisms and in fungi, the Pi regulates the expression of phosphate responsive genes as PHO genes for phosphatases [16]. Different carbon sources have been used for this purpose, including agroindustrial byproducts and residues as sugarcane bagasse, oatmeal and wheat bran [17]. In general, another aspect to be highlighted is the biotechnological potential of the alkaline phosphatases to be used in molecular biology experiments, removing 5′-phophate groups in the plasmids to block its recircularization, as well as in clinical diagnosis and as biosensor [18].
For an effective understanding of the physiological importance of the phosphatases in the mineralization process, as well as for an effective biotechnological application, the biochemical characteristics of the enzyme should be determined. According to this, the article describes the purification and characterization of an iron-activated alkaline phosphatase produced by the filamentous fungus Rhizopus microsporus var. microsporus.
2. MATERIALS AND METHODS
2.1. Culture Conditions and Preparation of Crude Extracts
The filamentous fungus R. microsporus var. microsporus (LH0706) is deposited in the Laboratory of Microbiology of the Faculty of Philosophy, Sciences and Letters of Ribeirão Preto, São Paulo, Brazil. The microorganism was grown in sterile PDA medium (potato dextrose agar) at 30°C for 6–8 days and stored in a refrigerator at 4°C until the use. Submerged fermentation (SbmF) was performed in 125-ml Erlenmeyer flasks containing 25 ml of Khanna medium [19] added with 0.4 mmol l-1 KH2PO4 at pH 6.3 and supplemented with 0.5% (m/v) rye flour as additional carbon source. The media were autoclaved at 120°C, 1.5 atm for 25 minutes and inoculated with 106 spores ml-1. Afterward, the cultures were maintained under orbital agitation (100 rpm) at 32°C for 76 hours. After this time, the cultures were harvested using No.1 Whatman filter paper and a vacuum pump. The free cell filtrate was dialyzed against distilled water for 24 hours at 4°C and used for the determination of the extracellular alkaline phosphatase activity and purification procedure.
2.2. Determination of Alkaline Phosphatase Activity
The alkaline phosphatase activity was determined discontinuously using 2.7 mmol l-1 p-nitrophenylphosphate (p-NPP) as substrate. The reaction mixture was composed of 100 μl of enzyme sample and 100 μl of p-NPP in 100 mmol l-1 Tris-HCl buffer, pH 8.5. The reaction was conducted at 45°C for different periods and it was stopped with 2 ml of a saturated solution of sodium tetraborate. The p-nitrophenolate ion was estimated at 405 nm. One unit of enzyme activity (U) was defined as the amount of enzyme required to hydrolyze one micromole of substrate per minute under the assay conditions. All experiments were carried out as triplicates and the values were expressed as the mean ± standard error of the mean.
2.3. Protein Quantification and Carbohydrate Content
Total protein was quantified according to the method described by Bradford [20] using bovine serum albumin (BSA) (20–200 mmol l-1) as standard, and expressed as milligram of protein per ml of sample. The specific enzyme activity was expressed as U/mg of protein.
The carbohydrate quantification was performed according to Dubois et al. [21] using phenol sulfuric acid method with mannose as standard and expressed as percentage of milligram of carbohydrate content per mg of protein.
All experiments were carried out as triplicates and the values were expressed as the mean ± standard error of the mean.
2.4. Purification of Alkaline Phosphatase
The crude filtrate containing alkaline phosphatase was loaded on DEAE-Cellulose chromatographic column (2 × 12 cm) equilibrated with 10 mmol l-1 Tris-HCl buffer, pH 7.5 (buffer A) and eluted by a linear gradient of NaCl (0–500 mmol l-1) in the same buffer. The 3 ml fractions were collected at a flow rate of 0.9 ml minutes-1. Fractions with alkaline phosphatase activity were pooled (25 ml), dialyzed against 5 l distilled water at 4°C for 24 hours, with 3 water exchanges, lyophilized and suspended in the buffer A added with 500 mmol l-1 NaCl, and loaded on Sepharose CL-6B chromatographic column (1 × 70 cm) equilibrated using the same buffer. The 1-ml fractions were eluted using the same buffer at a flow rate of 0.3 ml minutes-1. The fractions with phosphatase activity were pooled (16 ml), dialyzed for 24 hours against 5 l distilled water at 4°C, with 3 water exchanges, and used for electrophoretic analysis and enzyme characterization.
2.5. Denaturing Electrophoresis (SDS-PAGE)
The purified samples were analyzed by polyacrylamide gel electrophoresis in denaturing conditions (12% SDS-PAGE) [22] using the PowerPac™ (Bio-Rad®) power supply adjusted to 100 v and 40 mA. 187 μg of the purified protein sample were applied. After running for 90 min, the protein band in the gel was stained with Coomassie Blue Silver G-250 solution [23]. The Precision Plus Protein “Kaleidoscope” Standards (BIO-RAD®) was used as molecular mass marker with molecular mass ranging from 10 to 250 kDa.
2.6. Determination of the Native Molecular Mass
The native molecular mass for alkaline phosphatase was estimated using the Sepharose CL-6B chromatographic column (1 × 70 cm) as described above. The β-amylase (200 kDa), alcohol dehydrogenase (150 kDa), BSA (66 kDa) and carbonic anhydrase (29 kDa) were used as molecular mass makers.
2.7. Influence of the Temperature and pH on Enzyme Activity
Influence of the temperature on the activity of alkaline phosphatase was evaluated using temperature ranging from 30°C to 70°C, at pH 8.0. The thermal stability, using aqueous solution, was determined at a temperature ranging from 40°C to 60°C, for different periods (5–280 minutes). Aliquots were taken off for each predetermined period and kept in an ice bath until the enzyme assay.
The influence of pH on enzyme activity was analyzed using 100 mmol l-1 citric acid buffer (pH 3.0–4.0), 100 mmol l-1 sodium acetate buffer (pH 4.5–6.5) and 100 mmol l-1 Tris-HCl buffer (pH 7.5–9.0), at 45°C. The stability to pH was assayed using the same buffers at a pH range from 3.0 to 10.0 at 4°C for different periods (5 minutes to 24 hours). The enzymatic samples were added in the ratio of 1:1 (v/v) to the buffers. After incubation, aliquots of 100 μl were used to determine the residual enzymatic activity.
All experiments were carried out as triplicates and the values were expressed as the mean ± standard error of the mean.
2.8. Effect of Different Compounds on Alkaline Phosphatase Activity
The effects of chemical compounds on alkaline phosphatase activity were determined by adding 1 mmol l-1 of salts (AgNO3, BaCl2, CaCl2, CoCl2, CuCl2, CuSO4, FeCl3, Fe2(SO4)3, HgCl2, KCl, KH2PO4, MgCl2, MgSO4, MnCl2, MnSO4, NaCl, NaF, NH4Cl, NH4F, and ZnSO4), β-mercaptoethanol, ethylenediaminetetraacetic acid (EDTA), SDS and urea, 0.01% and 0.02% (v/v) of Triton X-100 and Tween-20, and 1% and 2% (v/v) of acetone, acetonitrile, ethanol, isopropanol, methanol and n-butanol. The enzymatic reactions were conducted at 45°C and pH 8.5. All experiments were carried out as triplicates and the values were expressed as the mean ± standard error of the mean.
2.9. Hydrolysis of Different Substrates by Alkaline Phosphatase
The hydrolytic activity of the alkaline phosphatase on the substrates β-glycerophosphate, ATP, phytic acid, D-fructose-6-phosphate, D-glucose-1-phosphate, D-glucose-6-phosphate and p-NPP was analyzed according to the Heinonen and Lahti methodology [24]. The inorganic phosphorus released was estimated at 355 nm. One unit of enzyme activity was defined as the amount of enzyme required to produce 1 μmol of inorganic phosphorus per minute under the assay conditions. All experiments were carried out as triplicates and the values were expressed as the mean ± standard error of the mean.
2.10. Determination of Kinetic Parameters
The Michaelis-Menten constant (Km), maximum velocity (Vmax) and kcat/Km were determined for the purified alkaline phosphatase using p-NPP (0.1–2.5 mmol l-1) as substrate in both, presence and absence of FeCl3. The reactions were conducted at 45°C, pH 8.5. The values of Km, Vmax and kcat were determined using the SigrafW® software [25]. All experiments were carried out as triplicates and the values were expressed as the mean ± standard error of the mean.
3. RESULTS
3.1. Enzyme Production and Purification
The Figure 1A depictures the DEAE-Cellulose chromatographic profile for the extracellular alkaline phosphatase. A single peak with enzymatic activity was eluted with 141 mmol l-1 of NaCl demonstrating that alkaline phosphatase interacted with the resin. Fractions with alkaline phosphatase activity were pooled, dialyzed, lyophilized and loaded into Sepharose CL-6B chromatographic column, and a single peak with alkaline phosphatase activity was observed (Figure 1B). After these chromatographic procedures, the alkaline phosphatase was purified 9.9-fold with recovery of 13% (Table 1).
3.1. Molecular Mass Determination and Carbohydrate Content
The native molecular mass of the alkaline phosphatase was 123 ± 1.8 kDa estimated with gel filtration using Sepharose CL-6B chromatographic column. In addition, the Figure 2 depicts the 12% sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE) profile for the purified alkaline phosphatase through DEAE-Cellulose and Sepharose CL-6B chromatographic columns. A single protein band with apparent molecular mass of 128 ± 0.7 kDa can be observed (lane C).
The carbohydrate content was estimated as 57 ± 0.6% characterizing the alkaline phosphatase as a glycoprotein.
3.3. Effects of Temperature and pH on Alkaline Phosphatase Activity
The activity of the purified alkaline phosphatase produced by R. microsporus var. microsporus was the highest at 45°C (Figure 3A). The purified alkaline phosphatase was able to maintain its activity above 80 ± 0.6% for 210 minutes at 40°C (Figure 3B). Phosphatase activity was reduced to 55 ± 1.8% in the same period at 45°C. When the phosphatase was maintained at 50°C and 60°C, the half-life (t50) was 40 and 5 minutes, respectively.
According to the Figure 3C, the high alkaline phosphatase activity was obtained at pH 8.5. Considering the enzyme stability to pH (Figure 3D), the enzymatic activity of the alkaline phosphatase was kept close to 90 ± 0.8% at pH 8.5 throughout the period analyzed. At pH 10.0, the enzymatic activity was reduced with t50 of 4 hours.
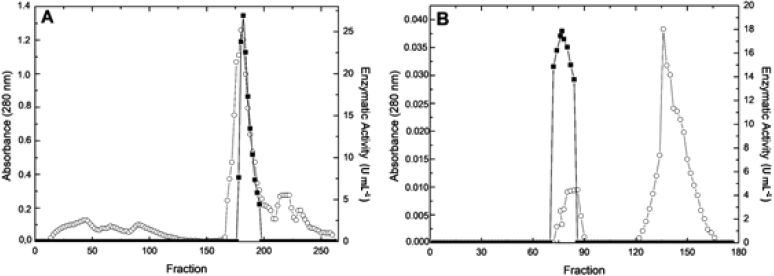 | Figure 1: DEAE-Cellulose (A) and Sepharose CL-6B (B) chromatographic profile for the extracellular alkaline p-nitrophenylphosphatase activity from R. microsporus var. microsporus. (?) alkaline p-nitrophenylphosphatase activity; (?) Absorbance 280 nm. [Click here to view] |
 | Table 1: Purification of extracellular alkaline phosphatase produced by R. microsporus var. microsporus. [Click here to view] |
3.5. Effect of Different Compounds on Alkaline Phosphatase Activity
The enzymatic activity of alkaline phosphatase was not significantly altered by most of the different salts used at 1 mmol l-1 (Table 2). However, the enzyme activity was inhibited 37%–39% in the presence of AgNO3, CuCl2 and CuSO4, 56 ± 0.5% in the presence of NH4F, and 79 ± 0% by NaF. KH2PO4 also inhibited the phosphatase activity (-54 ± 0.5%). The 1 mmol l-1 FeCl3 promoted an increase (+12 ± 1.6%) in enzyme activity.
According to this, the influence of different FeCl3 concentrations on enzyme activity was analyzed (Figure 4). The addition of 0.4 mmol l-1 FeCl3 stimulated the enzyme activity 1.2-fold if compared to the analysis without its addition.
The alkaline phosphatase activity was kept above 80 ± 1.6% in the presence of EDTA and β-mercaptoethanol. In addition, the activity was significantly inhibited by 1% acetonitrile (-30 ± 1.7%), but kept its activity above 80% for the other solvents. At 2% of solvents, the enzyme activity was reduced to 50%–54% except for the isopropanol (Table 3). The enzyme also kept its activity above 80% in both concentrations of the surfactants Triton X-100, SDS and Tween 20.
3.7. Hydrolysis of Different Substrates by Alkaline Phosphatase and Kinetic Parameters
The alkaline phosphatase from R. microsporus var. microsporus was able to hydrolyze different substrates (Table 4). The highest hydrolysis was observed for the synthetic substrate p-NPP (6.2 ± 0.8 U ml-1), followed by ATP (5.7 ± 0.9 U ml-1) and glucose-1-phosphate (4.2 ± 0.3 U ml-1). Hydrolysis of the phytic acid was reduced (0.7 ± 0.1 U ml-1). The hydrolysis of the substrate D-glucose-1-phosphate was higher than that observed for D-glucose-6-phosphate and D-fructose-6-phosphate.
The Km and Vmax values for the hydrolysis of the p-NPP by the alkaline phosphatase were estimated as 0.45 mmol l-1 and 3.95 U/mg of protein, respectively, in the absence of FeCl3, and 0.38 mmol l-1 and 5.12 U/mg of protein in the presence of this salt. The catalytic efficiency (kcat/Km) was 1.52-fold higher in the presence of 0.4 mmol l-1 FeCl3.
4. DISCUSSION
The use of submerged fermentation for the production of phosphatases by filamentous fungi has been reported. For this purpose, different carbon sources have been used, including expensive sources as xylan [26] and p-nitrophenylphosphate [13], and agroindustrial byproducts as wheat bran [17]. In spite of the use of agroindustrial byproducts, this is the first time that rye flour is used as carbon source targeting the production of a fungal alkaline phosphatase. Rye flour presents calcium, potassium, sodium and phosphorus as mineral constituents [27] and the presence of phosphorus was demonstrated to be positive for phosphatase production by R. microsporus [28]. Phosphorus impacts directly upon the production of phosphatases and for many filamentous fungi, the regulation of genes encoding phosphatases is similar to that described for Saccharomyces cerevisae where the PHO genes are upregulated. In the presence of ideal intracellular Pi concentration, the PHO genes are repressed while these genes are expressed under low Pi concentration [29].
 | Figure 2: 12% SDS-PAGE profile for extracellular alkaline phosphatase from R. microsporus var. microsporus purified through chromatographic columns. Proteins were stained by Coomassie Brilliant Blue R-250. Lane A, molecular weight marker BIO-RAD®; lane B, protein bands after DEAE-Cellulose; lane C, alkaline phosphatase band after Sepharose CL-6B (187 μg of protein were applied). [Click here to view] |
The cultivation of the fungus R. microsporus var. microsporus under submerged fermentation by using Khanna medium with 0.4 mmol l-1 KH2PO3, supplemented with 0.5% (w/v) rye flour, pH 6.3, for 76 hours at 32°C, as determined through preliminary studies (unpublished data), allowed the recovery of the alkaline phosphatase activity (21.7 ± 0.6 U ml-1) secreted into the culture broth. However, the fermentative parameters for maximal alkaline phosphatase production changed according to the microbial source. For instance, Guimarães et al. [26] cultured the fungus Aspergillus caespitosus into Segato Rizzatti (SR) medium, pH 6.0, at 40°C for 72 hours, obtaining an activity of 4.5 U ml-1. The fungus Aspergillus flavus Indian Type Culture Collection (ITCC) 6720 produced 12 U ml-1 of phosphatase in Mustard cake medium with initial pH 6.3, at 32°C for 76 hours [30].
The crude filtrate containing alkaline phosphatase was used for purification procedures and after two chromatographic steps, the factor of purification obtained was higher than the value observed (1.5-fold) for the purification of the alkaline phosphatase produced by A. fumigatus, with 46% recovery [13]. On the other hand, the acid phosphatases from A. caespitosus and Aspergillus nidulans were purified 33.4-fold and 120-fold with recovery of 14.7% and 3.8%, respectively [31,32]. Currently, the chromatography is the most powerful procedure for protein isolation considering the selectivity and recovery. However, it can be influenced by different factors, as the presence of particulate material in the samples, protein concentration and its physical-chemical properties, contaminant properties and characteristics of the chromatography medium [33].
The efficiency of the purification procedure was confirmed with SDS-PAGE analysis that presented a single protein band corresponding to the extracellular alkaline phosphatase. The molecular masses for alkaline phosphatase produced by R. microsporus var. microsporus estimated by gel filtration and SDS-PAGE were similar, indicating that this enzyme is a monomer, differing from that reported for the alkaline phosphatases from A. caespitosus (dimer) [26] and Neurospora crassa (tetramer) [34]. The molecular mass of the alkaline phosphatase from R. microsporus var. microsporus were similar to the values obtained for the enzymes produced by Scytalidium thermophilum (132 kDa) [35] and by Humicola grisea var. thermoidea with molecular mass of 126 kDa [36].
Different fungal alkaline phosphatases have also been reported as glycoprotein, as the enzymes produced by A. caespitosus with 56% carbohydrate content [26] and by N. crassa with 40% of carbohydrate content [34], as observed for the alkaline phosphatase from R. microsporus var. microsporus. The glycosylation of proteins is an interesting characteristic that protects the molecule under adverse conditions, as found by extracellular enzymes.
Temperature and pH of the enzyme activity are important parameters to be analyzed aiming not only to characterize the enzyme, but also targeting future applications. Under this view, the optimal temperature observed for the activity of alkaline phosphatase from R. microsporus var. microsporus resembles those described for the enzyme produced by A. fumigatus, which presented better activity between 40 and 45°C [37]. On the other hand, the best alkaline phosphatase activity from Penicillium citrinum was obtained at 60°C [38]. The increase in the temperature affects the degree of agitation of the molecule allowing an increase in the enzyme activity, but above a limit value, the protein molecule is denatured and the catalytic activity is depleted. Around 60%–70% of the enzymatic activity was maintained when the alkaline phosphatase from R. microsporus var. microsporus was incubated at 45°C, an optimal value for the enzyme activity, for long periods.
The optimal value of pH for the alkaline phosphatase activity (8.5) was lower than that reported by Guimarães et al. [35] for the conidial (10.0) and mycelial (9.5) phosphatases from S. thermophilum. On the other hand, phosphatase activity remained strictly acidic for A. fumigatus and P. citrinum, with highest activity at pH 5.5 and 4.8, respectively [37,38]. The pH affects the ionization degree of the molecule increasing or reducing the enzymatic activity. The stability observed at pH 8.5, the best pH value for activity, is attractive, allowing conduction of the reaction under this condition for a long period. Both the stability to temperature and pH can be justified by the high degree of glycosylation of the extracellular alkaline phosphatase from R. microsporus var. microsporus. Enzyme stability is an important parameter to be considered targeting the potential of application. In this sense, the commercial alkaline phosphatase from calf intestine has high specific activity, but its use is limited due to the low thermal stability observed, which hinders the transport, storage and application of this enzyme [39]. The alkaline phosphatase from R. microsporus var. microsporus can be considered as an alternative to overcome this situation.
 | Figure 3: Influence of temperature at pH 8.0 (A) and pH at 45°C (C), thermal (B) and pH stabilities (D) for alkaline phosphatase activity from R. microsporus var. microsporus. 100% corresponds to 14.38 U ml-1 (A); 16.09 U ml-1 (B); 14.38 U ml-1 (C); 12.06 U ml-1 (D). (?) citric acid buffer pH 3.0-4.0, (?) sodium acetate buffer pH 4.5-6.5 and (?) Tris-HCl buffer pH 7.5-9.0. The experiments were performed in triplicate and the standard errors varied between 0 and 1.8. [Click here to view] |
The enzymatic activity can also be affected by different chemical compounds. The divalent ions Mg+2, Zn+2 and Mn+2 have been reported as activators of the alkaline phosphatase activity from H. grisea var. thermoidea and S. thermophilum [35,36], differing from the data obtained for the activity from R. microsporus var. microsporus phosphatase, which was reduced in presence of these ions. Additionally, AgNO3, CuCl2, CuSO4, NH4F and NaF were the main inhibitors of the enzyme activity. NH4+ and NaF were also reported as inhibitors of alkaline phosphatase activity from R. microsporus var. rhizopodiformis and A. fumigatus, respectively [37,40].
On the other hand, addition of FeCl3 increased the alkaline phosphatase activity, especially at 0.4 mmol l-1. That is probably because there are enough ions to supply the enzyme demand, turning the substrate hydrolysis more efficient [41]. Enzyme activation by iron can be an advantage for microorganisms living in the environment containing iron. Fe minerals can be found in reducing conditions such as FeS2 and FeCO3 in acid soil as pyrite and alkaline soil as siderite, respectively [10]. Microorganisms and Fe oxides coexist in soils, providing possibilities of mutual interaction, where minerals can provide nutrients as Fe, P and S, while microbial community acts on mineral weathering [10]. The mineralization of organic phosphorus in soil containing Fe can be facilitated if the phosphatase is activated by Fe in spite of the low concentration of soluble Fe.
The EDTA is able to form stable complexes with various divalent metal ions, but there was no significant reduction in enzymatic activity of the alkaline phosphatase from R. microsporus var. microsporus, demonstrating that this enzyme did not depend on a specific divalent ion. The β-mercaptoethanol acts by disrupting enzyme disulfide bonds that may cause a decrease in the catalytic activity of the enzyme. However, the residual activity remained above 90% with this compound indicating that such interactions are not essential for the catalytic activity. The enzyme activity was not affected substantially by Triton X-100 and Tween 20. Organic solvents may increase the solubility of non-polar substrates and decrease water-dependent secondary reactions and surfactants, interfering in polar and nonpolar bonds of the protein [42].
 | Table 2: Effect of different compounds on alkaline extracellular phosphatase activity from R. microsporus var. microsporus. [Click here to view] |
The influence of different sources of phosphorus on the phosphatase activity has been reported, where inorganic phosphate is able either to inhibit the enzyme activity or to repress the enzyme expression [43]. The sensitivity to the repression by phosphate in an extracellular medium may suggest that this is phosphorus-scavenger enzyme with nutritional role. On other hand, Buainain et al. [36] reported an alkaline phosphatase activity from H. grisea var. thermoidea unaffected by inorganic phosphate, indicating that the alkaline phosphatases can present different functions in the organisms and in the environments.
Phosphatases are known as enzymes able to act on different substrates. Under this aspect, the best action of the enzyme was observed on the synthetic substrate p-NPP. In addition, the hydrolysis of glucose-1-phosphate was higher than that observed for glucose-6-phosphate, indicating a possible preference of the alkaline phosphatase to remove the phosphate groups in the position C-1 of the substrates. The acid phosphatase produced by A. caespitosus and the alkaline phosphatases from H. grisea var. thermoidea and S. thermophilum also presented preference for the synthetic substrate p-NPP [31,35,36].
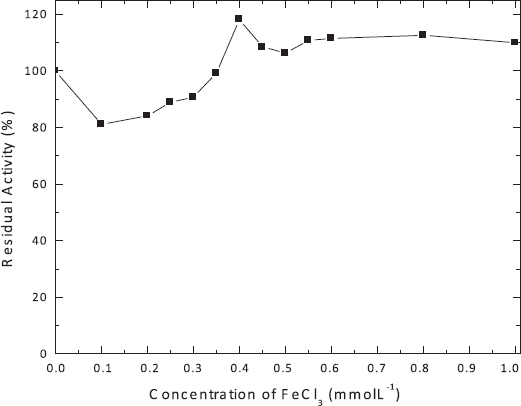 | Figure 4: Influence of different concentrations of FeCl3 on alkaline phosphatase activity produced by R. microsporus var. microsporus. 100% corresponds to 18.41 U ml-1. The experiments were performed in triplicate and the standard errors varied between 0 and 1.0. [Click here to view] |
 | Table 3: Effect of different solvents and surfactants on extracellular alkaline phosphatase activity from R. microsporus var. microsporus. [Click here to view] |
According to the kinetic parameters, the alkaline phosphatase was more efficient when FeCl3 was added to the reaction, with high affinity for the p-NPP as substrate. This fact is reinforced by the kcat value, which was larger in the presence of the salt. Buainain et al. [36] found Km = 0.33 mmol l-1 and Vmax = 89 U/mg of protein for the alkaline phosphatase from H. grisea var. thermoidea, while Morales et al. [44] reported Km = 0.35 mmol l-1 and Vmax = 241 U/mg of protein for the enzyme from N. crassa. Guimarães et al. [35] reported Km = 0.10 and 0.11 mmol l-1, and Vmax = 0.79 and 3.77 U/mg of protein for conidial and mycelial alkaline phosphates produced by S. thermophilum.
 | Table 4: Hydrolysis of different substrates by alkaline phosphatase from R. microsporus var. microsporus. [Click here to view] |
5. CONCLUSION
The fungus R. microsporus var. microsporus was able to produce a monomeric alkaline phosphatase activated by iron, which can be an important advantage for filamentous fungi growing in an iron-rich environment as some alkaline soil types, allowing better phosphorus uptake. The better catalytic efficiency in presence of iron is an evidence for this adaptation. The stability to temperature and pH highlights the potential of the enzyme for future application and indicates possible new studies with applied focus. This aspect is reinforced by the high residual activity observed in presence of solvents, surfactants, β-mercaptoethanol and urea. This first report of an iron-activated alkaline phosphatase is an important contribution for understanding of the phosphatase diversity and future application in the bioremediation of the alkaline soil.
ACKNOWLEDGMENT
The authors kindly acknowledge the financial support by the research scholarships from Coordenação de Aperfeiçoamento de Pessoal de Nível Superior and from Fundação de Amparo à Pesquisa do Estado de São Paulo (Process nº 2016-11311-5). This manuscript is part of the Master´s Degree dissertation from Ornela PHO (2017). We also thank Maurício de Oliveira for the technical assistance.
CONFLICT OF INTEREST
Authors declared that they do not have any conflicts of interest.
REFERENCES
1. Treseder KK, Lennon JT. Fungal traits that drive ecosystem dynamics on lad. Microbiol Mol Biol Rev 2015;79:243–62. CrossRef
2. Ferreira CV, Justo GZ, Souza AC, Queiroz KC, Zambuzzi WF, Aoyama H, et al. Natural compounds as a source of protein tyrosine phosphatase inhibitors: application to the rational design of small-molecule derivatives. Biochimie 2006;88:1859–73. CrossRef
3. Wang YP, Houlton BZ, Field CB. A model of biogeochemical cycles of carbon, nitrogen, and phosphorus including symbiotic nitrogen fixation and phosphatase production. Global Biogeochem Cycles 2007;21:1–15. CrossRef
4. Qureshi AS, Marium S, Khushk I, Bhutto MA. Production of alkaline phosphatase from newly isolated Aspergillus fumigatus EFRL05. Top Class J Microbiol 2013;1:67–73.
5. Mullaney EJ, Ullah AHJ. The term phytase comprises several different classes of enzymes. Biochem Biophys Res Commun 2003;312:179–84. CrossRef
6. Favarin BZ, Andrade MAR, Bolean M, Simão AMS, Ramos AP, Hoylaerts MF, et al. Effect of the presence of cholesterol in the interfacial microenvironment on the modulation of the alkaline phosphatase activity during in vitro mineralization. Colloids Surf B Biointerfaces 2017;155:466–76. CrossRef
7. Kapri A, Tewari L. Phosphate solubilization potential and phosphatase activity of rhizospheric Trichoderma spp. Braz J Microbiol 2010;41:787–95. CrossRef
8. Qin S, Hu1 C, Oenema O. Differentiating intracellular from extracellular alkaline phosphatase activity in soil by sonication. PloS One 2013;8:e58691. CrossRef
9. Acosta-Martínez V, Acosta-Mercado D, Sotomayor-Ramírez D, Cruz-Rodríguez L. Microbial communities and enzymatic activities under different management in semiarid soils. Appl Soil Ecol 2008;38(3):249–60. CrossRef
10. Colombo C, Palumbo G, He JZ, Pinton R, Cesco S. Review on iron availability in soil: interaction of Fe minerals, plants, and microbes. J Soils Sediments 2014;14:538–48. CrossRef
11. Roh Y, Zhang CL, Vali H, Lauf RJ, Zhou J, Phelps TJ. Biogeochemical and environmental factors in Fe biomineralization: magnetite and siderite formation. Clays Clay Miner 2003;51(1):83–95. CrossRef
12. Coban HB, Demirci A. Screening of phytase producers and optimization of culture conditions for submerged fermentation. Bioprocess Biosyst Eng 2014;37:609–16. CrossRef
13. Suresh N, Das A. Purification and characterization of phosphatases from Aspergillus fumigatus for applications in industrial uses. Rasayan J Chem 2014;7:118–28.
14. Singhania RR, Sukumaran RK, Patel AK, Larroche C, Pandey A. Advancement and comparative profiles in the production technologies using solid-state and submerged fermentation for microbial cellulases. Enzyme Microb Technol 2010;46(7):541–9. CrossRef
15. Sadh PK, Kumar S, Chawla P, Duhan JS. Fermentation: a boon for production of bioactive compounds by processing of food industries wastes (by-products). Molecules 2018;23:2560. CrossRef
16. Dick CF, Dos-Santos ALA, Meyer-Fernandes JR. Inorganic phosphate as an important regulator of phosphatases. Enzyme Res 2011;2011:1–7. CrossRef
17. Guimarães LHS, Jorge JA, Terenzi HF, Jamur MC, Oliver C, Polizeli MLTM. Effect of carbon source on alkaline phosphatase production and excretion in Aspergillus caespitosus. J Basic Microbiol 2003;43:210–7. CrossRef
18. Nalini P, Ellaiah P, Prabhakar T, Girijasankar G. Microbial alkaline phosphatases in bioprocessing. Int J Curr Microbiol Appl Sci 2015;4:384–96.
19. Khanna P, Sundari SS, Kumar NJ. Production, isolation and partial purification of xylanases from an Aspergillus sp. World J Microbiol Biotechnol 1995;11:242–3. CrossRef
20. Bradford MM. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 1976;72:248–54.
21. Dubois M, Gilles KA, Hamilton JK, Rebers PT, Smith F. Colorimetric method for determination of sugars and related substances. Anal Chem 1956;28:350–6. CrossRef
22. Laemmli UK. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 1970;227:680–5. CrossRef
23. Candiano G, Bruschi M, Musante L, Santucci L, Ghiggeri GM, Carnemolla B, et al. Blue silver: a very sensitive colloidal Coomassie G-250 staining for proteome analysis. Electrophoresis 2004;25:1327–33. CrossRef
24. Heinonen JK, Lahti RJ. A new and convenient colorimetric determination of inorganic orthophosphate and its application to the assay of inorganic pyrophosphatase. Anal Biochem 1981;113:313–7. CrossRef
25. Leone FA, Baranauskas JA, Furriel RPM, Borin IA. SigrafW: an easy-to-use program for fitting enzyme kinetic data. Biochem Mol Biol Educ 2005;33:399–403. CrossRef
26. Guimarães LHS, Terenzi HF, Jorge JA, Leone FA, Polizeli MLTM. Extracellular alkaline phosphatase from the filamentous fungus Aspergillus caespitosus: purification and biochemical characterization. Folia Microbiol 2003;48:627–32. CrossRef
27. Brites CM, Igrejas G, León AE. Centeno y triticale. In: León AE, Rosell CM (Eds.). De tales harinas, tales panes. Granos, harinas y productos de panificación en Iberoamérica. 1st edition, Hugo Báez Editor, Córdoba, Spain, pp 191–194, 2007.
28. Sato VS, Jorge JA, Guimarães LHS. Characterization of a thermotolerant produced by R. microsporus var. microsporus biofilm on an inert support using sugarcane bagasse as carbon source. Appl Biochem Biotechnol 2016;179:610–24. CrossRef
29. Gomes-Vieira AL, Widemanb JG, Paes-Vieira L, Gomes SL, Richard TA, Meyer-Fernandes JR. Evolutionary conservation of a core fungal phosphate homeostasis pathway coupled to development in Blastocladiella emersonii. Fungal Genet Biol 2018;115:20–32. CrossRef
30. Gaind S, Singh S. Production, purification and characterization of neutral phytase from thermotolerant Aspergillus flavus ITCC 6720. Int Biodeterior Biodegradation 2015;99:15–22. CrossRef
31. Guimarães LHS, Terenzi HF, Jorge JA, Leone FA, Polizeli MLTM. Characterization and properties of acid phosphatases with phytase activity produced by Aspergillus caespitocsus. Biotechnol Appl Biochem 2004;40:201–7. CrossRef
32. Nozawa SR, Macheroni Jr W, Stábeli RG, Thedei Jr G, Rossi A. Purification and properties of pi-repressible acid phosphatases from Aspergillus nidulans. Phytochemistry 1998;49:1517–23. CrossRef
33. Safarik I, Safarikova M. Magnetic techniques for the isolation and purification of proteins and peptides. Biomagn Res Technol 2004;2:7; doi:10.1186/1477-044X-2-7 CrossRef
34. Say JC, Furriel RPM, Ciancaglini P, Jorge JA, Polizeli MLTM, Pizauro JM, et al. Conidial alkaline phosphatase from Neurospora crassa. Phytochemistry 1996;41:71–5. CrossRef
35. Guimarães LHS, Terenzi HF, Jorge JA, Polizeli MLTM. Thermostable conidial and mycelial alkaline phosphatases from the thermophilic fungus Scytalidium thermophilum. J Ind Microbiol Biotechnol 2001;27:265–70. CrossRef
36. Buainain LB, Kadowaki MK, Polizeli ML, Terenzi HF, Jorge JA. Characterization of a conidial alkaline phosphatase from the thermophilic fungus Humicola grisea var. thermoidea. J Basic Microbiol 1998;38:85–94. CrossRef
37. Bernard M, Mouyna I, Dubreucq G, Debeaupuis JP, Fontaine T, Vorgais C, et al. Characterization of a cell-wall acid phosphatase (PhoAp) in Aspergillus fumigatus. Microbiology 2002;148:2819–29. CrossRef
38. Gawas-Sakhalkar P, Singh SM, Simantini N, Ravindra R. High-temperature optima phosphatases from the cold-tolerant Arctic fungus Penicillium citrinum. Polar Res 2012;31:11105. CrossRef
39. Dong G, Zeikus JG. Purification and characterization of alkaline phosphatase from Thermotoga neopolitana. Enzyme Microb Technol 1997;21:335–40. CrossRef
40. Barbosa-Júnior A, Guimarães LHS, Terenzi HF, Jorge JA, Leone FA, Polizeli MLTM. Purification and biochemical characterization of thermostable alkaline phosphatases produced by Rhizopus microsporus var. rhizopodiformis. Folia Microbiol 2008;53:509–16. CrossRef
41. Juma NG, Tabatabai MA. Effects of trace elements on phosphatase activity in soils. Soil Sci Soc Am J 1977;41:343–6. CrossRef
42. Chhokar V, Sangwan M, Beniwal V, Nehra K, Nehra KS. Effect of additives on the activity of tannase from Aspergillus awamori MTCC9299. Appl Biochem Biotechnol 2010;160:2256–64. CrossRef
43. Sánchez S, Demain AL. Metabolic regulation of fermentation processes. Enzyme Microbiol Technol 2002;31:895–906. CrossRef
44. Morales AC, Nozawa SR, Thedei Jr G, Maccheroni Jr W, Rossi A. Properties of a constitutive alkaline phosphatase from strain 74A of the mold Neurospora crassa. Braz J Med Biol Res 2000;33:905–12. CrossRef