1. INTRODUCTION
Biobutanol has gained popularity in the past decade as a renewable source of biofuel owing to its several advantages over conventional liquid fuels as well as the poor state of energy scenario due to depleting fossil fuels and rising demands which is leading to a stark rise in fuel prices. Butanol shows many advantageous properties as a fuel compared to ethanol. Frassoldati et al. [1] discussed the importance of butanol as a prospective substitute for other fuels in the industry. Studies carried out by Shahbakhti et al. [2] reached to a conclusion that the butanol is a potential fuel used as a substitute for traditional fuels, such as petrol and diesel. By 2020, market for butanol is likely to reach $247 billion because of its advantages [3].
There are several number of bacterial species found to be producing solvents acetone, butanol, and ethanol, with altering proportions through ABE fermentation process [4]. These cultures utilize a variety of substrates such as beet molasses, agricultural products, like wheat, rice straw, corn, etc [5,6]. It was concluded on the basis of a study carried out by four species using different substrates that the bacteria C. acetobutylicum was the most appropriate strain for starch containing medium [7]. In recent years, researchers have focused on the genetic engineering aspects for increasing the product yield from the microorganisms used for the fermentation. For instance, Tracy et al. [8] studied the ability of engineered C. cellulolyticum and C. thermocellum to directly convert cellulose to butanol.
For an efficient ABE fermentation, the substrate should be supplemented with extra nutrients, such as vitamins, minerals, and salts which eventually lead to improved production of solvent [9,10]. These nutrients act as enzyme co-factors and contribute in the metabolism process which leads to solvent production.
The previous researchers had innumerable studies on growth media for clostridial culture. The growth media comprises amino acids, vitamins, carbon sources, growth factors, etc. A few of these components are critical for the cell growth. For example, Yeast extract encourages phase shift from acidogenesis to solventogenesis by escalating gene transcription which finally increases butanol production. There are certain factors, such as Zn2+, Fe, and other vitamins found in yeast extract, PABA, iron and zinc salts, etc. which are necessary for the efficient functioning of enzymes involved in clostridial metabolism [11]. For the enzyme Thiolase activation, Haapalainen et al. [12] studied the implication of Cl-and K+ ion which facilitates clostridial metabolism for butanol production.
Optimization of fermentation parameters can be analyzed for improving the biobutanol yield through Central composite design (CCD) of Response surface methodology (RSM). Conventional methods require more time and are not able to demonstrate prospective interactions between the various independent variables of study [13]. Optimization using RSM was used to overcome the various disadvantages associated with the conventional methods for optimization and also saves a lot of time for optimizing the multiple variables. In the past, RSM has been employed for the optimization of biobutanol production [5].
In the present work, first the optimization and comparison of different nitrogen and carbon sources was done using the one variable at a time approach. Following this, the optimization of biobutanol production from macroalgae Gracilaria edulis as the substrate was done using RSM with CCD by considering the effects temperature, pH and glucose concentration as well as their effects on biobutanol production using C. acetobutylicum Microbial Type Culture Collection (MTCC) 11274.
2. MATERIAL AND METHODS
2.1. Culture maintenance and growth
Lyophilized cells of C. acetobutylicum MTCC 11274 was procured from MTCC, IMTECH (Chandigarh, India). The strain was then grown in Reinforced Clostridial Medium and kept for incubation in an incubator shaker (REMI, India) at 120 rpm and 28oC for 72 hours. For the preparation of culture medium, 5 g of RCA medium was dissolved in 100 ml of distilled water in 250 ml Erlenmeyer flask and autoclaved at a temperature of 121oC and 15 lb pressure for 25 minutes.
2.2. Fermentation conditions
ABE fermentation was carried out with the help of clostridial strain C. acetobutylicum MTCC 11274. To maintain anaerobic conditions during fermentation, Sodium Sulfide (1.5%) was supplemented as an oxygen scavenger [14]. P2 medium was used for fermentation having the following composition [15,16]—Glucose 50.0 g/l; Yeast extract 1.0 g/l; K2HPO40.5 g/l; KH2PO40.5 g/l; Thiamin 0.001; PABA 0.001 g/l; Biotin 0.001; MnSO4.7H2O 0.01 g/l; Ammonium acetate 2.2 g/l; Fe SO4.7H2O 0.01 g/l and NaCl 0.01 g/l.
2.3. Optimization of fermentation medium components for biobutanol production
In our study, foremost we considered the impact of particular nutritional factors on biobutanol production by C. acetobutylicum MTCC 11274 using the OVAT, i.e., ‘one-variable-at-a-time’ method. The importance of the consequential effect of these parameters on production of biobutanol was then analyzed using a statistical optimization approach known as RSM. Fermentation was carried out at various different concentrations of carbon and nitrogen sources with the help of C. acetobutylicum MTCC 11274 carried out in 100 ml P2 medium. 2 ml of 2 % Sodium Sulfide was added to maintain anaerobic conditions. The P2 medium containing bottles were autoclaved at 121°C for 15 minutes for sterilization.
2.3.1. Effect of different carbon sources on biobutanol production from C. acetobutylicum MTCC 11274
Different carbon sources Xylose, Starch, Dextrin, Glucose, and Mannose were taken and supplemented in P2 medium and fermentation was carried out for 96 hours in an attempt to find out the best carbon source for production of biobutanol from C. acetobutylicum MTCC 11274.
2.3.2 Effect of different nitrogen sources on biobutanol Production by C. acetobutylicum MTCC 11274
Different nitrogen sources, such as yeast extract, peptone, beef extract, and Soya protein were taken and supplemented in P2 medium and fermentation was done for 96 hours in an attempt to find out the best nitrogen source for production of biobutanol from C. acetobutylicum MTCC 11274.
2.4. Analytical methods
The analysis of biobutanol produced by ABE fermentation was done by spectrophotometric method. It is a cost-effective method for biobutanol detection in comparison to conventional Gas chromatography or HPLC techniques. For this analysis, first the centrifugation of fermented broth was carried out for 10 minutes at 5,000 rpm and the supernatant was employed for detecting the presence of butanol. In analysis, 25% (v/v) butanol was added and results were compared with blank having saturated salt solution and 25% (v/v) of butanol.
The estimation of butanol content in the broth can be carried out by the process of salting out extraction with saturated K3PO4 solution at high pH (>13) followed by measuring the absorbance by employing the diquat reagent. A standard curve was plotted for the estimation of butanol in acetone-ethanol-butanol mixture under optimized conditions [17].
3. RESULTS AND DISCUSSION
3.1. Optimization of fermentation medium components for biobutanol production
3.1.1. Effect of varying carbon sources on biobutanol production from C. acetobutylicum MTCC 11274
In this section, we had attempted to study the effect of varied amount of glucose on clostridial fermentation and production of butanol from C. acetobutylicum MTCC 11274. Different concentrations of glucose (20, 30, 40, 50, 60, 70, and 80 g/l) were supplemented to P2 medium and production of biobutanol was obtained by the OVAT approach. Fermentation was performed for 96 hours at pH 7 and a temperature of 30áµ’C. Supplementation with 20 g/l glucose with C. acetobutylicum MTCC 11274 produced 1.03 g/l of biobutanol. Similarly, fermentation of medium containing enhanced glucose concentration of 30 and 40 g/l produced 2.44 and 3.57 g/l biobutanol, respectively. Increasing glucose concentration further to 50 and 60 g/l increased biobutanol concentration to 5.61 and 6.27 g/l. Furthermore, rise in glucose concentration to 70 and 80 g/l reduced biobutanol content to 5.13 and 2.48 g/l. Therefore, 60 g/l was the optimum glucose concentration for biobutanol production. Figure 1 illustrates the comparative assessment of butanol production at varying glucose concentrations.
We further compared different carbon sources on butanol production from C. acetobutylicum MTCC11274, and it was found that glucose produced optimum amount of biobutanol at 60 g/l concentration. Therefore, further studies were done by taking 60 g/l concentration of carbon sources supplemented in P2 medium and production of biobutanol was done by using the OVAT approach. Fermentation was carried out at a pH 6 and a temperature of 30áµ’C. 60 g/l mannose when used as a carbon source obtained 6.07 g/l biobutanol content after fermentation with C. acetobutylicum MTCC 11274followed by mannose which obtained 5.73 g/l butanol content. Also, 60 g/l starch and dextrin produced 6.12 and 5.53 g/l biobutanol, respectively. Figure 2 illustrates the comparative assessment of butanol production at different carbon sources.
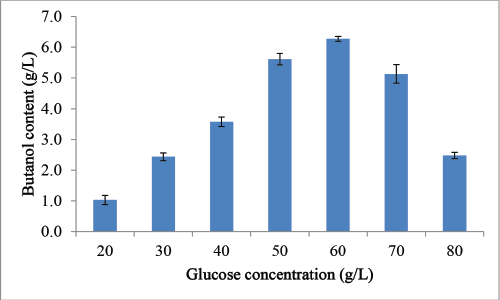 | Figure 1: Biobutanol Production by C. acetobutylicum MTCC 11274 under varied glucose concentration. Experiments were conducted in triplicates value represents average value ± SD. [Click here to view] |
 | Figure 2: Butanol Production by C. acetobutylicum MTCC 11274 at varying carbon sources. Experiments were conducted in triplicates and value represents avg. value ± SD. [Click here to view] |
Clostridial strains are renowned for their potential of utilizing a variety of different sugars. One of the most abundant sugars, available in major quantity in most of the biomass, is glucose [18]. All bacteria utilize the substrate from their environment in order to generate energy, mainly in the form of ATP. All bacterial biosynthetic processes, such as reproduction and respiration require ATP [19]. D–glucose is an important carbohydrate of biological sciences. Cells use it as a source of energy and a metabolic intermediate. Glucose supplemented to complex bacterial growth media is expected to affect the production of a certain protein under the temperature control, which should further affect the ABE solvents yield [20,21].
It has also been reported that during a 96 hours of fermentation with rice bran as a substrate produced 2.31 g/l butanol and 2.74 g/l total solvents. However, during the same experiment with 30 g/l glucose in P2 medium with pretreated rice bran produced 6.27 g/l total ABE solvents and 4.86 g/l butanol [22]. A previous study demonstrated that when 4 g/l in fed-batch cultures of C. acetobutylicum were added per day and 10 g/l glucose in batch cultures were able to produce only acids and there is no shift to ABE solvents [23]. However, it has also been reported in the past that higher glucose concentrations have an inhibitory effect on the production of butanol due to the inhibition of enzyme glucose permease at 12 g/l and above butanol concentration [24].
Clostridial species C. acetobutylicum, C. pasteurianum, and C. beijerinckii were found to ferment both hexose and pentose sugars. However, higher solvent concentrations were obtained from hexoses mainly glucose on the other hand considerably lower solvent concentrations were obtained with pentose sugars, such as xylose [25,26]. A butanol production of 11.25 g/l took place in P2 medium supplemented with 60 g/l of glucose by C. acetobutylicum, whereas only 4.23 g/l butanol was obtained with 36.73 g/l of xylose [27].
3.1.2. Effect of varying nitrogen sources on biobutanol production from C. acetobutylicum MTCC 11274
Present work aimed to study the optimal amount of yeast extract for C. acetobutylicum MTCC 11274 for enhanced butanol production using macroalgae G. edulis based medium. Fermentation of optimized macroalgae at varied yeast extract composition of 1, 2, 3, 4, 5, and 6 mg/l was performed keeping all the other medium constituents constant. At 1, 2, and 3 g/l yeast extract concentration, C. acetobutylicum MTCC 11274 showed a butanol production of 1.17, 3.83, and 5.15 g/l, respectively. Maximum biobutanol production of 7.10 and 7.40 g/l occurred at 4 and 5 g/l yeast extract concentrations. Reduced biobutanol production of 6.52 g/l was seen at a yeast extract concentration of 6 g/l. Figure 3 illustrates the comparative assessment of biobutanol production at varying yeast extract concentrations. To conclude, it was observed that 5.5 mg concentration of yeast extract was most suitable for biobutanol production by C. acetobutylicum MTCC 11274.
 | Figure 3: Comparison of butanol content achieved by fermentation of G. edulis by C. acetobutylicum MTCC 11274 at different yeast extract concentrations. Experiments were conducted in triplicates and value represents average value ± SD. [Click here to view] |
We also studied the effect of varying nitrogen sources on C. acetobutylicum MTCC 11274 and found that yeast extract produced an optimum amount of butanol at 5 g/l yeast extract concentration. Therefore, further studies on other nitrogen sources were done by taking 5 g/l concentration of nitrogen sources supplemented in P2 medium and production of biobutanol was obtained by studying one variable at a time approach. Fermentation was carried out at a temperature of 30áµ’C and 6.0 ± 0.1pH. It was found that the maximum amount of biobutanol content 7.42 g/l was attained with 5 g/l yeast extract. 7.06 and 7.03 g/l butanol content were obtained when 5 g/l peptone and beef extract were used as a nitrogen source. The least amount of butanol content, i.e., 6.44 g/l was obtained from 5 g/l soya peptone as a nitrogen source. Figure 4 illustrates the comparative assessment of biobutanol production at varying nitrogen sources.
Yeast extract is a common nitrogen source which provides proteins, minerals, growth factors, etc. which promotes the microorganism growth and enhances solvent production [28]. Li et al. [29] stated that yeast extract in medium encourages phase shift (acidogenesis to solventogenesis) and thus enhance butanol synthesis. It enhances butanol synthesis by increasing gene transcription 16-folds and indirectly enhances synthesis of butanol in the course of hastening the accumulation of aspartic acid and histidine families. It has been reported in the past that an increase in yeast extract concentration from 5.1 to 7.5 g/l increased butanol yield from 7.6 to 9.9 g/l [30]. Li et al. [30] reported that the presence of yeast extract in the medium encourages phase shift from acidogenesis to solventogenesis and thus enhance butanol synthesis. It enhances butanol synthesis by increasing gene transcription 16 folds and indirectly enhances synthesis of butanol through hastening the accumulation of aspartic acid and histidine families. It has also been reported that maximum biobutanol yield of 0.3054 g/g was attained by C. acetobutylicum from oil palm frond (OPF) juice at a yeast extract concentration of 5.5 g/l after 144 hours of incubation [31].
 | Figure 4: Butanol Production by C. acetobutylicum MTCC 11274 at varying nitrogen sources (5 g/l). Experiments were conducted in triplicates and value represents average value ± SD. [Click here to view] |
Ranjan et al. [32] obtained 6 g/l concentration of biobutanol by addition of 3 g/l yeast extract in P2 medium in a rice straw hydrolysate containing fermentation medium. In another experiment by Ranjan et al. [5] 9.28 g/l butanol concentration was obtained when rice straw hydrolysate containing fermentation medium was supplemented with 2 mg/l PABA + 3 g/l yeast extract [14]. Chua et al. [33] studied effect of yeast extract concentration on biobutanol production and found that increasing its concentration from 0.4% to 1% increased butanol concentration from 8.52 to 8.61 g/l [33].
3.2. Optimization of fermentation parameters for biobutanol production via RSM
Optimization of fermentation parameters were analyzed for improving the biobutanol yield through CCD of RSM. Conventional methods require more time and are not able to demonstrate prospective interactions between the various independent variables of study [34]. Optimization using RSM was used to overcome the various disadvantages associated with the conventional methods for optimization and also saves a lot of time for optimizing the multiple variables.
 | Table 1: Experimental design matrix generated through RSM and biobutanol yield as response. [Click here to view] |
The fermentation parameters temperature, pH and glucose concentration were optimized for biobutanol production using pretreated macroalgae (G. edulis) with C. acetobutylicum MTCC 11274. CCD design was used for optimization of the above-mentioned parameters and low and high values of variables were chosen by performing the steepest ascent experiment (Table 1). The Analysis of variance (ANOVA) model was used for estimating the significance of the model coefficients shown in Table 2.
 | Table 2: ANOVA for optimization of fermentation condition for enhanced biobutanol yield from the pre-hydrolysate of G. edulis. [Click here to view] |
 | Figure 5: Response surface and contour plot for biobutanol yield as a result of the interaction between temperature and pH. [Click here to view] |
 | Figure 6: Response surface and contour plots for biobutanol yield as a result of the interaction between glucose concentration and pH. [Click here to view] |
 | Figure 7: Response surface and contour plots for biobutanol yield as a result of the interaction between temperature and glucose concentration. [Click here to view] |
The coefficient of determination (R2) is 0.9225 and 0.9630, respectively, for both model equations, which means that the model is fit for the variability of the response variable. The adequate precision value measures the signal-to-noise ratio, and a value greater than four is desirable for the appropriate fitness of the quadratic model. The adequate precision value of 15.071 and 10.591 indicates desirable fitness.
Response surface graphs showing the pretreatment as a result of the interaction between temperature and pH (Fig. 5), the interaction between glucose concentration and pH (Fig. 6), and the interaction between glucose concentration and temperature (Fig. 7). Optimized fermentation conditions were 30áµ’C temperature, 40 g/l glucose concentration, and pH 6. p-value showed that the temperature, glucose concentration, and pH have significant effects (p < 0.05) on biobutanol yield.
4. CONCLUSION
In this research work, we have presented a study on the optimization of macroalgae based ABE fermentation process for biobutanol production. The optimized process conditions which were determined with the help of RSM approach help in scaling up the biobutanol production.
Optimization of biobutanol production was carried out by Design of experiments software. The fermentation parameters temperature, pH, and glucose concentration were optimized for biobutanol production using pretreated macroalgae (G. edulis) biomass with C. acetobutylicum MTCC 11274. CCD design was used for optimization of said parameters and low and high values of variables were chosen by performing steepest experiment. The ANOVA was used for estimating the model coefficients significance. It was found that nearly 8.56 g/l biobutanol was obtained via optimization using RSM.
ACKNOWLEDGMENTS
The author is grateful to TEQIP-II (World Bank Project) for providing fellowship during the research work. The authors would also like to acknowledge Aquagri Ltd. Tamil Nadu, India for providing the seaweed sample.
CONFLICT OF INTEREST
The authors have declared no conflict of interest.
REFERENCES
1. Frassoldati A, Grana R, Faravalli T, Ranzi E, Oßwald P and KohsehÖinghaus K. Detailed kinetic- modelling of the combustion of four butanol isomers in premixed low-pressure flames. Combust Flame 2012;159(7):2295–311; doi:10.1016/j.combustflame.2012.03.002
2. Shahbakhti M, Ghazimirsaied A, Audet A, Koch CR. Combustion characteristics of Butanol/n-Heptane blend fuels in an HCCI engine. In Proceedings of Combustion Institute − Canadian Section Spring Technical Meeting Carleton, 2010.
3. Bharathiraja B, Jayamuthunagai J, Sudharsanaa T, Bharghavi A, Praveen kumar R, Chakravarthy M, et al. Biobutanol—an impending biofuel for future: a review on upstream and downstream processing techniques. Renew Sustain Energy Rev 2017;68:788–807; doi:10.1016/j.rser.2016.10.017
4. Beesch SC. Acetone butanol fermentation of sugars. Ind Eng Chem 1952;44:1677–82; doi:10.1021/ie50511a054
5. Ranjan A, Mayank R, Maholkar VS. Process optimization for butanol production from developed rice straw hydrolysate using Clostridium acetobutylicum MTCC 481 strain. Biomass Convers Bior 2013;3(2):143–55; doi:10.1007/s13399-012-0062-2
6. Bergqvist MM, Wardh KS, Das A, Ahlgren EO. A techno-economic assessment of rice husk- based power generation in the Mekong river delta of Vietnam. Int J Energy Res 2008;32:1136–50; doi:10.1002/er.1451
7. Shaheen R., Shirley M., Jones DT. Comparative fermentation studies of industrial strains belonging to four species of solvent producing Clostridia. J Mol Microbiol Biotechnol 2000;2(1):115–24.
8. Tracy B, Jones S, Fast A, Indurthi D, Papoutsakis E. Clostridia: the importance of their exceptional substrate and metabolite diversity for biofuel and bio-refinery applications. Curr Opin Biotechnol 2012;23(3):364–81; doi:10.1016/j.copbio.2011.10.008
9. Moon C, Lee CH, Sang BI, Um Y. Optimization of medium compositions favouring butanol and 1, 3-propanediol production from glycerol by Clostridium pasteurianum. Bioresour Technol 2011;102(22):10561–8; doi:10.1016/j.biortech.2011.08.094
10. Dabrock B, Bahl H, Gottschalk G. Parameters affecting solvent production by Clostridium pasteurianum. Appl Environ Microbiol 1992;58:1233–9.
11. Walter KA, Bennett GN, Papoutsakis ET. Molecular characterization of two Clostridium acetobutylicum ATCC 824 butanol dehydrogenase isozyme genes. J Bacteriol 1992;174:7149–58; doi:10.1128/jb.174.22.7149-7158.1992
12. Haapalainen AM, Meriläinen G, Pirilä PL, Kondo N, Fukao T, Wierenga RK. Crystallographic and kinetic studies of human mitochondrial acetoacetyl-CoA thiolase (T2): the importance of potassium and chloride ions for its structure and function. Biochemistry 2007;46:4305–21.
13. Ibrahim MF, Abd-Aziz S. Razak MNA, Phang LY, Hassan MA. Oil palm empty food bunch as alternative substrate for Acetone-Butanol-Ethanol production by Clostridium butyricum EB6; Appl Biochem Biotechnol 2012;166:1615–25; doi:10.1007/s12010-012-9538-6
14. Potts TM. The production of biobutanol from biomass via a hybrid biological/chemical process. Theses and Dissertations, 2015. Available via http://scholarworks.uark.edu/etd/2050
15. Maiti S, Sarma SJ, Brar SK, Bihan YL, Drougui P, Buelna G, et al. Agro-Industrial Wastes as Feedstock for Sustainable bio-Production of Butanol by Clostridium Beijerinckii. Food Bioprod Process 2016;98:217–26; doi:10.1016/j.fbp.2016.01.002.
16. Khamaiseh EI, Hamid AA, Abdeshahian P, Yusoff WMW, Kalil MS. Enhanced butanol production by Clostridium acetobutylicum NCIMB 13357 grown on date fruit as carbon source in P2 medium. Sci World J 2014;ID 395754; doi:10.1155/2014/395754
17. Maiti S, Saurabh JS, Satinder KB, Yann LB. Novel spectrophotometric method for detection and estimation of butanol in acetone-butanol-ethanol fermenter. Talanta 2015;141:116–21; doi:10.1016/j.talanta.2015.03.062
18. Mitchell WJ, Shaw JE, Andrews L. Properties of the glucose phosphotransferase system of Clostridium acetobutylicum NCIB 8052. Appl Environ Microbiol 1991;57:534–9.
19. Girbal L, Soucaille P. Regulation of Clostridium acetobutylicum metabolism as revealed by mixed–substrate steady–state continuous cultures: role of NADH/NAD ratio and ATP pool. J Bacteriol 1994;176(21):6433–8.
20. Junelles AM, Janati–Idrissi R, Petitdemange H, Gay R. Effect of pyruvate on glucose metabolism in Clostridium acetobutylicum. Biochimie 1987;69(11–12):1183–90.
21. Lehmann D, Radomski N, Lütke–Eversloh T. New insights into the butyric acid metabolism of Clostridium acetobutylicum. Appl Microbiol Biotechnol 2012;96(5):1325–39; doi:10.1007/s00253-012-4109-x
22. Al-Shorgani NKN, Kalil MS, Yusoff WMW. Biobutanol production from rice bran and de-oiled rice bran by Clostridium saccharoperbutylacetonicum N1-4. Bioprocess Biosyst Eng 2012;35:817–26; doi:10.1007/s00449-011-0664-2
23. Fond O, Matta-Ammouri G, Petitdemange H, Engasser JM. The role of acids on the production of acetone and butanol by Clostridium acetobutylicum. Appl Microbiol Biotechnol 1985;22(3):195–200.
24. Ounine K, Petitdemange H, Raval G, Gay R. Regulation and butanol inhibition of D-Xylose and D-Glucose uptake in Clostridium acetobutylicum. Appl Environ Microbiol 1985;49:874–8.
25. Dai Z, Dong H, Zhu Y, Zhang Y, Li Y, Ma Y. Introducing a single secondary alcohol dehydrogenase into butanol-tolerant Clostridium acetobutylicum Rh8 switches ABE fermentation to high level IBE fermentation. Biotechnol Biofuels 2012;5;https://doi.org/5:44; doi:10.1186/1754-6834-5-44
26. Gone F, Bao G, Zhao C, Zhang Y, Li Y, Dong H. Fermentation and genomic analysis of acetone uncoupled butanol production by Clostridium tetanomorphum. Appl Microbiol Biotechnol 2016;100:1523–9; doi:10.1007/s00253-015-7121-0
27. Gu Y, Li J, Zhang L, Chen J, Niu LX, Yang YL, Yang S, Jiang HW. Improvement of D-xylose utilization in Clostridium acetobutylicum via expression of the tal A gene encoding transaldolase from Escherichia coli. J Biotechnol 2009;143(4):284–7; doi:10.1016/j.jbiotec.2009.08.009
28. Shorgani, Hamid AA, Mohtar W, Yosuff W, Kalil MS. Pre optimization of medium for biobutanol production by a new isolate of solvent producing Clostridium. Bioresources 2013;8(1):1420–30.
29. Li M, Liao X, Zhang D, Du G, Chen J. Yeast extract promotes cell growth and induces production of polyvinyl alcohol-degrading enzymes. Enzyme Res 2011;ID 179819:1–8; doi:10.4061/2011/179819
30. Singh KG, Lapsiya KL, Gophane RR, Ranade DR. Optimization for butanol production using Plackett-Burman Design coupled with Central Composite Design by Clostridium beijerenckii strain CHTa isolated from distillery waste manure. J Biochem Technol 2016;7(1):1063–8.
31. Nasrah NSM, Zahari MAKM, Masngut N, Ariffin H. Statistical optimization for biobutanol production by Clostridium acetobutylicum ATCC 824 from Oil Palm Frond (OPF) juice using response surface methodology. MATEC Web Conf 2017;111:13001.
32. Ranjan A, Mayank R, Maholkar VS. Development of semi-defined rice straw- based medium for butanol production and its kinetic study. 3biotech. 2013;3(5):353–64; doi:10.1007/213205-013-0120-x
33. Chua TK, Liang DW, Qi C, Yang KL, He J. Characterization of a butanol acetone producing Clostridium strain and identification of its solventogenic genes. Bioresour Technol 2013;135:372–8; doi:0.1016/j.biortech.2012.08.085
34. Ibrahim MF, Abd-Aziz S, Razak MNA, Phang LY, Hassan MA. Oil palm empty food bunch as alternative substrate for Acetone-Butanol-Ethanol production by Clostridium butyricum EB6. Appl Biochem Biotechnol 2012;166(7):1615–1625. https://doi.org/10.1007/s12010-012-9538-6.